Yirong Zhou
Error Bound Analysis of Physics-Informed Neural Networks-Driven T2 Quantification in Cardiac Magnetic Resonance Imaging
Dec 16, 2025



Abstract:Physics-Informed Neural Networks (PINN) are emerging as a promising approach for quantitative parameter estimation of Magnetic Resonance Imaging (MRI). While existing deep learning methods can provide an accurate quantitative estimation of the T2 parameter, they still require large amounts of training data and lack theoretical support and a recognized gold standard. Thus, given the absence of PINN-based approaches for T2 estimation, we propose embedding the fundamental physics of MRI, the Bloch equation, in the loss of PINN, which is solely based on target scan data and does not require a pre-defined training database. Furthermore, by deriving rigorous upper bounds for both the T2 estimation error and the generalization error of the Bloch equation solution, we establish a theoretical foundation for evaluating the PINN's quantitative accuracy. Even without access to the ground truth or a gold standard, this theory enables us to estimate the error with respect to the real quantitative parameter T2. The accuracy of T2 mapping and the validity of the theoretical analysis are demonstrated on a numerical cardiac model and a water phantom, where our method exhibits excellent quantitative precision in the myocardial T2 range. Clinical applicability is confirmed in 94 acute myocardial infarction (AMI) patients, achieving low-error quantitative T2 estimation under the theoretical error bound, highlighting the robustness and potential of PINN.
Simultaneous Deep Learning of Myocardium Segmentation and T2 Quantification for Acute Myocardial Infarction MRI
May 17, 2024



Abstract:In cardiac Magnetic Resonance Imaging (MRI) analysis, simultaneous myocardial segmentation and T2 quantification are crucial for assessing myocardial pathologies. Existing methods often address these tasks separately, limiting their synergistic potential. To address this, we propose SQNet, a dual-task network integrating Transformer and Convolutional Neural Network (CNN) components. SQNet features a T2-refine fusion decoder for quantitative analysis, leveraging global features from the Transformer, and a segmentation decoder with multiple local region supervision for enhanced accuracy. A tight coupling module aligns and fuses CNN and Transformer branch features, enabling SQNet to focus on myocardium regions. Evaluation on healthy controls (HC) and acute myocardial infarction patients (AMI) demonstrates superior segmentation dice scores (89.3/89.2) compared to state-of-the-art methods (87.7/87.9). T2 quantification yields strong linear correlations (Pearson coefficients: 0.84/0.93) with label values for HC/AMI, indicating accurate mapping. Radiologist evaluations confirm SQNet's superior image quality scores (4.60/4.58 for segmentation, 4.32/4.42 for T2 quantification) over state-of-the-art methods (4.50/4.44 for segmentation, 3.59/4.37 for T2 quantification). SQNet thus offers accurate simultaneous segmentation and quantification, enhancing cardiac disease diagnosis, such as AMI.
Deep Separable Spatiotemporal Learning for Fast Dynamic Cardiac MRI
Feb 24, 2024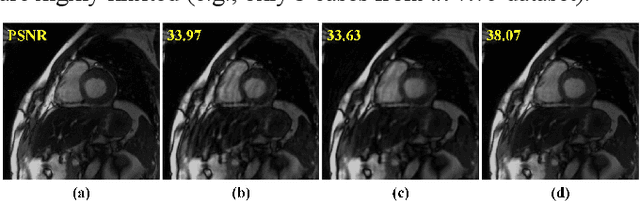
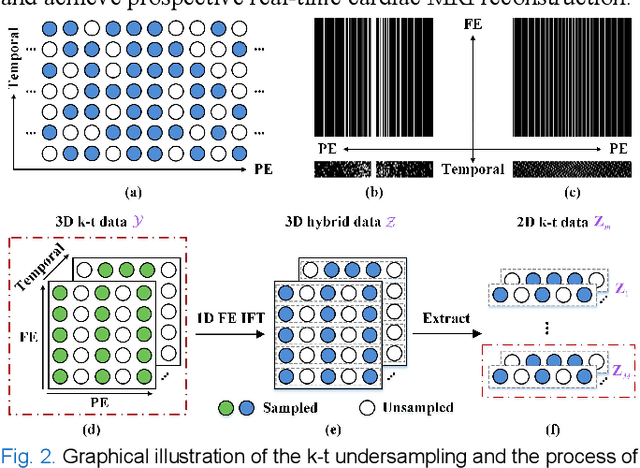
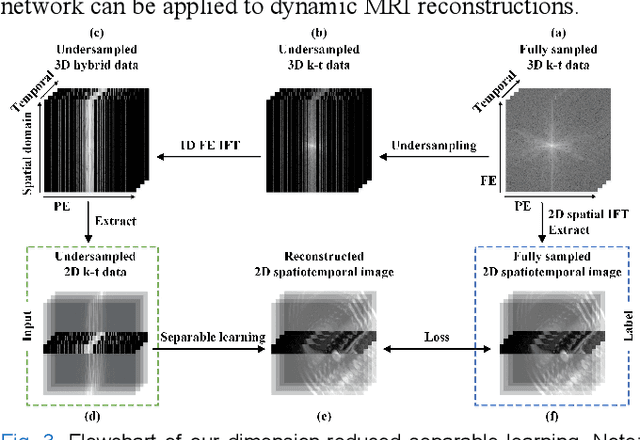
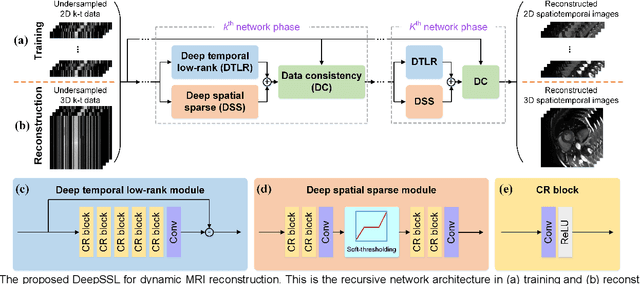
Abstract:Dynamic magnetic resonance imaging (MRI) plays an indispensable role in cardiac diagnosis. To enable fast imaging, the k-space data can be undersampled but the image reconstruction poses a great challenge of high-dimensional processing. This challenge leads to necessitate extensive training data in many deep learning reconstruction methods. This work proposes a novel and efficient approach, leveraging a dimension-reduced separable learning scheme that excels even with highly limited training data. We further integrate it with spatiotemporal priors to develop a Deep Separable Spatiotemporal Learning network (DeepSSL), which unrolls an iteration process of a reconstruction model with both temporal low-rankness and spatial sparsity. Intermediate outputs are visualized to provide insights into the network's behavior and enhance its interpretability. Extensive results on cardiac cine datasets show that the proposed DeepSSL is superior to the state-of-the-art methods visually and quantitatively, while reducing the demand for training cases by up to 75%. And its preliminary adaptability to cardiac patients has been verified through experienced radiologists' and cardiologists' blind reader study. Additionally, DeepSSL also benefits for achieving the downstream task of cardiac segmentation with higher accuracy and shows robustness in prospective real-time cardiac MRI.
Cloud-Magnetic Resonance Imaging System: In the Era of 6G and Artificial Intelligence
Oct 18, 2023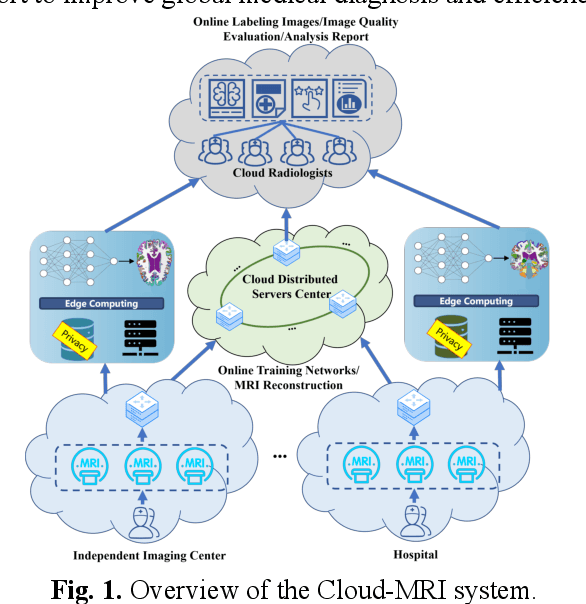
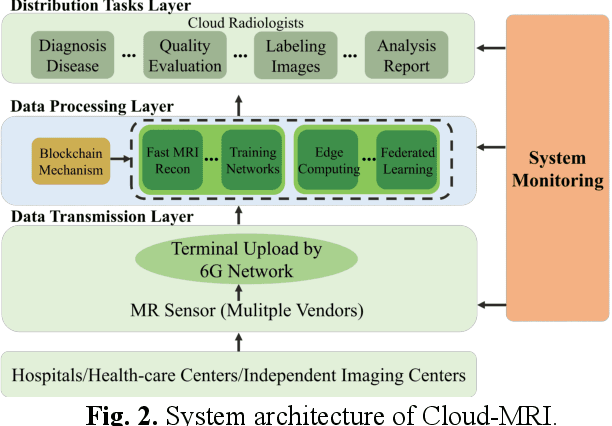
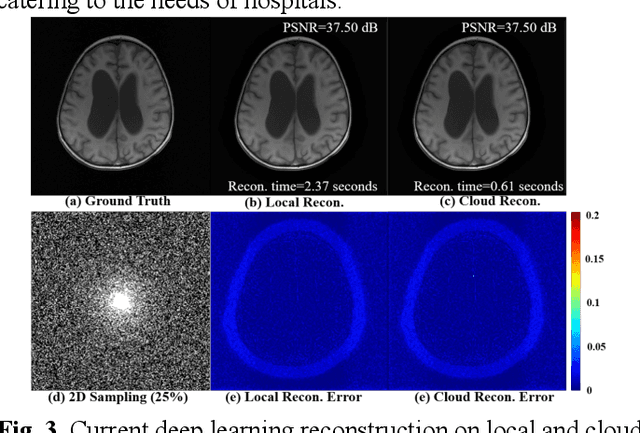
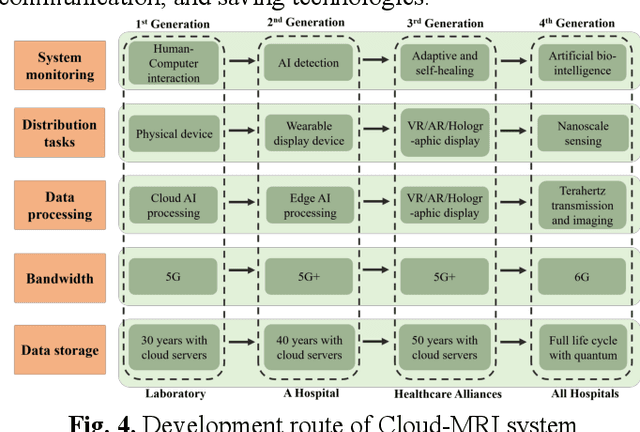
Abstract:Magnetic Resonance Imaging (MRI) plays an important role in medical diagnosis, generating petabytes of image data annually in large hospitals. This voluminous data stream requires a significant amount of network bandwidth and extensive storage infrastructure. Additionally, local data processing demands substantial manpower and hardware investments. Data isolation across different healthcare institutions hinders cross-institutional collaboration in clinics and research. In this work, we anticipate an innovative MRI system and its four generations that integrate emerging distributed cloud computing, 6G bandwidth, edge computing, federated learning, and blockchain technology. This system is called Cloud-MRI, aiming at solving the problems of MRI data storage security, transmission speed, AI algorithm maintenance, hardware upgrading, and collaborative work. The workflow commences with the transformation of k-space raw data into the standardized Imaging Society for Magnetic Resonance in Medicine Raw Data (ISMRMRD) format. Then, the data are uploaded to the cloud or edge nodes for fast image reconstruction, neural network training, and automatic analysis. Then, the outcomes are seamlessly transmitted to clinics or research institutes for diagnosis and other services. The Cloud-MRI system will save the raw imaging data, reduce the risk of data loss, facilitate inter-institutional medical collaboration, and finally improve diagnostic accuracy and work efficiency.
CloudBrain-MRS: An Intelligent Cloud Computing Platform for in vivo Magnetic Resonance Spectroscopy Preprocessing, Quantification, and Analysis
Jun 19, 2023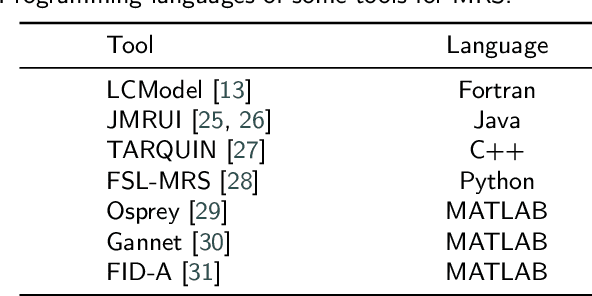
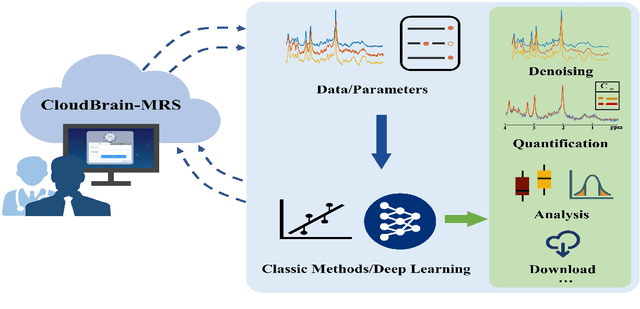
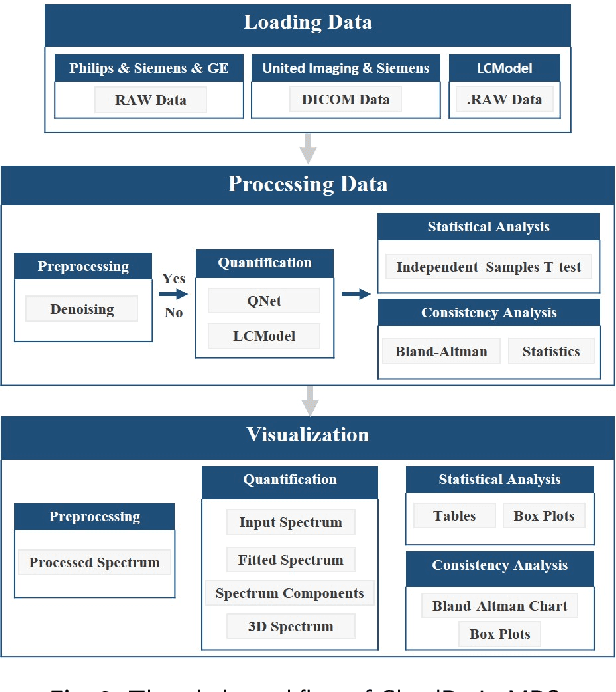
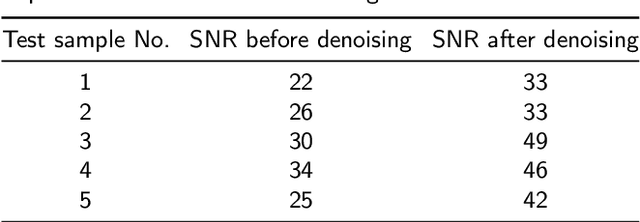
Abstract:Magnetic resonance spectroscopy (MRS) is an important clinical imaging method for the diagnosis of diseases. Spectrum is used to observe the signal intensity of metabolites or further infer their concentrations. Although the magnetic resonance vendors commonly provide basic functions of spectra plots and metabolite quantification, the widespread clinical research of MRS is still limited due to the lack of easy-to-use processing software or platform. To address this issue, we have developed CloudBrain-MRS, a cloud-based online platform that provides powerful hardware and advanced algorithms. The platform can be accessed simply through a web browser, without requiring any program installation on the user side. CloudBrain-MRS also integrates the classic LCModel and advanced artificial intelligence algorithms and supports batch preprocessing, quantification, and analysis of MRS data. Additionally, the platform offers useful functions: 1) Automatically statistical analysis to find biomarkers from the health and patient groups; 2) Consistency verification between the classic and artificial intelligence quantification algorithms; 3) Colorful and three-dimensional visualization for the easy observation of individual metabolite spectrum. Last, both healthy and mild cognitive impairment patient data are used to demonstrate the usefulness of the platform. To the best of our knowledge, this is the first cloud computing platform for in vivo MRS with artificial intelligence processing. We sincerely hope that this platform will facilitate efficient clinical research for MRS. CloudBrain-MRS is open-accessed at https://csrc.xmu.edu.cn/CloudBrain.html.
Magnetic Resonance Spectroscopy Quantification Aided by Deep Estimations of Imperfection Factors and Overall Macromolecular Signal
Jun 16, 2023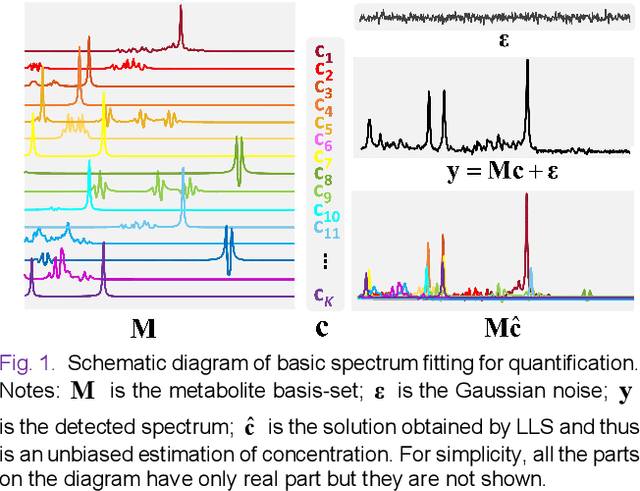
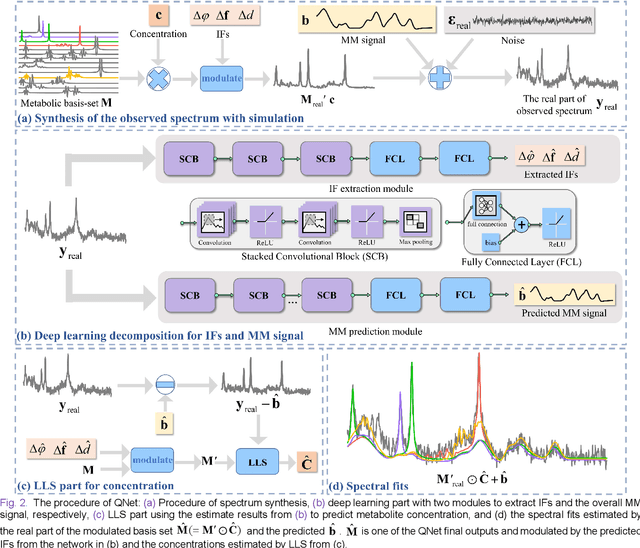
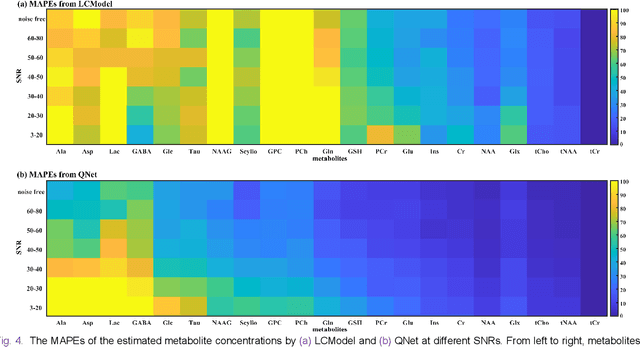
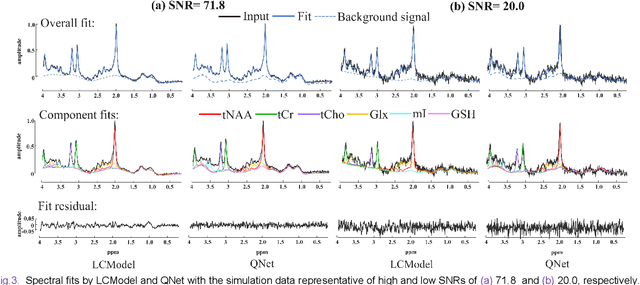
Abstract:Magnetic Resonance Spectroscopy (MRS) is an important non-invasive technique for in vivo biomedical detection. However, it is still challenging to accurately quantify metabolites with proton MRS due to three problems: Serious overlaps of metabolite signals, signal distortions due to non-ideal acquisition conditions and interference with strong background signals including macromolecule signals. The most popular software, LCModel, adopts the non-linear least square to quantify metabolites and addresses these problems by introducing regularization terms, imperfection factors of non-ideal acquisition conditions, and designing several empirical priors such as basissets of both metabolites and macromolecules. However, solving such a large non-linear quantitative problem is complicated. Moreover, when the signal-to-noise ratio of an input MRS signal is low, the solution may have a large deviation. In this work, deep learning is introduced to reduce the complexity of solving this overall quantitative problem. Deep learning is designed to predict directly the imperfection factors and the overall signal from macromolecules. Then, the remaining part of the quantification problem becomes a much simpler effective fitting and is easily solved by Linear Least Squares (LLS), which greatly improves the generalization to unseen concentration of metabolites in the training data. Experimental results show that compared with LCModel, the proposed method has smaller quantification errors for 700 sets of simulated test data, and presents more stable quantification results for 20 sets of healthy in vivo data at a wide range of signal-to-noise ratio. Qnet also outperforms other deep learning methods in terms of lower quantification error on most metabolites. Finally, QNet has been deployed on a cloud computing platform, CloudBrain-MRS, which is open accessed at https://csrc.xmu.edu.cn/CloudBrain.html.
CloudBrain-ReconAI: An Online Platform for MRI Reconstruction and Image Quality Evaluation
Dec 04, 2022



Abstract:Efficient collaboration between engineers and radiologists is important for image reconstruction algorithm development and image quality evaluation in magnetic resonance imaging (MRI). Here, we develop CloudBrain-ReconAI, an online cloud computing platform, for algorithm deployment, fast and blind reader study. This platform supports online image reconstruction using state-of-the-art artificial intelligence and compressed sensing algorithms with applications to fast imaging and high-resolution diffusion imaging. Through visiting the website, radiologists can easily score and mark the images. Then, automatic statistical analysis will be provided. CloudBrain-ReconAI is now open accessed at https://csrc.xmu.edu.cn/CloudBrain.html and will be continually improved to serve the MRI research community.
XCloud-pFISTA: A Medical Intelligence Cloud for Accelerated MRI
Apr 18, 2021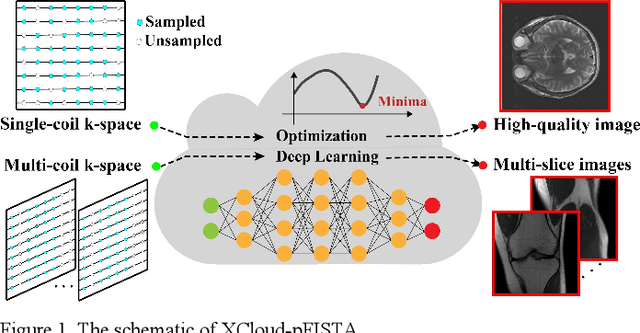
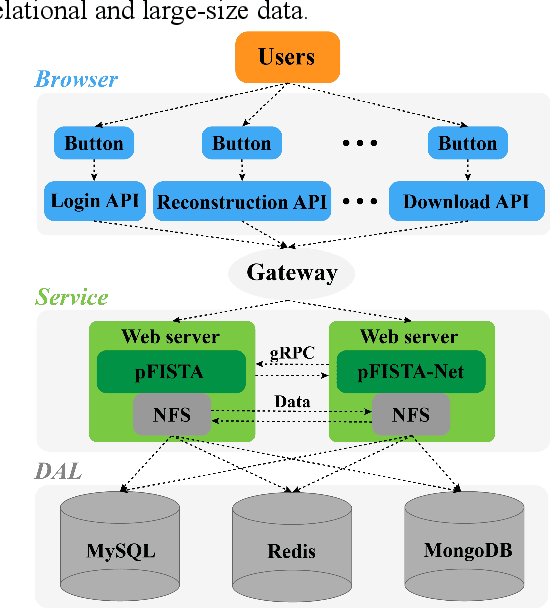
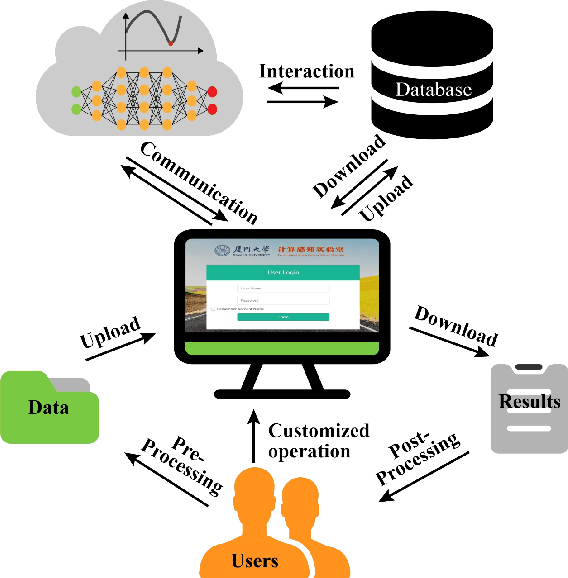
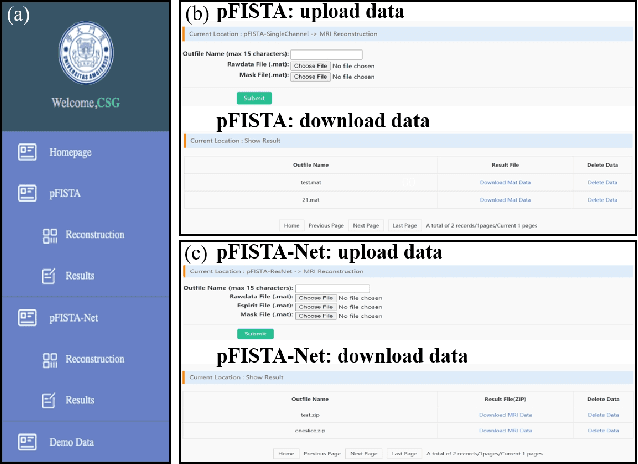
Abstract:Machine learning and artificial intelligence have shown remarkable performance in accelerated magnetic resonance imaging (MRI). Cloud computing technologies have great advantages in building an easily accessible platform to deploy advanced algorithms. In this work, we develop an open-access, easy-to-use and high-performance medical intelligence cloud computing platform (XCloud-pFISTA) to reconstruct MRI images from undersampled k-space data. Two state-of-the-art approaches of the Projected Fast Iterative Soft-Thresholding Algorithm (pFISTA) family have been successfully implemented on the cloud. This work can be considered as a good example of cloud-based medical image reconstruction and may benefit the future development of integrated reconstruction and online diagnosis system.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge