Kawin Setsompop
Fully 3D Unrolled Magnetic Resonance Fingerprinting Reconstruction via Staged Pretraining and Implicit Gridding
Jan 23, 2026Abstract:Magnetic Resonance Fingerprinting (MRF) enables fast quantitative imaging, yet reconstructing high-resolution 3D data remains computationally demanding. Non-Cartesian reconstructions require repeated non-uniform FFTs, and the commonly used Locally Low Rank (LLR) prior adds computational overhead and becomes insufficient at high accelerations. Learned 3D priors could address these limitations, but training them at scale is challenging due to memory and runtime demands. We propose SPUR-iG, a fully 3D deep unrolled subspace reconstruction framework that integrates efficient data consistency with a progressive training strategy. Data consistency leverages implicit GROG, which grids non-Cartesian data onto a Cartesian grid with an implicitly learned kernel, enabling FFT-based updates with minimal artifacts. Training proceeds in three stages: (1) pretraining a denoiser with extensive data augmentation, (2) greedy per-iteration unrolled training, and (3) final fine-tuning with gradient checkpointing. Together, these stages make large-scale 3D unrolled learning feasible within a reasonable compute budget. On a large in vivo dataset with retrospective undersampling, SPUR-iG improves subspace coefficient maps quality and quantitative accuracy at 1-mm isotropic resolution compared with LLR and a hybrid 2D/3D unrolled baseline. Whole-brain reconstructions complete in under 15-seconds, with up to $\times$111 speedup for 2-minute acquisitions. Notably, $T_1$ maps with our method from 30-second scans achieve accuracy on par with or exceeding LLR reconstructions from 2-minute scans. Overall, the framework improves both accuracy and speed in large-scale 3D MRF reconstruction, enabling efficient and reliable accelerated quantitative imaging.
Fast Reconstruction of Motion-Corrupted Data with Mobile-GRAPPA: Motion and dB0 Inhomogeneity Correction Leveraging Efficient GRAPPA
Nov 09, 2025Abstract:Advanced motion navigations now enable rapid tracking of subject motion and dB0-induced phase, but accurately incorporating this high-temporal-resolution information into SENSE (Aligned-SENSE) is often computationally prohibitive. We propose "Mobile-GRAPPA", a k-space "cleaning" approach that uses local GRAPPA operators to remove motion and dB0 related corruption so that the resulting data can be reconstructed with standard SENSE. We efficiently train a family of k-space-position-specific Mobile-GRAPPA kernels via a lightweight multilayer perceptron (MLP) and apply them across k-space to generate clean data. In experiments on highly motion-corrupted 1-mm whole-brain GRE (Tacq = 10 min; 1,620 motion/dB0 trackings) and EPTI (Tacq = 2 min; 544 trackings), Mobile-GRAPPA enabled accurate reconstruction with negligible time penalty, whereas full Aligned-SENSE was impractical (reconstruction times > 10 h for GRE and > 10 days for EPTI). These results show that Mobile-GRAPPA incorporates detailed motion and dB0 tracking into SENSE with minimal computational overhead, enabling fast, high-quality reconstructions of challenging data.
Enhancing Diffusion-Weighted Images (DWI) for Diffusion MRI: Is it Enough without Non-Diffusion-Weighted B=0 Reference?
May 19, 2025Abstract:Diffusion MRI (dMRI) is essential for studying brain microstructure, but high-resolution imaging remains challenging due to the inherent trade-offs between acquisition time and signal-to-noise ratio (SNR). Conventional methods often optimize only the diffusion-weighted images (DWIs) without considering their relationship with the non-diffusion-weighted (b=0) reference images. However, calculating diffusion metrics, such as the apparent diffusion coefficient (ADC) and diffusion tensor with its derived metrics like fractional anisotropy (FA) and mean diffusivity (MD), relies on the ratio between each DWI and the b=0 image, which is crucial for clinical observation and diagnostics. In this study, we demonstrate that solely enhancing DWIs using a conventional pixel-wise mean squared error (MSE) loss is insufficient, as the error in ratio between generated DWIs and b=0 diverges. We propose a novel ratio loss, defined as the MSE loss between the predicted and ground-truth log of DWI/b=0 ratios. Our results show that incorporating the ratio loss significantly improves the convergence of this ratio error, achieving lower ratio MSE and slightly enhancing the peak signal-to-noise ratio (PSNR) of generated DWIs. This leads to improved dMRI super-resolution and better preservation of b=0 ratio-based features for the derivation of diffusion metrics.
OPTIKS: Optimized Gradient Properties Through Timing in K-Space
May 11, 2025



Abstract:A customizable method (OPTIKS) for designing fast trajectory-constrained gradient waveforms with optimized time domain properties was developed. Given a specified multidimensional k-space trajectory, the method optimizes traversal speed (and therefore timing) with position along the trajectory. OPTIKS facilitates optimization of objectives dependent on the time domain gradient waveform and the arc-length domain k-space speed. OPTIKS is applied to design waveforms which limit peripheral nerve stimulation (PNS), minimize mechanical resonance excitation, and reduce acoustic noise. A variety of trajectory examples are presented including spirals, circular echo-planar-imaging, and rosettes. Design performance is evaluated based on duration, standardized PNS models, field measurements, gradient coil back-EMF measurements, and calibrated acoustic measurements. We show reductions in back-EMF of up to 94% and field oscillations up to 91.1%, acoustic noise decreases of up to 9.22 dB, and with efficient use of PNS models speed increases of up to 11.4%. The design method implementation is made available as an open source Python package through GitHub.
ACE-Net: AutofoCus-Enhanced Convolutional Network for Field Imperfection Estimation with application to high b-value spiral Diffusion MRI
Nov 21, 2024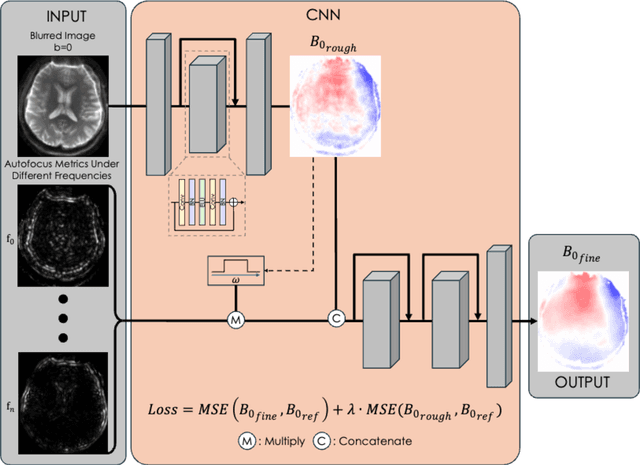
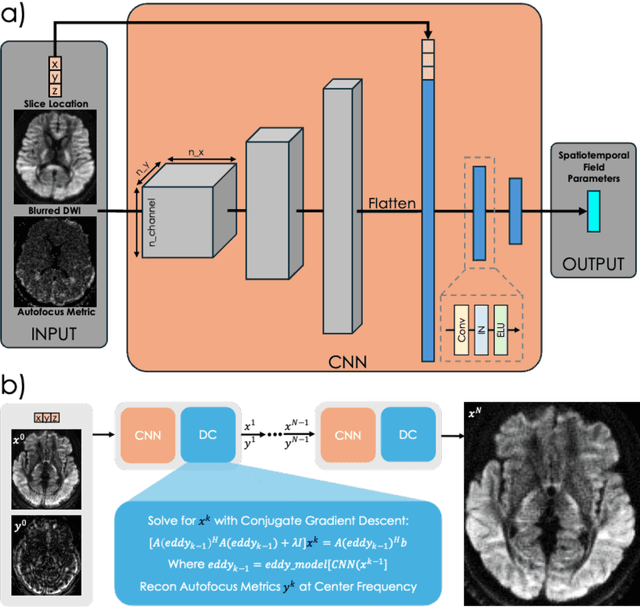
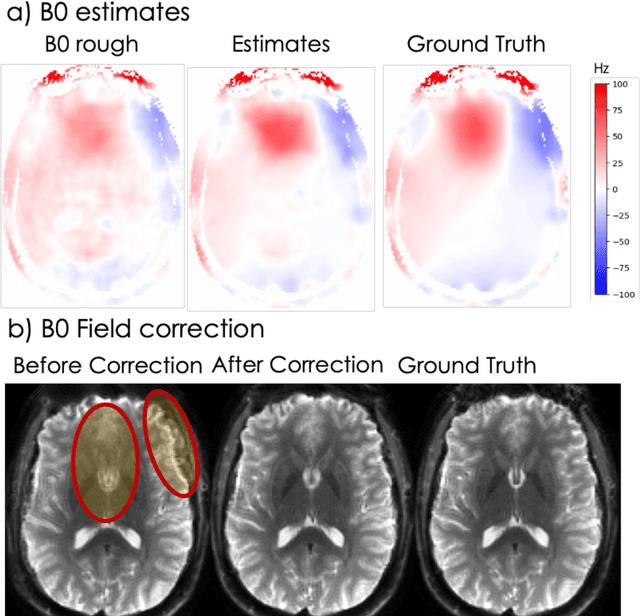
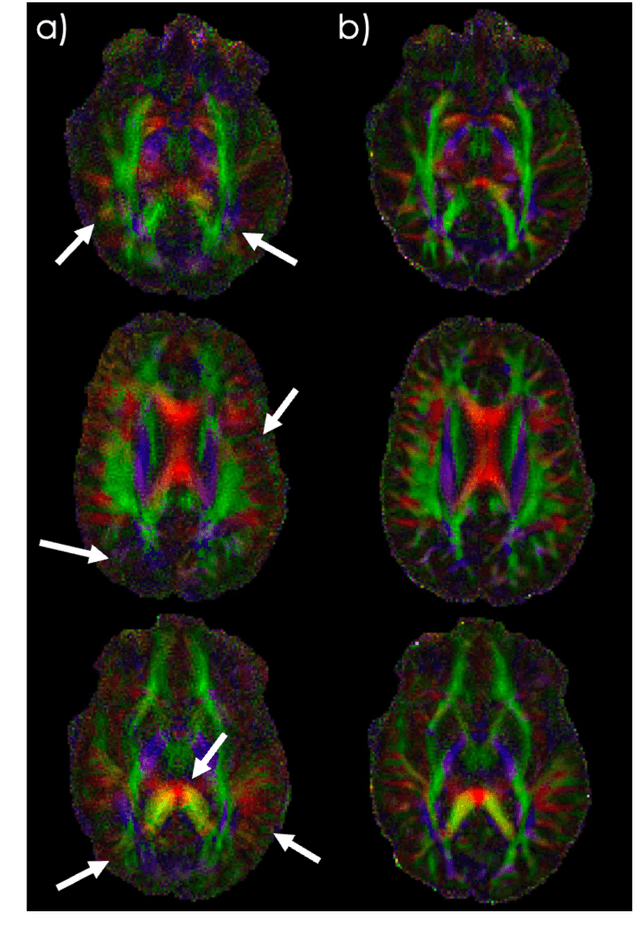
Abstract:Spatiotemporal magnetic field variations from B0-inhomogeneity and diffusion-encoding-induced eddy-currents can be detrimental to rapid image-encoding schemes such as spiral, EPI and 3D-cones, resulting in undesirable image artifacts. In this work, a data driven approach for automatic estimation of these field imperfections is developed by combining autofocus metrics with deep learning, and by leveraging a compact basis representation of the expected field imperfections. The method was applied to single-shot spiral diffusion MRI at high b-values where accurate estimation of B0 and eddy were obtained, resulting in high quality image reconstruction without need for additional external calibrations.
Accelerating Longitudinal MRI using Prior Informed Latent Diffusion
Jun 29, 2024



Abstract:MRI is a widely used ionization-free soft-tissue imaging modality, often employed repeatedly over a patient's lifetime. However, prolonged scanning durations, among other issues, can limit availability and accessibility. In this work, we aim to substantially reduce scan times by leveraging prior scans of the same patient. These prior scans typically contain considerable shared information with the current scan, thereby enabling higher acceleration rates when appropriately utilized. We propose a prior informed reconstruction method with a trained diffusion model in conjunction with data-consistency steps. Our method can be trained with unlabeled image data, eliminating the need for a dataset of either k-space measurements or paired longitudinal scans as is required of other learning-based methods. We demonstrate superiority of our method over previously suggested approaches in effectively utilizing prior information without over-biasing prior consistency, which we validate on both an open-source dataset of healthy patients as well as several longitudinal cases of clinical interest.
Data and Physics driven Deep Learning Models for Fast MRI Reconstruction: Fundamentals and Methodologies
Jan 29, 2024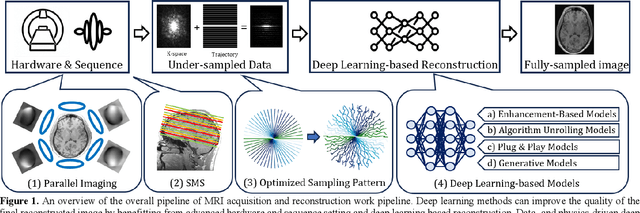


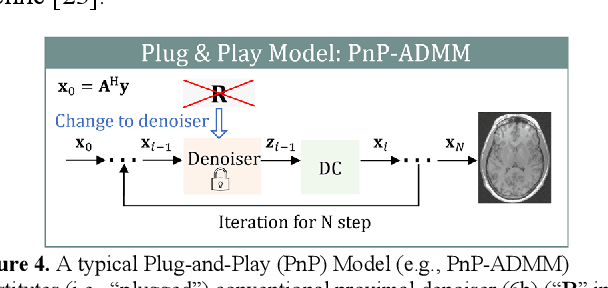
Abstract:Magnetic Resonance Imaging (MRI) is a pivotal clinical diagnostic tool, yet its extended scanning times often compromise patient comfort and image quality, especially in volumetric, temporal and quantitative scans. This review elucidates recent advances in MRI acceleration via data and physics-driven models, leveraging techniques from algorithm unrolling models, enhancement-based models, and plug-and-play models to emergent full spectrum of generative models. We also explore the synergistic integration of data models with physics-based insights, encompassing the advancements in multi-coil hardware accelerations like parallel imaging and simultaneous multi-slice imaging, and the optimization of sampling patterns. We then focus on domain-specific challenges and opportunities, including image redundancy exploitation, image integrity, evaluation metrics, data heterogeneity, and model generalization. This work also discusses potential solutions and future research directions, emphasizing the role of data harmonization, and federated learning for further improving the general applicability and performance of these methods in MRI reconstruction.
An Efficient Algorithm for Spatial-Spectral Partial Volume Compartment Mapping with Applications to Multicomponent Diffusion and Relaxation MRI
Jan 23, 2024



Abstract:It has been previously shown that high-quality partial volume tissue compartment maps can be obtained by combining multiparametric contrast-encoded MRI data acquisition methods with spatially-regularized spectroscopic image estimation techniques. However, the advantages of this combined approach generally come at the expense of substantial computational complexity. In this work, we propose a new algorithm to solve this kind of estimation problem more efficiently. Our algorithm is based on the linearized alternating directions method of multipliers (LADMM), and relies on the introduction of novel quadratic penalty terms to substantially simplify the subproblems that must be solved at each iteration. We evaluate this algorithm on a variety of different estimation problems (diffusion-relaxation, relaxation-relaxation, relaxometry, and magnetic resonance fingerprinting), where we consistently observe substantial (roughly 5$\times$-80$\times$) speed improvements. We expect that this new faster algorithm will lower practical barriers to using spatial regularization and multiparametric contrast-encoded MRI data acquisition methods for partial volume compartment mapping.
High-resolution myelin-water fraction and quantitative relaxation mapping using 3D ViSTa-MR fingerprinting
Dec 21, 2023Abstract:Purpose: This study aims to develop a high-resolution whole-brain multi-parametric quantitative MRI approach for simultaneous mapping of myelin-water fraction (MWF), T1, T2, and proton-density (PD), all within a clinically feasible scan time. Methods: We developed 3D ViSTa-MRF, which combined Visualization of Short Transverse relaxation time component (ViSTa) technique with MR Fingerprinting (MRF), to achieve high-fidelity whole-brain MWF and T1/T2/PD mapping on a clinical 3T scanner. To achieve fast acquisition and memory-efficient reconstruction, the ViSTa-MRF sequence leverages an optimized 3D tiny-golden-angle-shuffling spiral-projection acquisition and joint spatial-temporal subspace reconstruction with optimized preconditioning algorithm. With the proposed ViSTa-MRF approach, high-fidelity direct MWF mapping was achieved without a need for multi-compartment fitting that could introduce bias and/or noise from additional assumptions or priors. Results: The in-vivo results demonstrate the effectiveness of the proposed acquisition and reconstruction framework to provide fast multi-parametric mapping with high SNR and good quality. The in-vivo results of 1mm- and 0.66mm-iso datasets indicate that the MWF values measured by the proposed method are consistent with standard ViSTa results that are 30x slower with lower SNR. Furthermore, we applied the proposed method to enable 5-minute whole-brain 1mm-iso assessment of MWF and T1/T2/PD mappings for infant brain development and for post-mortem brain samples. Conclusions: In this work, we have developed a 3D ViSTa-MRF technique that enables the acquisition of whole-brain MWF, quantitative T1, T2, and PD maps at 1mm and 0.66mm isotropic resolution in 5 and 15 minutes, respectively. This advancement allows for quantitative investigations of myelination changes in the brain.
Sequence adaptive field-imperfection estimation (SAFE): retrospective estimation and correction of $B_1^+$ and $B_0$ inhomogeneities for enhanced MRF quantification
Dec 15, 2023



Abstract:$B_1^+$ and $B_0$ field-inhomogeneities can significantly reduce accuracy and robustness of MRF's quantitative parameter estimates. Additional $B_1^+$ and $B_0$ calibration scans can mitigate this but add scan time and cannot be applied retrospectively to previously collected data. Here, we proposed a calibration-free sequence-adaptive deep-learning framework, to estimate and correct for $B_1^+$ and $B_0$ effects of any MRF sequence. We demonstrate its capability on arbitrary MRF sequences at 3T, where no training data were previously obtained. Such approach can be applied to any previously-acquired and future MRF-scans. The flexibility in directly applying this framework to other quantitative sequences is also highlighted.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge