Fuyixue Wang
Time-efficient, High Resolution 3T Whole Brain Quantitative Relaxometry using 3D-QALAS with Wave-CAIPI Readouts
Nov 08, 2022

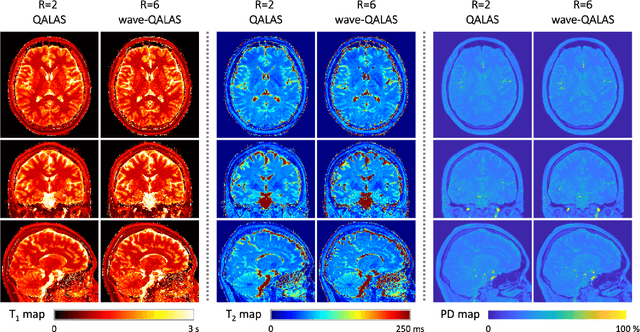

Abstract:Purpose: Volumetric, high resolution, quantitative mapping of brain tissues relaxation properties is hindered by long acquisition times and SNR challenges. This study, for the first time, combines the time efficient wave-CAIPI readouts into the 3D-QALAS acquisition scheme, enabling full brain quantitative T1, T2 and PD maps at 1.15 isotropic voxels in only 3 minutes. Methods: Wave-CAIPI readouts were embedded in the standard 3d-QALAS encoding scheme, enabling full brain quantitative parameter maps (T1, T2 and PD) at acceleration factors of R=3x2 with minimum SNR loss due to g-factor penalties. The quantitative maps using the accelerated protocol were quantitatively compared against those obtained from conventional 3D-QALAS sequence using GRAPPA acceleration of R=2 in the ISMRM NIST phantom, and ten healthy volunteers. To show the feasibility of the proposed methods in clinical settings, the accelerated wave-CAIPI 3D-QALAS sequence was also employed in pediatric patients undergoing clinical MRI examinations. Results: When tested in both the ISMRM/NIST phantom and 7 healthy volunteers, the quantitative maps using the accelerated protocol showed excellent agreement against those obtained from conventional 3D-QALAS at R=2. Conclusion: 3D-QALAS enhanced with wave-CAIPI readouts enables time-efficient, full brain quantitative T1, T2 and PD mapping at 1.15 in 3 minutes at R=3x2 acceleration. When tested on the NIST phantom and 7 healthy volunteers, the quantitative maps obtained from the accelerated wave-CAIPI 3D-QALAS protocol showed very similar values to those obtained from the standard 3D-QALAS (R=2) protocol, alluding to the robustness and reliability of the proposed methods. This study also shows that the accelerated protocol can be effectively employed in pediatric patient populations, making high-quality high-resolution full brain quantitative imaging feasible in clinical settings.
Latent Signal Models: Learning Compact Representations of Signal Evolution for Improved Time-Resolved, Multi-contrast MRI
Aug 27, 2022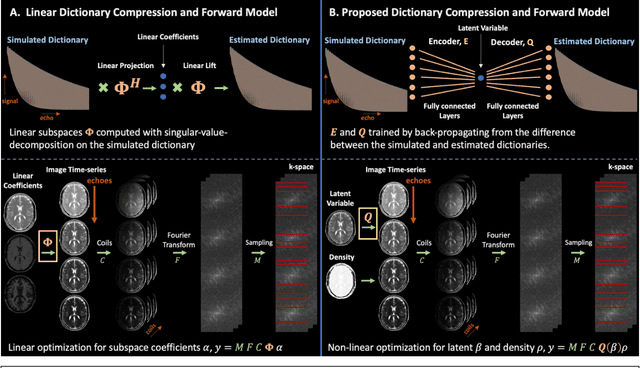
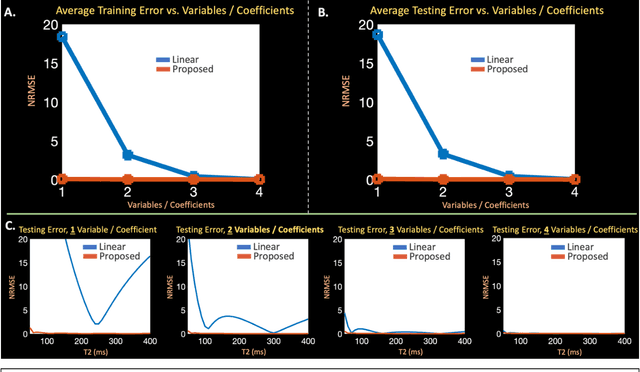
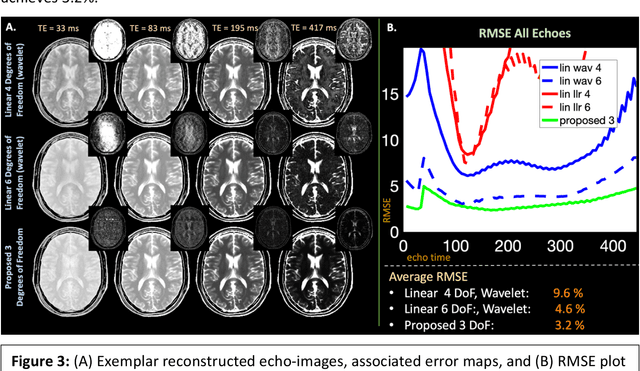
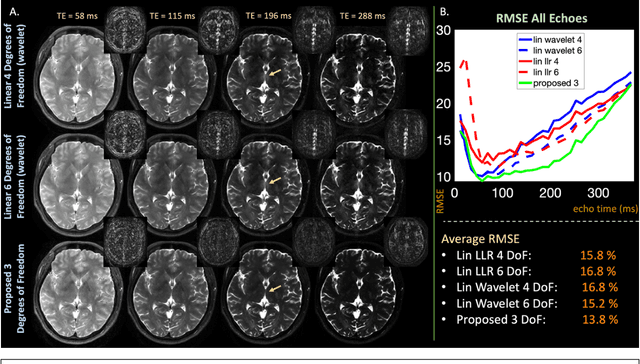
Abstract:Purpose: Training auto-encoders on simulated signal evolution and inserting the decoder into the forward model improves reconstructions through more compact, Bloch-equation-based representations of signal in comparison to linear subspaces. Methods: Building on model-based nonlinear and linear subspace techniques that enable reconstruction of signal dynamics, we train auto-encoders on dictionaries of simulated signal evolution to learn more compact, non-linear, latent representations. The proposed Latent Signal Model framework inserts the decoder portion of the auto-encoder into the forward model and directly reconstructs the latent representation. Latent Signal Models essentially serve as a proxy for fast and feasible differentiation through the Bloch-equations used to simulate signal. This work performs experiments in the context of T2-shuffling, gradient echo EPTI, and MPRAGE-shuffling. We compare how efficiently auto-encoders represent signal evolution in comparison to linear subspaces. Simulation and in-vivo experiments then evaluate if reducing degrees of freedom by inserting the decoder into the forward model improves reconstructions in comparison to subspace constraints. Results: An auto-encoder with one real latent variable represents FSE, EPTI, and MPRAGE signal evolution as well as linear subspaces characterized by four basis vectors. In simulated/in-vivo T2-shuffling and in-vivo EPTI experiments, the proposed framework achieves consistent quantitative NRMSE and qualitative improvement over linear approaches. From qualitative evaluation, the proposed approach yields images with reduced blurring and noise amplification in MPRAGE shuffling experiments. Conclusion: Directly solving for non-linear latent representations of signal evolution improves time-resolved MRI reconstructions through reduced degrees of freedom.
SRDTI: Deep learning-based super-resolution for diffusion tensor MRI
Feb 17, 2021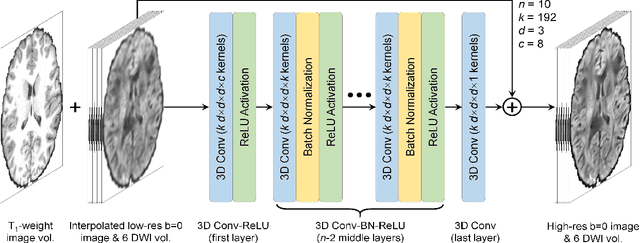
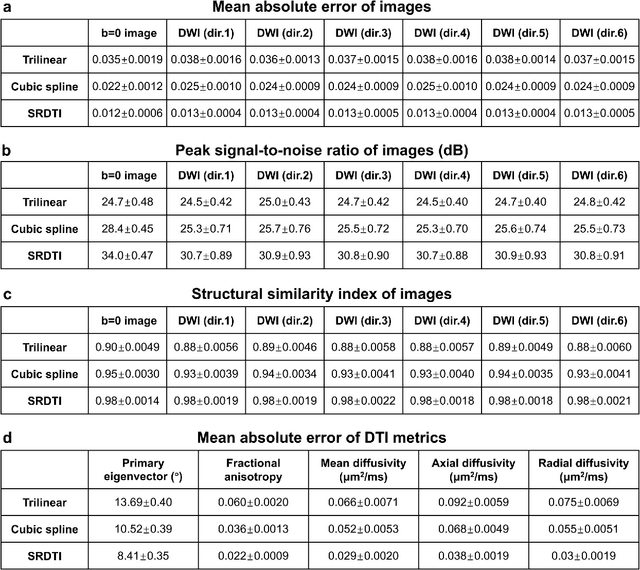
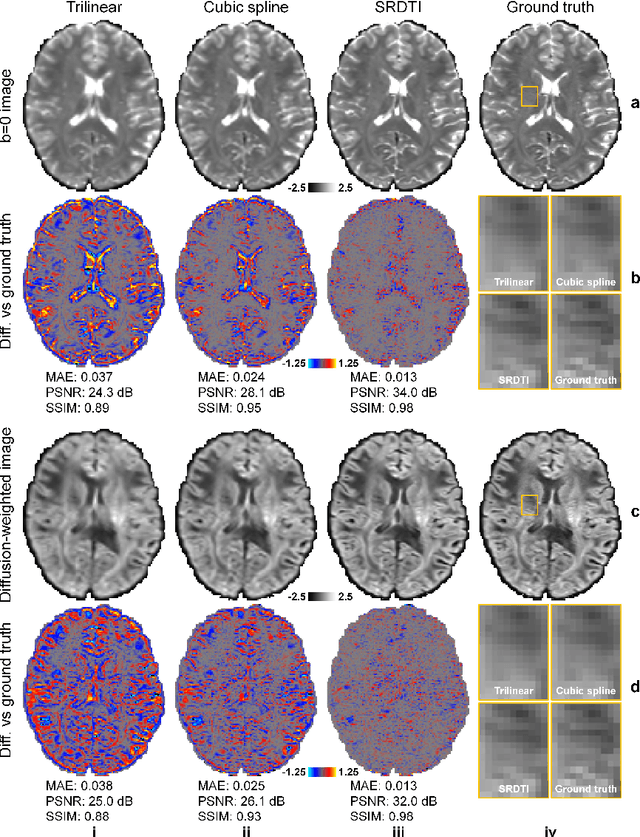
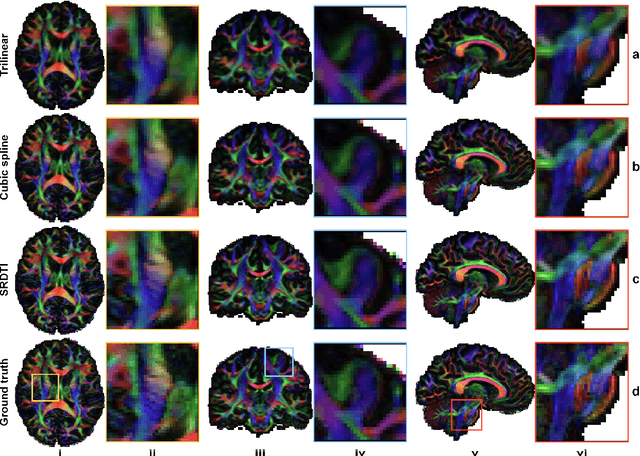
Abstract:High-resolution diffusion tensor imaging (DTI) is beneficial for probing tissue microstructure in fine neuroanatomical structures, but long scan times and limited signal-to-noise ratio pose significant barriers to acquiring DTI at sub-millimeter resolution. To address this challenge, we propose a deep learning-based super-resolution method entitled "SRDTI" to synthesize high-resolution diffusion-weighted images (DWIs) from low-resolution DWIs. SRDTI employs a deep convolutional neural network (CNN), residual learning and multi-contrast imaging, and generates high-quality results with rich textural details and microstructural information, which are more similar to high-resolution ground truth than those from trilinear and cubic spline interpolation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge