Michael Barnett
Enhancing Angular Resolution via Directionality Encoding and Geometric Constraints in Brain Diffusion Tensor Imaging
Sep 11, 2024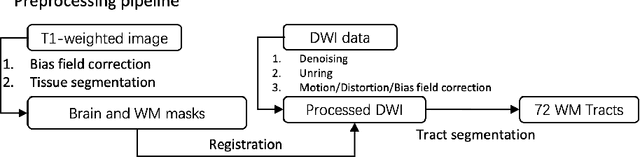

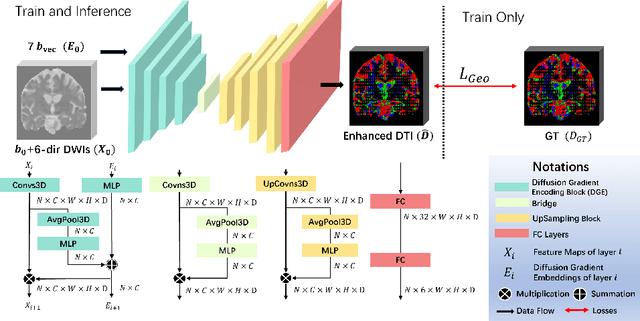

Abstract:Diffusion-weighted imaging (DWI) is a type of Magnetic Resonance Imaging (MRI) technique sensitised to the diffusivity of water molecules, offering the capability to inspect tissue microstructures and is the only in-vivo method to reconstruct white matter fiber tracts non-invasively. The DWI signal can be analysed with the diffusion tensor imaging (DTI) model to estimate the directionality of water diffusion within voxels. Several scalar metrics, including axial diffusivity (AD), mean diffusivity (MD), radial diffusivity (RD), and fractional anisotropy (FA), can be further derived from DTI to quantitatively summarise the microstructural integrity of brain tissue. These scalar metrics have played an important role in understanding the organisation and health of brain tissue at a microscopic level in clinical studies. However, reliable DTI metrics rely on DWI acquisitions with high gradient directions, which often go beyond the commonly used clinical protocols. To enhance the utility of clinically acquired DWI and save scanning time for robust DTI analysis, this work proposes DirGeo-DTI, a deep learning-based method to estimate reliable DTI metrics even from a set of DWIs acquired with the minimum theoretical number (6) of gradient directions. DirGeo-DTI leverages directional encoding and geometric constraints to facilitate the training process. Two public DWI datasets were used for evaluation, demonstrating the effectiveness of the proposed method. Extensive experimental results show that the proposed method achieves the best performance compared to existing DTI enhancement methods and potentially reveals further clinical insights with routine clinical DWI scans.
Symmetry Awareness Encoded Deep Learning Framework for Brain Imaging Analysis
Jul 12, 2024
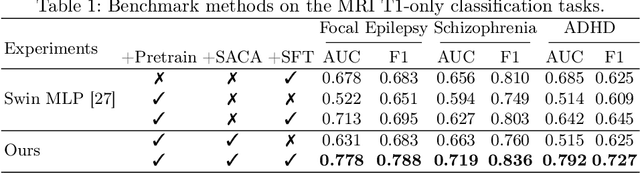


Abstract:The heterogeneity of neurological conditions, ranging from structural anomalies to functional impairments, presents a significant challenge in medical imaging analysis tasks. Moreover, the limited availability of well-annotated datasets constrains the development of robust analysis models. Against this backdrop, this study introduces a novel approach leveraging the inherent anatomical symmetrical features of the human brain to enhance the subsequent detection and segmentation analysis for brain diseases. A novel Symmetry-Aware Cross-Attention (SACA) module is proposed to encode symmetrical features of left and right hemispheres, and a proxy task to detect symmetrical features as the Symmetry-Aware Head (SAH) is proposed, which guides the pretraining of the whole network on a vast 3D brain imaging dataset comprising both healthy and diseased brain images across various MRI and CT. Through meticulous experimentation on downstream tasks, including both classification and segmentation for brain diseases, our model demonstrates superior performance over state-of-the-art methodologies, particularly highlighting the significance of symmetry-aware learning. Our findings advocate for the effectiveness of incorporating symmetry awareness into pretraining and set a new benchmark for medical imaging analysis, promising significant strides toward accurate and efficient diagnostic processes. Code is available at https://github.com/bitMyron/sa-swin.
How Much Data are Enough? Investigating Dataset Requirements for Patch-Based Brain MRI Segmentation Tasks
Apr 04, 2024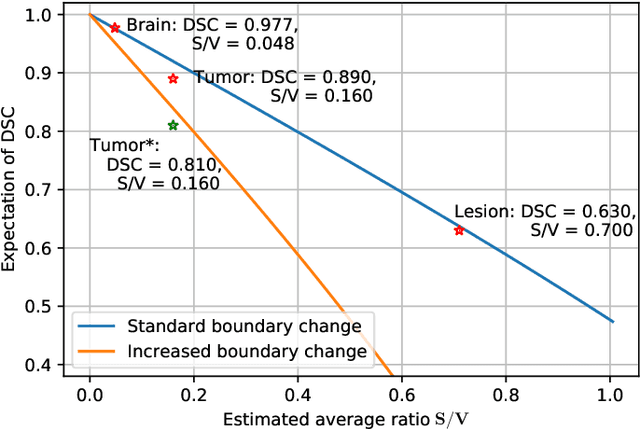
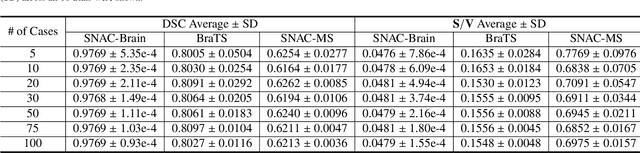

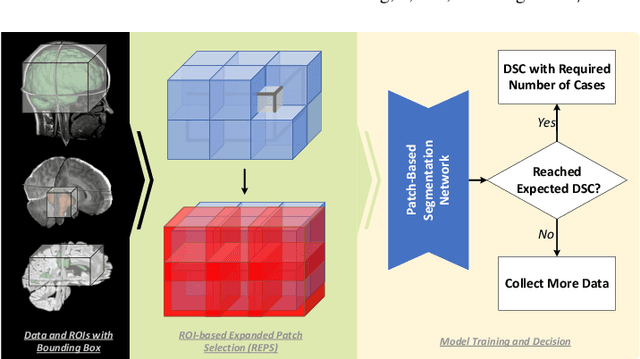
Abstract:Training deep neural networks reliably requires access to large-scale datasets. However, obtaining such datasets can be challenging, especially in the context of neuroimaging analysis tasks, where the cost associated with image acquisition and annotation can be prohibitive. To mitigate both the time and financial costs associated with model development, a clear understanding of the amount of data required to train a satisfactory model is crucial. This paper focuses on an early stage phase of deep learning research, prior to model development, and proposes a strategic framework for estimating the amount of annotated data required to train patch-based segmentation networks. This framework includes the establishment of performance expectations using a novel Minor Boundary Adjustment for Threshold (MinBAT) method, and standardizing patch selection through the ROI-based Expanded Patch Selection (REPS) method. Our experiments demonstrate that tasks involving regions of interest (ROIs) with different sizes or shapes may yield variably acceptable Dice Similarity Coefficient (DSC) scores. By setting an acceptable DSC as the target, the required amount of training data can be estimated and even predicted as data accumulates. This approach could assist researchers and engineers in estimating the cost associated with data collection and annotation when defining a new segmentation task based on deep neural networks, ultimately contributing to their efficient translation to real-world applications.
A Deep Learning Analysis of Climate Change, Innovation, and Uncertainty
Oct 19, 2023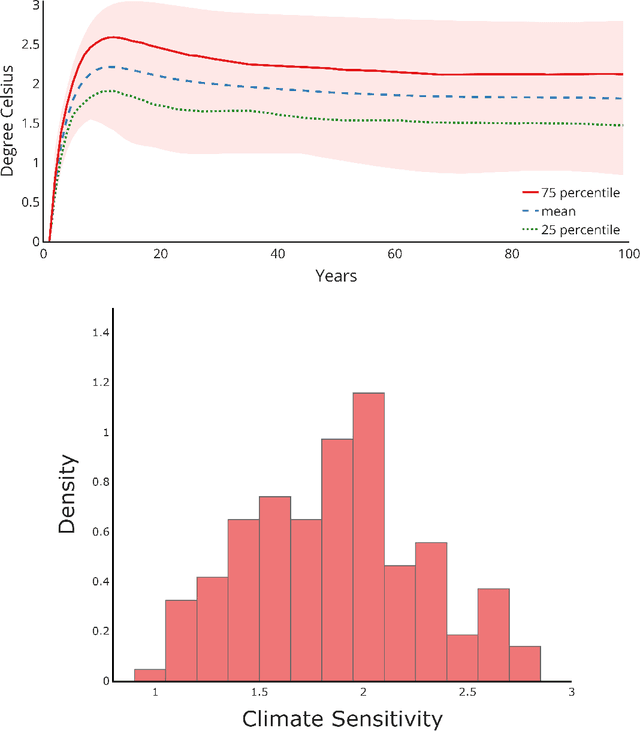
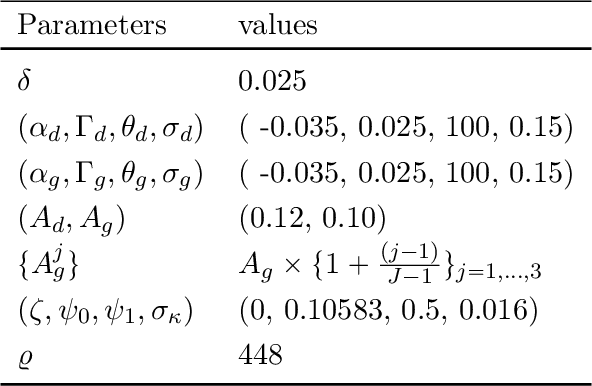

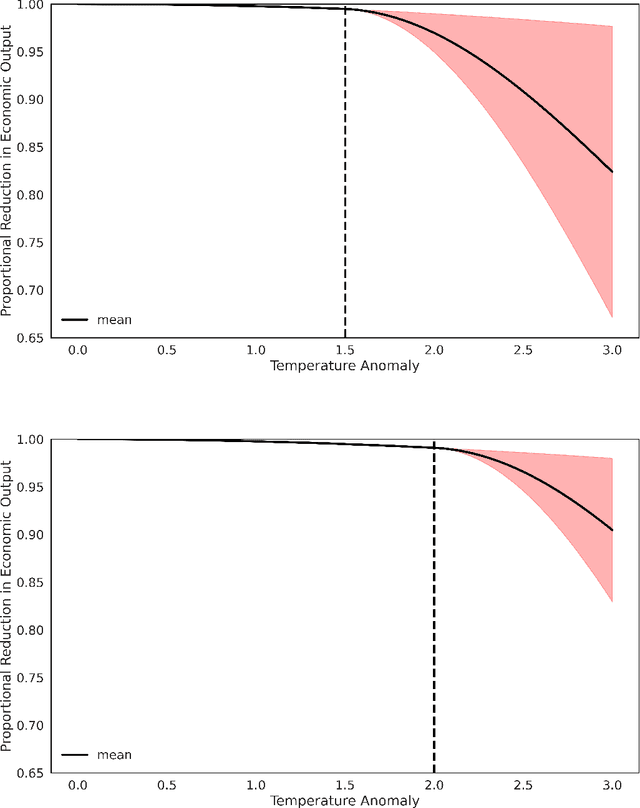
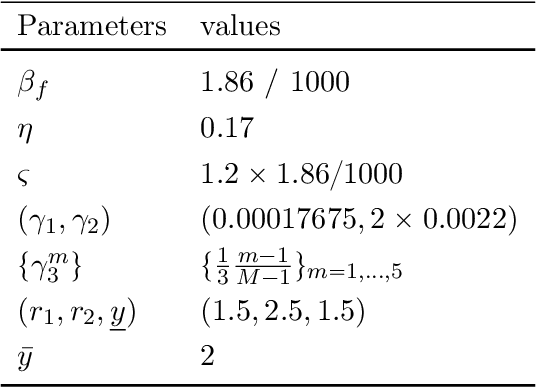
Abstract:We study the implications of model uncertainty in a climate-economics framework with three types of capital: "dirty" capital that produces carbon emissions when used for production, "clean" capital that generates no emissions but is initially less productive than dirty capital, and knowledge capital that increases with R\&D investment and leads to technological innovation in green sector productivity. To solve our high-dimensional, non-linear model framework we implement a neural-network-based global solution method. We show there are first-order impacts of model uncertainty on optimal decisions and social valuations in our integrated climate-economic-innovation framework. Accounting for interconnected uncertainty over climate dynamics, economic damages from climate change, and the arrival of a green technological change leads to substantial adjustments to investment in the different capital types in anticipation of technological change and the revelation of climate damage severity.
Improving Multiple Sclerosis Lesion Segmentation Across Clinical Sites: A Federated Learning Approach with Noise-Resilient Training
Aug 31, 2023



Abstract:Accurately measuring the evolution of Multiple Sclerosis (MS) with magnetic resonance imaging (MRI) critically informs understanding of disease progression and helps to direct therapeutic strategy. Deep learning models have shown promise for automatically segmenting MS lesions, but the scarcity of accurately annotated data hinders progress in this area. Obtaining sufficient data from a single clinical site is challenging and does not address the heterogeneous need for model robustness. Conversely, the collection of data from multiple sites introduces data privacy concerns and potential label noise due to varying annotation standards. To address this dilemma, we explore the use of the federated learning framework while considering label noise. Our approach enables collaboration among multiple clinical sites without compromising data privacy under a federated learning paradigm that incorporates a noise-robust training strategy based on label correction. Specifically, we introduce a Decoupled Hard Label Correction (DHLC) strategy that considers the imbalanced distribution and fuzzy boundaries of MS lesions, enabling the correction of false annotations based on prediction confidence. We also introduce a Centrally Enhanced Label Correction (CELC) strategy, which leverages the aggregated central model as a correction teacher for all sites, enhancing the reliability of the correction process. Extensive experiments conducted on two multi-site datasets demonstrate the effectiveness and robustness of our proposed methods, indicating their potential for clinical applications in multi-site collaborations.
Precise Few-shot Fat-free Thigh Muscle Segmentation in T1-weighted MRI
Apr 27, 2023Abstract:Precise thigh muscle volumes are crucial to monitor the motor functionality of patients with diseases that may result in various degrees of thigh muscle loss. T1-weighted MRI is the default surrogate to obtain thigh muscle masks due to its contrast between muscle and fat signals. Deep learning approaches have recently been widely used to obtain these masks through segmentation. However, due to the insufficient amount of precise annotations, thigh muscle masks generated by deep learning approaches tend to misclassify intra-muscular fat (IMF) as muscle impacting the analysis of muscle volumetrics. As IMF is infiltrated inside the muscle, human annotations require expertise and time. Thus, precise muscle masks where IMF is excluded are limited in practice. To alleviate this, we propose a few-shot segmentation framework to generate thigh muscle masks excluding IMF. In our framework, we design a novel pseudo-label correction and evaluation scheme, together with a new noise robust loss for exploiting high certainty areas. The proposed framework only takes $1\%$ of the fine-annotated training dataset, and achieves comparable performance with fully supervised methods according to the experimental results.
TW-BAG: Tensor-wise Brain-aware Gate Network for Inpainting Disrupted Diffusion Tensor Imaging
Oct 31, 2022Abstract:Diffusion Weighted Imaging (DWI) is an advanced imaging technique commonly used in neuroscience and neurological clinical research through a Diffusion Tensor Imaging (DTI) model. Volumetric scalar metrics including fractional anisotropy, mean diffusivity, and axial diffusivity can be derived from the DTI model to summarise water diffusivity and other quantitative microstructural information for clinical studies. However, clinical practice constraints can lead to sub-optimal DWI acquisitions with missing slices (either due to a limited field of view or the acquisition of disrupted slices). To avoid discarding valuable subjects for group-wise studies, we propose a novel 3D Tensor-Wise Brain-Aware Gate network (TW-BAG) for inpainting disrupted DTIs. The proposed method is tailored to the problem with a dynamic gate mechanism and independent tensor-wise decoders. We evaluated the proposed method on the publicly available Human Connectome Project (HCP) dataset using common image similarity metrics derived from the predicted tensors and scalar DTI metrics. Our experimental results show that the proposed approach can reconstruct the original brain DTI volume and recover relevant clinical imaging information.
MS Lesion Segmentation: Revisiting Weighting Mechanisms for Federated Learning
May 03, 2022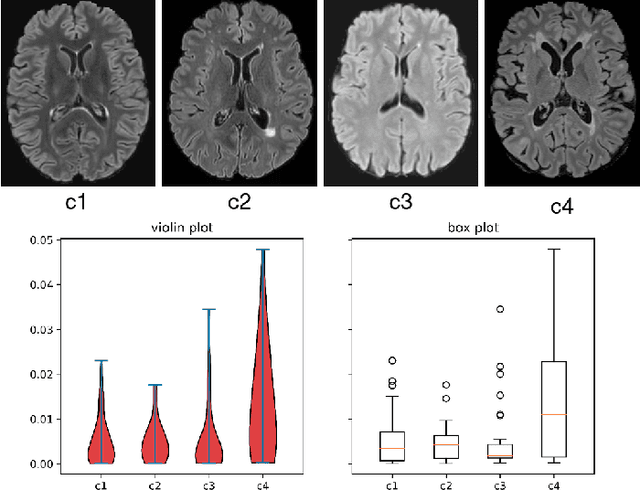
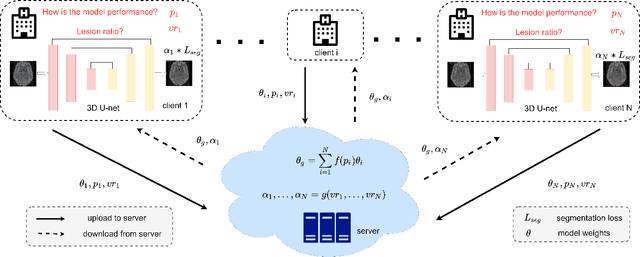
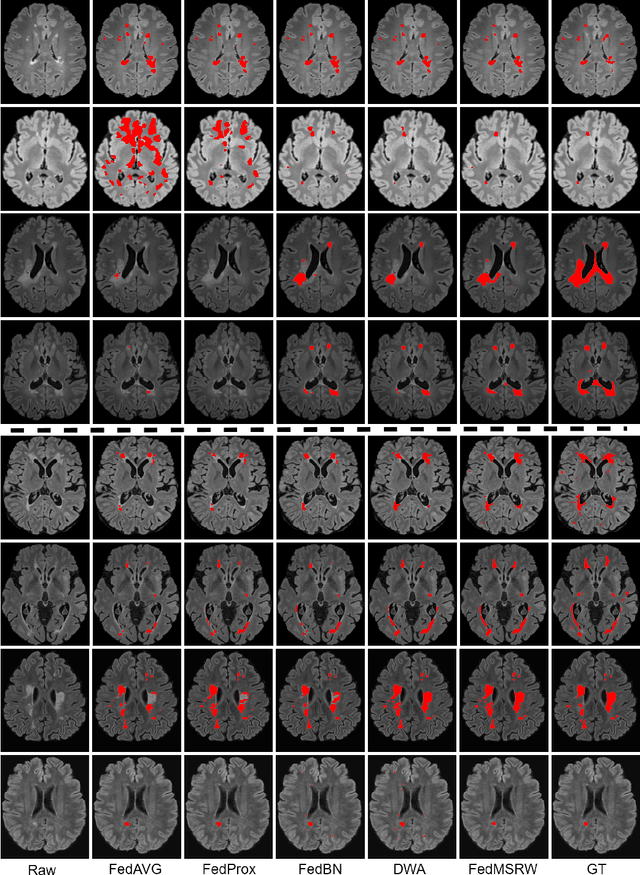
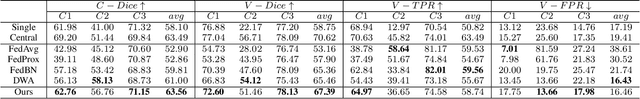
Abstract:Federated learning (FL) has been widely employed for medical image analysis to facilitate multi-client collaborative learning without sharing raw data. Despite great success, FL's performance is limited for multiple sclerosis (MS) lesion segmentation tasks, due to variance in lesion characteristics imparted by different scanners and acquisition parameters. In this work, we propose the first FL MS lesion segmentation framework via two effective re-weighting mechanisms. Specifically, a learnable weight is assigned to each local node during the aggregation process, based on its segmentation performance. In addition, the segmentation loss function in each client is also re-weighted according to the lesion volume for the data during training. Comparison experiments on two FL MS segmentation scenarios using public and clinical datasets have demonstrated the effectiveness of the proposed method by outperforming other FL methods significantly. Furthermore, the segmentation performance of FL incorporating our proposed aggregation mechanism can exceed centralised training with all the raw data. The extensive evaluation also indicated the superiority of our method when estimating brain volume differences estimation after lesion inpainting.
Decompose to Adapt: Cross-domain Object Detection via Feature Disentanglement
Jan 06, 2022

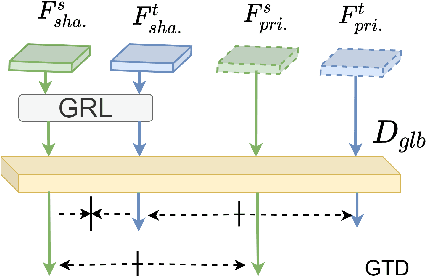

Abstract:Recent advances in unsupervised domain adaptation (UDA) techniques have witnessed great success in cross-domain computer vision tasks, enhancing the generalization ability of data-driven deep learning architectures by bridging the domain distribution gaps. For the UDA-based cross-domain object detection methods, the majority of them alleviate the domain bias by inducing the domain-invariant feature generation via adversarial learning strategy. However, their domain discriminators have limited classification ability due to the unstable adversarial training process. Therefore, the extracted features induced by them cannot be perfectly domain-invariant and still contain domain-private factors, bringing obstacles to further alleviate the cross-domain discrepancy. To tackle this issue, we design a Domain Disentanglement Faster-RCNN (DDF) to eliminate the source-specific information in the features for detection task learning. Our DDF method facilitates the feature disentanglement at the global and local stages, with a Global Triplet Disentanglement (GTD) module and an Instance Similarity Disentanglement (ISD) module, respectively. By outperforming state-of-the-art methods on four benchmark UDA object detection tasks, our DDF method is demonstrated to be effective with wide applicability.
Multiple Sclerosis Lesion Analysis in Brain Magnetic Resonance Images: Techniques and Clinical Applications
Apr 20, 2021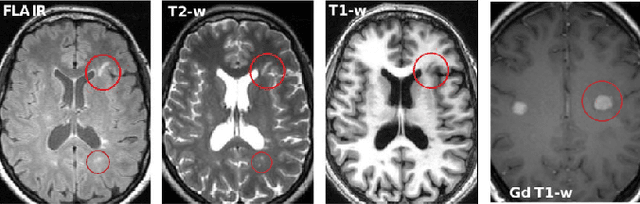
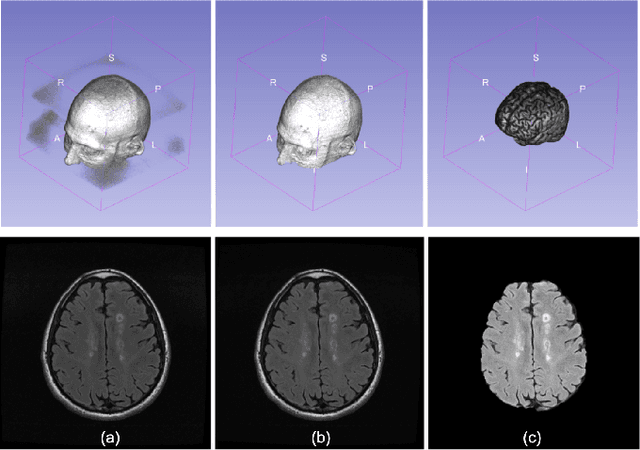
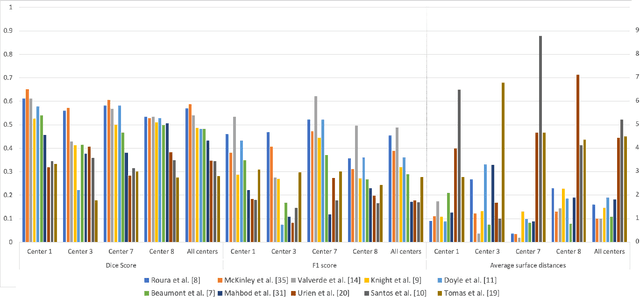
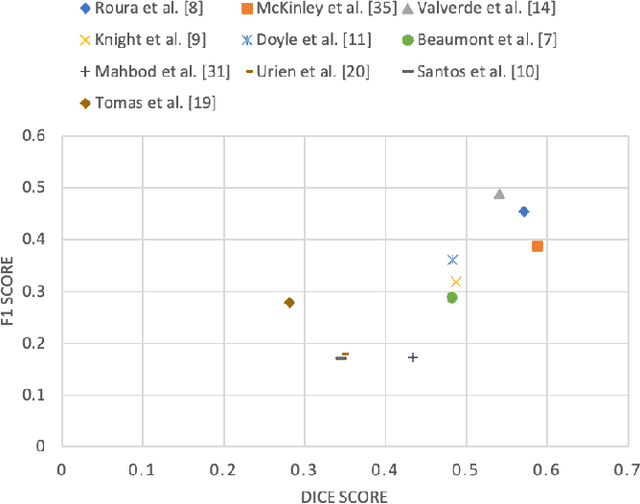
Abstract:Multiple sclerosis (MS) is a chronic inflammatory and degenerative disease of the central nervous system, characterized by the appearance of focal lesions in the white and gray matter that topographically correlate with an individual patient's neurological symptoms and signs. Magnetic resonance imaging (MRI) provides detailed in-vivo structural information, permitting the quantification and categorization of MS lesions that critically inform disease management. Traditionally, MS lesions have been manually annotated on 2D MRI slices, a process that is inefficient and prone to inter-/intra-observer errors. Recently, automated statistical imaging analysis techniques have been proposed to extract and segment MS lesions based on MRI voxel intensity. However, their effectiveness is limited by the heterogeneity of both MRI data acquisition techniques and the appearance of MS lesions. By learning complex lesion representations directly from images, deep learning techniques have achieved remarkable breakthroughs in the MS lesion segmentation task. Here, we provide a comprehensive review of state-of-the-art automatic statistical and deep-learning MS segmentation methods and discuss current and future clinical applications. Further, we review technical strategies, such as domain adaptation, to enhance MS lesion segmentation in real-world clinical settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge