Congyu Liao
Enhancing Diffusion-Weighted Images (DWI) for Diffusion MRI: Is it Enough without Non-Diffusion-Weighted B=0 Reference?
May 19, 2025Abstract:Diffusion MRI (dMRI) is essential for studying brain microstructure, but high-resolution imaging remains challenging due to the inherent trade-offs between acquisition time and signal-to-noise ratio (SNR). Conventional methods often optimize only the diffusion-weighted images (DWIs) without considering their relationship with the non-diffusion-weighted (b=0) reference images. However, calculating diffusion metrics, such as the apparent diffusion coefficient (ADC) and diffusion tensor with its derived metrics like fractional anisotropy (FA) and mean diffusivity (MD), relies on the ratio between each DWI and the b=0 image, which is crucial for clinical observation and diagnostics. In this study, we demonstrate that solely enhancing DWIs using a conventional pixel-wise mean squared error (MSE) loss is insufficient, as the error in ratio between generated DWIs and b=0 diverges. We propose a novel ratio loss, defined as the MSE loss between the predicted and ground-truth log of DWI/b=0 ratios. Our results show that incorporating the ratio loss significantly improves the convergence of this ratio error, achieving lower ratio MSE and slightly enhancing the peak signal-to-noise ratio (PSNR) of generated DWIs. This leads to improved dMRI super-resolution and better preservation of b=0 ratio-based features for the derivation of diffusion metrics.
Artificial Intelligence for Neuro MRI Acquisition: A Review
Jun 10, 2024Abstract:Magnetic resonance imaging (MRI) has significantly benefited from the resurgence of artificial intelligence (AI). By leveraging AI's capabilities in large-scale optimization and pattern recognition, innovative methods are transforming the MRI acquisition workflow, including planning, sequence design, and correction of acquisition artifacts. These emerging algorithms demonstrate substantial potential in enhancing the efficiency and throughput of acquisition steps. This review discusses several pivotal AI-based methods in neuro MRI acquisition, focusing on their technological advances, impact on clinical practice, and potential risks.
Advancing low-field MRI with a universal denoising imaging transformer: Towards fast and high-quality imaging
Apr 30, 2024Abstract:Recent developments in low-field (LF) magnetic resonance imaging (MRI) systems present remarkable opportunities for affordable and widespread MRI access. A robust denoising method to overcome the intrinsic low signal-noise-ratio (SNR) barrier is critical to the success of LF MRI. However, current data-driven MRI denoising methods predominantly handle magnitude images and rely on customized models with constrained data diversity and quantity, which exhibit limited generalizability in clinical applications across diverse MRI systems, pulse sequences, and organs. In this study, we present ImT-MRD: a complex-valued imaging transformer trained on a vast number of clinical MRI scans aiming at universal MR denoising at LF systems. Compared with averaging multiple-repeated scans for higher image SNR, the model obtains better image quality from fewer repetitions, demonstrating its capability for accelerating scans under various clinical settings. Moreover, with its complex-valued image input, the model can denoise intermediate results before advanced post-processing and prepare high-quality data for further MRI research. By delivering universal and accurate denoising across clinical and research tasks, our model holds great promise to expedite the evolution of LF MRI for accessible and equal biomedical applications.
An Efficient Algorithm for Spatial-Spectral Partial Volume Compartment Mapping with Applications to Multicomponent Diffusion and Relaxation MRI
Jan 23, 2024



Abstract:It has been previously shown that high-quality partial volume tissue compartment maps can be obtained by combining multiparametric contrast-encoded MRI data acquisition methods with spatially-regularized spectroscopic image estimation techniques. However, the advantages of this combined approach generally come at the expense of substantial computational complexity. In this work, we propose a new algorithm to solve this kind of estimation problem more efficiently. Our algorithm is based on the linearized alternating directions method of multipliers (LADMM), and relies on the introduction of novel quadratic penalty terms to substantially simplify the subproblems that must be solved at each iteration. We evaluate this algorithm on a variety of different estimation problems (diffusion-relaxation, relaxation-relaxation, relaxometry, and magnetic resonance fingerprinting), where we consistently observe substantial (roughly 5$\times$-80$\times$) speed improvements. We expect that this new faster algorithm will lower practical barriers to using spatial regularization and multiparametric contrast-encoded MRI data acquisition methods for partial volume compartment mapping.
High-resolution myelin-water fraction and quantitative relaxation mapping using 3D ViSTa-MR fingerprinting
Dec 21, 2023Abstract:Purpose: This study aims to develop a high-resolution whole-brain multi-parametric quantitative MRI approach for simultaneous mapping of myelin-water fraction (MWF), T1, T2, and proton-density (PD), all within a clinically feasible scan time. Methods: We developed 3D ViSTa-MRF, which combined Visualization of Short Transverse relaxation time component (ViSTa) technique with MR Fingerprinting (MRF), to achieve high-fidelity whole-brain MWF and T1/T2/PD mapping on a clinical 3T scanner. To achieve fast acquisition and memory-efficient reconstruction, the ViSTa-MRF sequence leverages an optimized 3D tiny-golden-angle-shuffling spiral-projection acquisition and joint spatial-temporal subspace reconstruction with optimized preconditioning algorithm. With the proposed ViSTa-MRF approach, high-fidelity direct MWF mapping was achieved without a need for multi-compartment fitting that could introduce bias and/or noise from additional assumptions or priors. Results: The in-vivo results demonstrate the effectiveness of the proposed acquisition and reconstruction framework to provide fast multi-parametric mapping with high SNR and good quality. The in-vivo results of 1mm- and 0.66mm-iso datasets indicate that the MWF values measured by the proposed method are consistent with standard ViSTa results that are 30x slower with lower SNR. Furthermore, we applied the proposed method to enable 5-minute whole-brain 1mm-iso assessment of MWF and T1/T2/PD mappings for infant brain development and for post-mortem brain samples. Conclusions: In this work, we have developed a 3D ViSTa-MRF technique that enables the acquisition of whole-brain MWF, quantitative T1, T2, and PD maps at 1mm and 0.66mm isotropic resolution in 5 and 15 minutes, respectively. This advancement allows for quantitative investigations of myelination changes in the brain.
Rapid Non-cartesian Reconstruction Using an Implicit Representation of GROG Kernels
Oct 16, 2023



Abstract:MRI data is acquired in Fourier space. Data acquisition is typically performed on a Cartesian grid in this space to enable the use of a fast Fourier transform algorithm to achieve fast and efficient reconstruction. However, it has been shown that for multiple applications, non-Cartesian data acquisition can improve the performance of MR imaging by providing fast and more efficient data acquisition, and improving motion robustness. Nonetheless, the image reconstruction process of non-Cartesian data is more involved and can be time-consuming, even through the use of efficient algorithms such as non-uniform FFT (NUFFT). This work provides an efficient approach (iGROG) to transform the non-Cartesian data into Cartesian data, to achieve simpler and faster reconstruction which should help enable non-Cartesian data sampling to be performed more widely in MRI.
3D-EPI Blip-Up/Down Acquisition with CAIPI and Joint Hankel Structured Low-Rank Reconstruction for Rapid Distortion-Free High-Resolution T2* Mapping
Dec 01, 2022Abstract:Purpose: This work aims to develop a novel distortion-free 3D-EPI acquisition and image reconstruction technique for fast and robust, high-resolution, whole-brain imaging as well as quantitative T2* mapping. Methods: 3D-Blip-Up and -Down Acquisition (3D-BUDA) sequence is designed for both single- and multi-echo 3D GRE-EPI imaging using multiple shots with blip-up and -down readouts to encode B0 field map information. Complementary k-space coverage is achieved using controlled aliasing in parallel imaging (CAIPI) sampling across the shots. For image reconstruction, an iterative hard-thresholding algorithm is employed to minimize the cost function that combines field map information informed parallel imaging with the structured low-rank constraint for multi-shot 3D-BUDA data. Extending 3D-BUDA to multi-echo imaging permits T2* mapping. For this, we propose constructing a joint Hankel matrix along both echo and shot dimensions to improve the reconstruction. Results: Experimental results on in vivo multi-echo data demonstrate that, by performing joint reconstruction along with both echo and shot dimensions, reconstruction accuracy is improved compared to standard 3D-BUDA reconstruction. CAIPI sampling is further shown to enhance the image quality. For T2* mapping, T2* values from 3D-Joint-CAIPI-BUDA and reference multi-echo GRE are within limits of agreement as quantified by Bland-Altman analysis. Conclusions: The proposed technique enables rapid 3D distortion-free high-resolution imaging and T2* mapping. Specifically, 3D-BUDA enables 1-mm isotropic whole-brain imaging in 22 s at 3 T and 9 s on a 7 T scanner. The combination of multi-echo 3D-BUDA with CAIPI acquisition and joint reconstruction enables distortion-free whole-brain T2* mapping in 47 s at 1.1x1.1x1.0 mm3 resolution.
BUDA-SAGE with self-supervised denoising enables fast, distortion-free, high-resolution T2, T2*, para- and dia-magnetic susceptibility mapping
Sep 09, 2021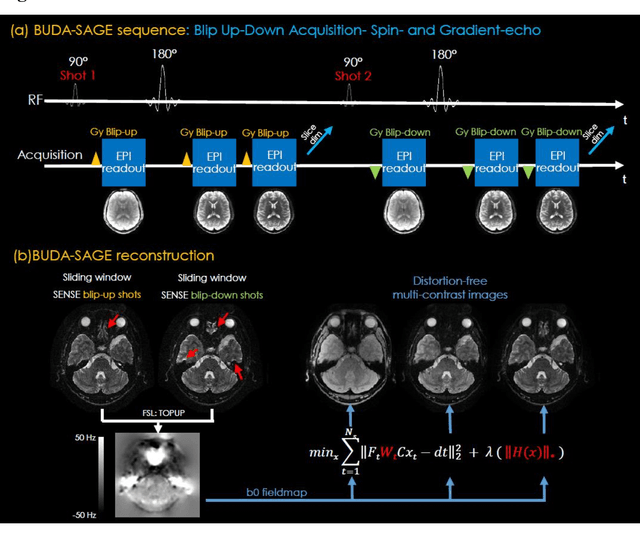
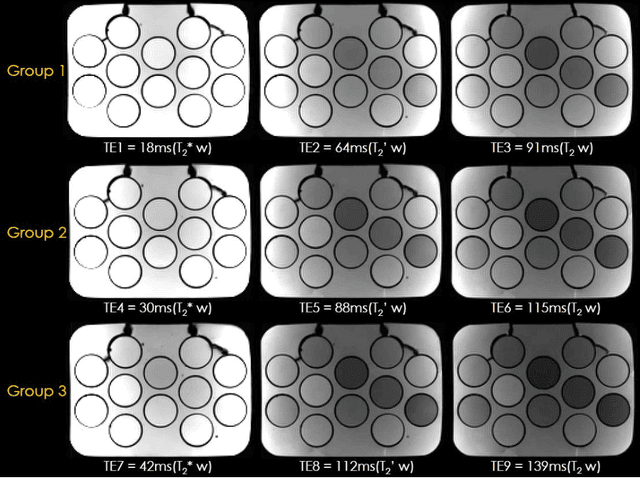
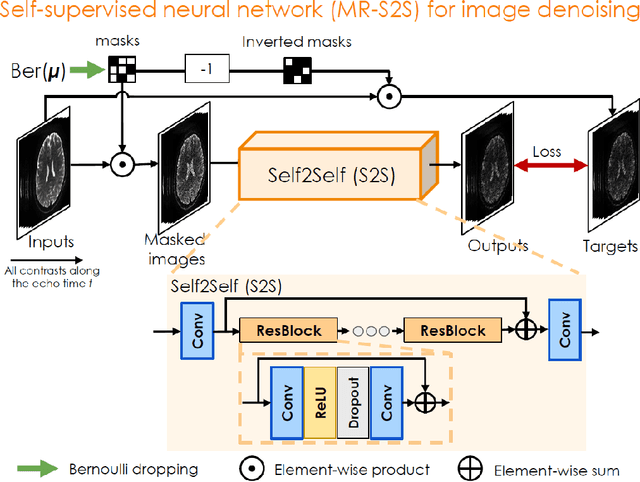

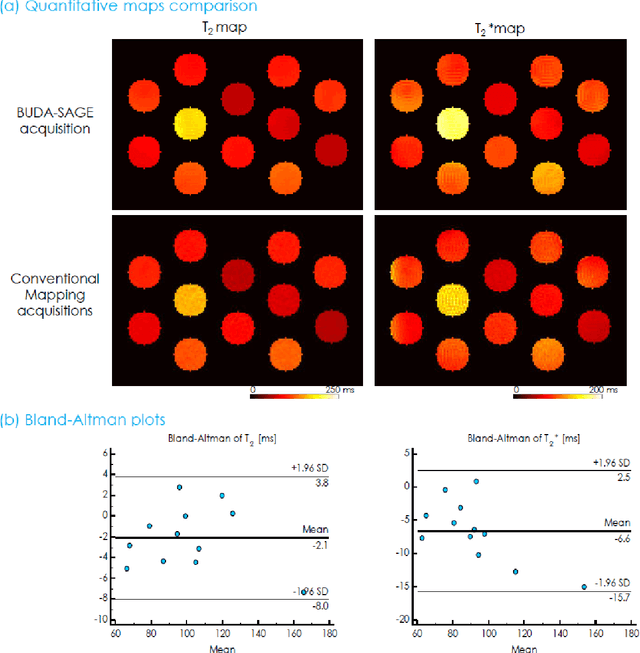
Abstract:To rapidly obtain high resolution T2, T2* and quantitative susceptibility mapping (QSM) source separation maps with whole-brain coverage and high geometric fidelity. We propose Blip Up-Down Acquisition for Spin And Gradient Echo imaging (BUDA-SAGE), an efficient echo-planar imaging (EPI) sequence for quantitative mapping. The acquisition includes multiple T2*-, T2'- and T2-weighted contrasts. We alternate the phase-encoding polarities across the interleaved shots in this multi-shot navigator-free acquisition. A field map estimated from interim reconstructions was incorporated into the joint multi-shot EPI reconstruction with a structured low rank constraint to eliminate geometric distortion. A self-supervised MR-Self2Self (MR-S2S) neural network (NN) was utilized to perform denoising after BUDA reconstruction to boost SNR. Employing Slider encoding allowed us to reach 1 mm isotropic resolution by performing super-resolution reconstruction on BUDA-SAGE volumes acquired with 2 mm slice thickness. Quantitative T2 and T2* maps were obtained using Bloch dictionary matching on the reconstructed echoes. QSM was estimated using nonlinear dipole inversion (NDI) on the gradient echoes. Starting from the estimated R2 and R2* maps, R2' information was derived and used in source separation QSM reconstruction, which provided additional para- and dia-magnetic susceptibility maps. In vivo results demonstrate the ability of BUDA-SAGE to provide whole-brain, distortion-free, high-resolution multi-contrast images and quantitative T2 and T2* maps, as well as yielding para- and dia-magnetic susceptibility maps. Derived quantitative maps showed comparable values to conventional mapping methods in phantom and in vivo measurements. BUDA-SAGE acquisition with self-supervised denoising and Slider encoding enabled rapid, distortion-free, whole-brain T2, T2* mapping at 1 mm3 isotropic resolution in 90 seconds.
Optimized multi-axis spiral projection MR fingerprinting with subspace reconstruction for rapid whole-brain high-isotropic-resolution quantitative imaging
Aug 12, 2021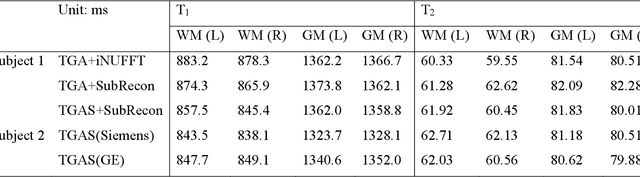

Abstract:Purpose: To improve image quality and accelerate the acquisition of 3D MRF. Methods: Building on the multi-axis spiral-projection MRF technique, a subspace reconstruction with locally low rank (LLR) constraint and a modified spiral-projection spatiotemporal encoding scheme termed tiny-golden-angle-shuffling (TGAS) were implemented for rapid whole-brain high-resolution quantitative mapping. The LLR regularization parameter and the number of subspace bases were tuned using retrospective in-vivo data and simulated examinations, respectively. B0 inhomogeneity correction using multi-frequency interpolation was incorporated into the subspace reconstruction to further improve the image quality by mitigating blurring caused by off-resonance effect. Results: The proposed MRF acquisition and reconstruction framework can produce provide high quality 1-mm isotropic whole-brain quantitative maps in a total acquisition time of 1 minute 55 seconds, with higher-quality results than ones obtained from the previous approach in 6 minutes. The comparison of quantitative results indicates that neither the subspace reconstruction nor the TGAS trajectory induce bias for T1 and T2 mapping. High quality whole-brain MRF data were also obtained at 0.66-mm isotropic resolution in 4 minutes using the proposed technique, where the increased resolution was shown to improve visualization of subtle brain structures. Conclusion: The proposed TGAS-SPI-MRF with optimized spiral-projection trajectory and subspace reconstruction can enable high-resolution quantitative mapping with faster acquisition speed.
eRAKI: Fast Robust Artificial neural networks for K-space Interpolation (RAKI) with Coil Combination and Joint Reconstruction
Jul 07, 2021Abstract:RAKI can perform database-free MRI reconstruction by training models using only auto-calibration signal (ACS) from each specific scan. As it trains a separate model for each individual coil, learning and inference with RAKI can be computationally prohibitive, particularly for large 3D datasets. In this abstract, we accelerate RAKI more than 200 times by directly learning a coil-combined target and further improve the reconstruction performance using joint reconstruction across multiple echoes together with an elliptical-CAIPI sampling approach. We further deploy these improvements in quantitative imaging and rapidly obtain T2 and T2* parameter maps from a fast EPTI scan.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge