Zihao Tang
Mnemis: Dual-Route Retrieval on Hierarchical Graphs for Long-Term LLM Memory
Feb 17, 2026Abstract:AI Memory, specifically how models organizes and retrieves historical messages, becomes increasingly valuable to Large Language Models (LLMs), yet existing methods (RAG and Graph-RAG) primarily retrieve memory through similarity-based mechanisms. While efficient, such System-1-style retrieval struggles with scenarios that require global reasoning or comprehensive coverage of all relevant information. In this work, We propose Mnemis, a novel memory framework that integrates System-1 similarity search with a complementary System-2 mechanism, termed Global Selection. Mnemis organizes memory into a base graph for similarity retrieval and a hierarchical graph that enables top-down, deliberate traversal over semantic hierarchies. By combining the complementary strength from both retrieval routes, Mnemis retrieves memory items that are both semantically and structurally relevant. Mnemis achieves state-of-the-art performance across all compared methods on long-term memory benchmarks, scoring 93.9 on LoCoMo and 91.6 on LongMemEval-S using GPT-4.1-mini.
Sigma: Differential Rescaling of Query, Key and Value for Efficient Language Models
Jan 23, 2025



Abstract:We introduce Sigma, an efficient large language model specialized for the system domain, empowered by a novel architecture including DiffQKV attention, and pre-trained on our meticulously collected system domain data. DiffQKV attention significantly enhances the inference efficiency of Sigma by optimizing the Query (Q), Key (K), and Value (V) components in the attention mechanism differentially, based on their varying impacts on the model performance and efficiency indicators. Specifically, we (1) conduct extensive experiments that demonstrate the model's varying sensitivity to the compression of K and V components, leading to the development of differentially compressed KV, and (2) propose augmented Q to expand the Q head dimension, which enhances the model's representation capacity with minimal impacts on the inference speed. Rigorous theoretical and empirical analyses reveal that DiffQKV attention significantly enhances efficiency, achieving up to a 33.36% improvement in inference speed over the conventional grouped-query attention (GQA) in long-context scenarios. We pre-train Sigma on 6T tokens from various sources, including 19.5B system domain data that we carefully collect and 1T tokens of synthesized and rewritten data. In general domains, Sigma achieves comparable performance to other state-of-arts models. In the system domain, we introduce the first comprehensive benchmark AIMicius, where Sigma demonstrates remarkable performance across all tasks, significantly outperforming GPT-4 with an absolute improvement up to 52.5%.
Predictive Inference With Fast Feature Conformal Prediction
Dec 01, 2024Abstract:Conformal prediction is widely adopted in uncertainty quantification, due to its post-hoc, distribution-free, and model-agnostic properties. In the realm of modern deep learning, researchers have proposed Feature Conformal Prediction (FCP), which deploys conformal prediction in a feature space, yielding reduced band lengths. However, the practical utility of FCP is limited due to the time-consuming non-linear operations required to transform confidence bands from feature space to output space. In this paper, we introduce Fast Feature Conformal Prediction (FFCP), which features a novel non-conformity score and is convenient for practical applications. FFCP serves as a fast version of FCP, in that it equivalently employs a Taylor expansion to approximate the aforementioned non-linear operations in FCP. Empirical validations showcase that FFCP performs comparably with FCP (both outperforming the vanilla version) while achieving a significant reduction in computational time by approximately 50x. The code is available at https://github.com/ElvisWang1111/FastFeatureCP
Unsupervised dMRI Artifact Detection via Angular Resolution Enhancement and Cycle Consistency Learning
Sep 24, 2024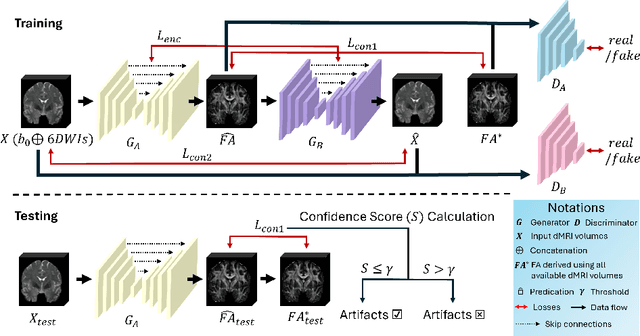
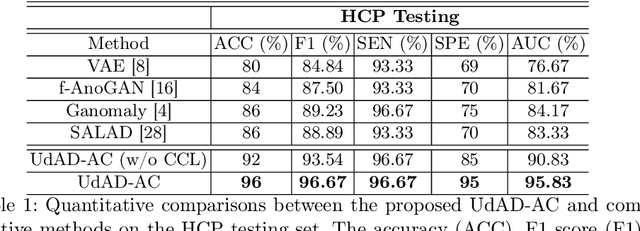
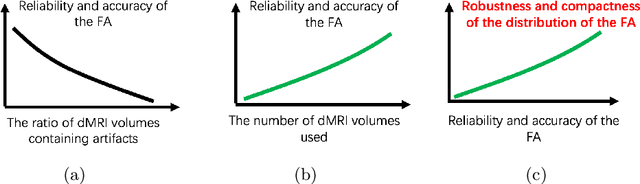
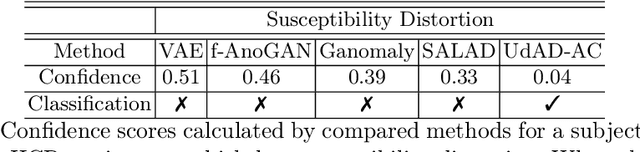
Abstract:Diffusion magnetic resonance imaging (dMRI) is a crucial technique in neuroimaging studies, allowing for the non-invasive probing of the underlying structures of brain tissues. Clinical dMRI data is susceptible to various artifacts during acquisition, which can lead to unreliable subsequent analyses. Therefore, dMRI preprocessing is essential for improving image quality, and manual inspection is often required to ensure that the preprocessed data is sufficiently corrected. However, manual inspection requires expertise and is time-consuming, especially with large-scale dMRI datasets. Given these challenges, an automated dMRI artifact detection tool is necessary to increase the productivity and reliability of dMRI data analysis. To this end, we propose a novel unsupervised deep learning framework called $\textbf{U}$nsupervised $\textbf{d}$MRI $\textbf{A}$rtifact $\textbf{D}$etection via $\textbf{A}$ngular Resolution Enhancement and $\textbf{C}$ycle Consistency Learning (UdAD-AC). UdAD-AC leverages dMRI angular resolution enhancement and cycle consistency learning to capture the effective representation of artifact-free dMRI data during training, and it identifies data containing artifacts using designed confidence score during inference. To assess the capability of UdAD-AC, several commonly reported dMRI artifacts, including bias field, susceptibility distortion, and corrupted volume, were added to the testing data. Experimental results demonstrate that UdAD-AC achieves the best performance compared to competitive methods in unsupervised dMRI artifact detection.
Enhancing Angular Resolution via Directionality Encoding and Geometric Constraints in Brain Diffusion Tensor Imaging
Sep 11, 2024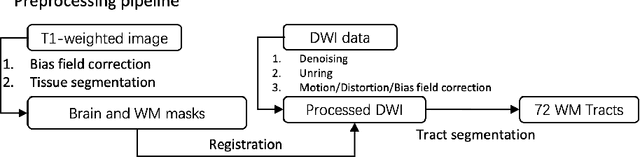

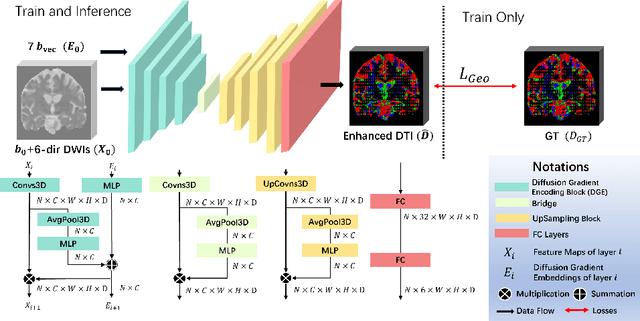

Abstract:Diffusion-weighted imaging (DWI) is a type of Magnetic Resonance Imaging (MRI) technique sensitised to the diffusivity of water molecules, offering the capability to inspect tissue microstructures and is the only in-vivo method to reconstruct white matter fiber tracts non-invasively. The DWI signal can be analysed with the diffusion tensor imaging (DTI) model to estimate the directionality of water diffusion within voxels. Several scalar metrics, including axial diffusivity (AD), mean diffusivity (MD), radial diffusivity (RD), and fractional anisotropy (FA), can be further derived from DTI to quantitatively summarise the microstructural integrity of brain tissue. These scalar metrics have played an important role in understanding the organisation and health of brain tissue at a microscopic level in clinical studies. However, reliable DTI metrics rely on DWI acquisitions with high gradient directions, which often go beyond the commonly used clinical protocols. To enhance the utility of clinically acquired DWI and save scanning time for robust DTI analysis, this work proposes DirGeo-DTI, a deep learning-based method to estimate reliable DTI metrics even from a set of DWIs acquired with the minimum theoretical number (6) of gradient directions. DirGeo-DTI leverages directional encoding and geometric constraints to facilitate the training process. Two public DWI datasets were used for evaluation, demonstrating the effectiveness of the proposed method. Extensive experimental results show that the proposed method achieves the best performance compared to existing DTI enhancement methods and potentially reveals further clinical insights with routine clinical DWI scans.
AuG-KD: Anchor-Based Mixup Generation for Out-of-Domain Knowledge Distillation
Mar 18, 2024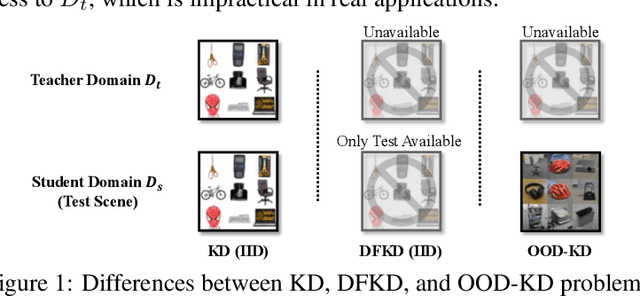
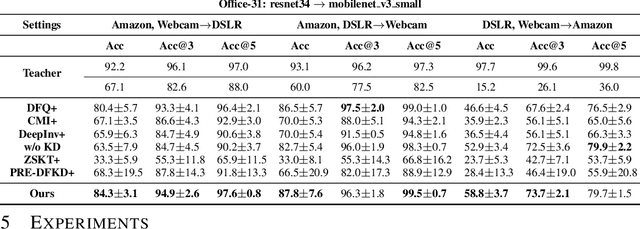
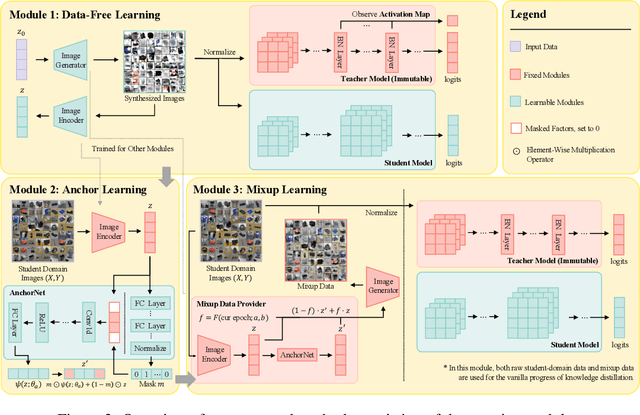
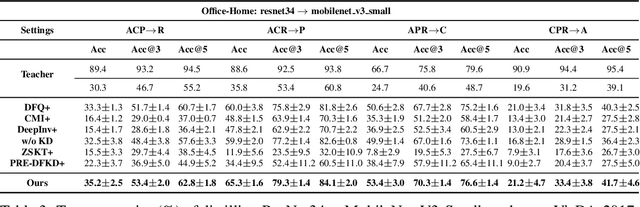
Abstract:Due to privacy or patent concerns, a growing number of large models are released without granting access to their training data, making transferring their knowledge inefficient and problematic. In response, Data-Free Knowledge Distillation (DFKD) methods have emerged as direct solutions. However, simply adopting models derived from DFKD for real-world applications suffers significant performance degradation, due to the discrepancy between teachers' training data and real-world scenarios (student domain). The degradation stems from the portions of teachers' knowledge that are not applicable to the student domain. They are specific to the teacher domain and would undermine students' performance. Hence, selectively transferring teachers' appropriate knowledge becomes the primary challenge in DFKD. In this work, we propose a simple but effective method AuG-KD. It utilizes an uncertainty-guided and sample-specific anchor to align student-domain data with the teacher domain and leverages a generative method to progressively trade off the learning process between OOD knowledge distillation and domain-specific information learning via mixup learning. Extensive experiments in 3 datasets and 8 settings demonstrate the stability and superiority of our approach. Code available at https://github.com/IshiKura-a/AuG-KD .
ModelGPT: Unleashing LLM's Capabilities for Tailored Model Generation
Feb 18, 2024



Abstract:The rapid advancement of Large Language Models (LLMs) has revolutionized various sectors by automating routine tasks, marking a step toward the realization of Artificial General Intelligence (AGI). However, they still struggle to accommodate the diverse and specific needs of users and simplify the utilization of AI models for the average user. In response, we propose ModelGPT, a novel framework designed to determine and generate AI models specifically tailored to the data or task descriptions provided by the user, leveraging the capabilities of LLMs. Given user requirements, ModelGPT is able to provide tailored models at most 270x faster than the previous paradigms (e.g. all-parameter or LoRA finetuning). Comprehensive experiments on NLP, CV, and Tabular datasets attest to the effectiveness of our framework in making AI models more accessible and user-friendly. Our code is available at https://github.com/IshiKura-a/ModelGPT.
Precise Few-shot Fat-free Thigh Muscle Segmentation in T1-weighted MRI
Apr 27, 2023Abstract:Precise thigh muscle volumes are crucial to monitor the motor functionality of patients with diseases that may result in various degrees of thigh muscle loss. T1-weighted MRI is the default surrogate to obtain thigh muscle masks due to its contrast between muscle and fat signals. Deep learning approaches have recently been widely used to obtain these masks through segmentation. However, due to the insufficient amount of precise annotations, thigh muscle masks generated by deep learning approaches tend to misclassify intra-muscular fat (IMF) as muscle impacting the analysis of muscle volumetrics. As IMF is infiltrated inside the muscle, human annotations require expertise and time. Thus, precise muscle masks where IMF is excluded are limited in practice. To alleviate this, we propose a few-shot segmentation framework to generate thigh muscle masks excluding IMF. In our framework, we design a novel pseudo-label correction and evaluation scheme, together with a new noise robust loss for exploiting high certainty areas. The proposed framework only takes $1\%$ of the fine-annotated training dataset, and achieves comparable performance with fully supervised methods according to the experimental results.
TW-BAG: Tensor-wise Brain-aware Gate Network for Inpainting Disrupted Diffusion Tensor Imaging
Oct 31, 2022Abstract:Diffusion Weighted Imaging (DWI) is an advanced imaging technique commonly used in neuroscience and neurological clinical research through a Diffusion Tensor Imaging (DTI) model. Volumetric scalar metrics including fractional anisotropy, mean diffusivity, and axial diffusivity can be derived from the DTI model to summarise water diffusivity and other quantitative microstructural information for clinical studies. However, clinical practice constraints can lead to sub-optimal DWI acquisitions with missing slices (either due to a limited field of view or the acquisition of disrupted slices). To avoid discarding valuable subjects for group-wise studies, we propose a novel 3D Tensor-Wise Brain-Aware Gate network (TW-BAG) for inpainting disrupted DTIs. The proposed method is tailored to the problem with a dynamic gate mechanism and independent tensor-wise decoders. We evaluated the proposed method on the publicly available Human Connectome Project (HCP) dataset using common image similarity metrics derived from the predicted tensors and scalar DTI metrics. Our experimental results show that the proposed approach can reconstruct the original brain DTI volume and recover relevant clinical imaging information.
MS Lesion Segmentation: Revisiting Weighting Mechanisms for Federated Learning
May 03, 2022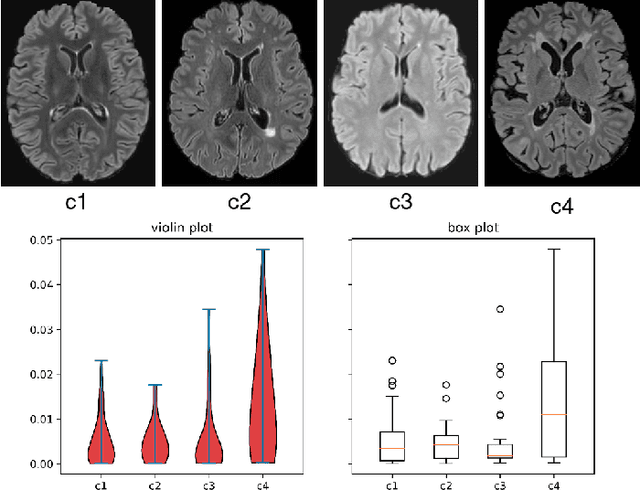
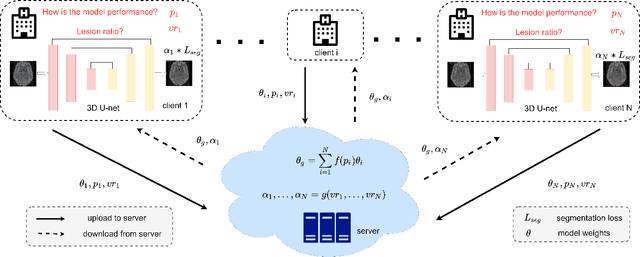
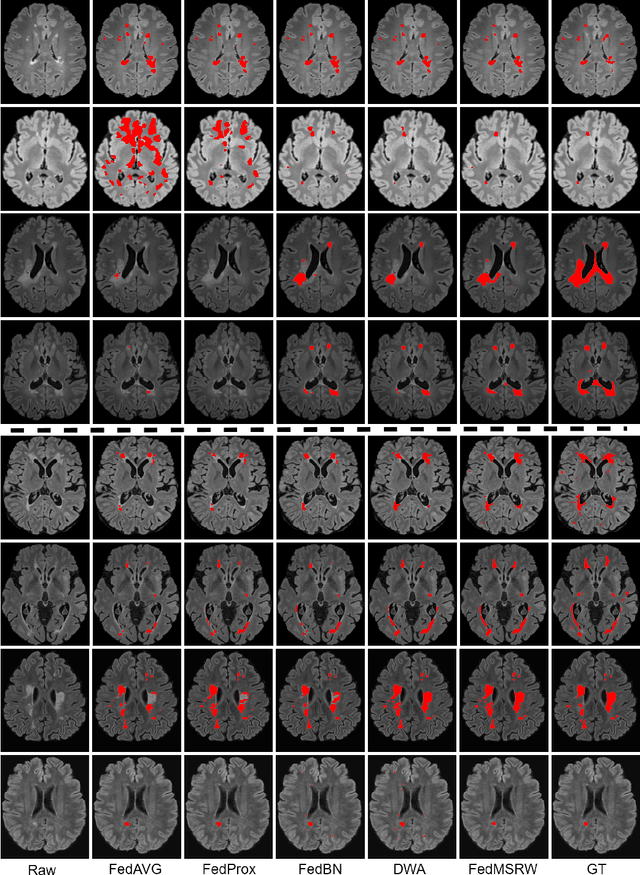
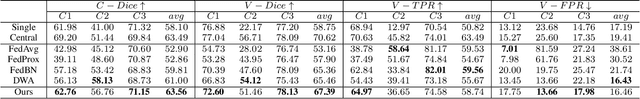
Abstract:Federated learning (FL) has been widely employed for medical image analysis to facilitate multi-client collaborative learning without sharing raw data. Despite great success, FL's performance is limited for multiple sclerosis (MS) lesion segmentation tasks, due to variance in lesion characteristics imparted by different scanners and acquisition parameters. In this work, we propose the first FL MS lesion segmentation framework via two effective re-weighting mechanisms. Specifically, a learnable weight is assigned to each local node during the aggregation process, based on its segmentation performance. In addition, the segmentation loss function in each client is also re-weighted according to the lesion volume for the data during training. Comparison experiments on two FL MS segmentation scenarios using public and clinical datasets have demonstrated the effectiveness of the proposed method by outperforming other FL methods significantly. Furthermore, the segmentation performance of FL incorporating our proposed aggregation mechanism can exceed centralised training with all the raw data. The extensive evaluation also indicated the superiority of our method when estimating brain volume differences estimation after lesion inpainting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge