Huihui Ye
MIMOSA: Multi-parametric Imaging using Multiple-echoes with Optimized Simultaneous Acquisition for highly-efficient quantitative MRI
Aug 13, 2025



Abstract:Purpose: To develop a new sequence, MIMOSA, for highly-efficient T1, T2, T2*, proton density (PD), and source separation quantitative susceptibility mapping (QSM). Methods: MIMOSA was developed based on 3D-quantification using an interleaved Look-Locker acquisition sequence with T2 preparation pulse (3D-QALAS) by combining 3D turbo Fast Low Angle Shot (FLASH) and multi-echo gradient echo acquisition modules with a spiral-like Cartesian trajectory to facilitate highly-efficient acquisition. Simulations were performed to optimize the sequence. Multi-contrast/-slice zero-shot self-supervised learning algorithm was employed for reconstruction. The accuracy of quantitative mapping was assessed by comparing MIMOSA with 3D-QALAS and reference techniques in both ISMRM/NIST phantom and in-vivo experiments. MIMOSA's acceleration capability was assessed at R = 3.3, 6.5, and 11.8 in in-vivo experiments, with repeatability assessed through scan-rescan studies. Beyond the 3T experiments, mesoscale quantitative mapping was performed at 750 um isotropic resolution at 7T. Results: Simulations demonstrated that MIMOSA achieved improved parameter estimation accuracy compared to 3D-QALAS. Phantom experiments indicated that MIMOSA exhibited better agreement with the reference techniques than 3D-QALAS. In-vivo experiments demonstrated that an acceleration factor of up to R = 11.8-fold can be achieved while preserving parameter estimation accuracy, with intra-class correlation coefficients of 0.998 (T1), 0.973 (T2), 0.947 (T2*), 0.992 (QSM), 0.987 (paramagnetic susceptibility), and 0.977 (diamagnetic susceptibility) in scan-rescan studies. Whole-brain T1, T2, T2*, PD, source separation QSM were obtained with 1 mm isotropic resolution in 3 min at 3T and 750 um isotropic resolution in 13 min at 7T. Conclusion: MIMOSA demonstrated potential for highly-efficient multi-parametric mapping.
Artificial Intelligence for Neuro MRI Acquisition: A Review
Jun 10, 2024Abstract:Magnetic resonance imaging (MRI) has significantly benefited from the resurgence of artificial intelligence (AI). By leveraging AI's capabilities in large-scale optimization and pattern recognition, innovative methods are transforming the MRI acquisition workflow, including planning, sequence design, and correction of acquisition artifacts. These emerging algorithms demonstrate substantial potential in enhancing the efficiency and throughput of acquisition steps. This review discusses several pivotal AI-based methods in neuro MRI acquisition, focusing on their technological advances, impact on clinical practice, and potential risks.
Accelerated MR Fingerprinting with Low-Rank and Generative Subspace Modeling
May 25, 2023



Abstract:Magnetic Resonance (MR) Fingerprinting is an emerging multi-parametric quantitative MR imaging technique, for which image reconstruction methods utilizing low-rank and subspace constraints have achieved state-of-the-art performance. However, this class of methods often suffers from an ill-conditioned model-fitting issue, which degrades the performance as the data acquisition lengths become short and/or the signal-to-noise ratio becomes low. To address this problem, we present a new image reconstruction method for MR Fingerprinting, integrating low-rank and subspace modeling with a deep generative prior. Specifically, the proposed method captures the strong spatiotemporal correlation of contrast-weighted time-series images in MR Fingerprinting via a low-rank factorization. Further, it utilizes an untrained convolutional generative neural network to represent the spatial subspace of the low-rank model, while estimating the temporal subspace of the model from simulated magnetization evolutions generated based on spin physics. Here the architecture of the generative neural network serves as an effective regularizer for the ill-conditioned inverse problem without additional spatial training data that are often expensive to acquire. The proposed formulation results in a non-convex optimization problem, for which we develop an algorithm based on variable splitting and alternating direction method of multipliers.We evaluate the performance of the proposed method with numerical simulations and in vivo experiments and demonstrate that the proposed method outperforms the state-of-the-art low-rank and subspace reconstruction.
Accurate parameter estimation using scan-specific unsupervised deep learning for relaxometry and MR fingerprinting
Dec 13, 2021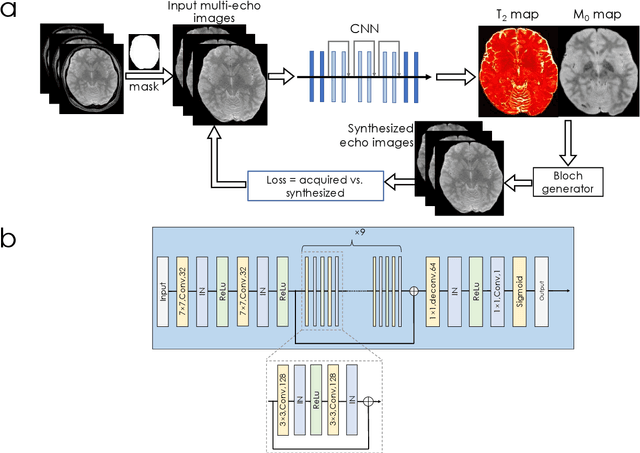
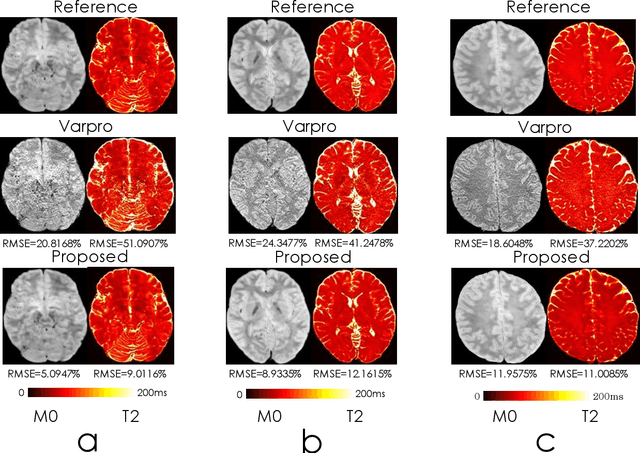
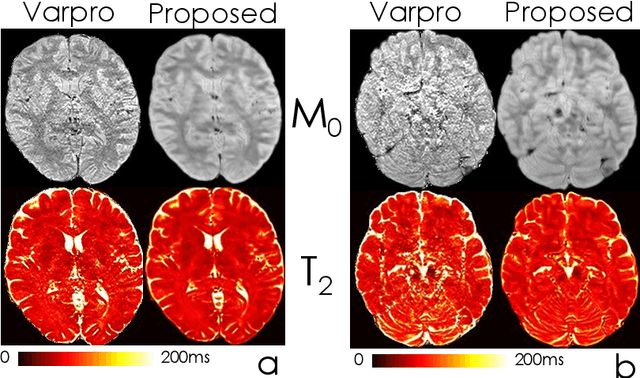
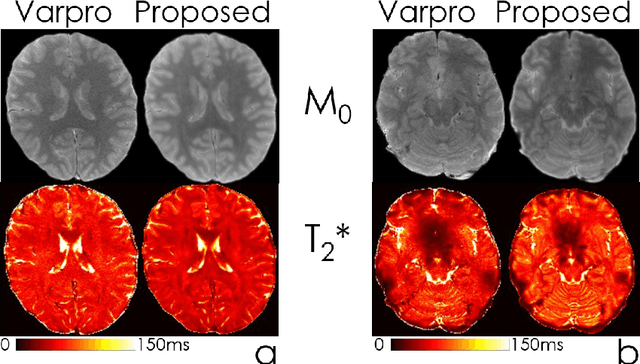
Abstract:We propose an unsupervised convolutional neural network (CNN) for relaxation parameter estimation. This network incorporates signal relaxation and Bloch simulations while taking advantage of residual learning and spatial relations across neighboring voxels. Quantification accuracy and robustness to noise is shown to be significantly improved compared to standard parameter estimation methods in numerical simulations and in vivo data for multi-echo T2 and T2* mapping. The combination of the proposed network with subspace modeling and MR fingerprinting (MRF) from highly undersampled data permits high quality T1 and T2 mapping.
BUDA-SAGE with self-supervised denoising enables fast, distortion-free, high-resolution T2, T2*, para- and dia-magnetic susceptibility mapping
Sep 09, 2021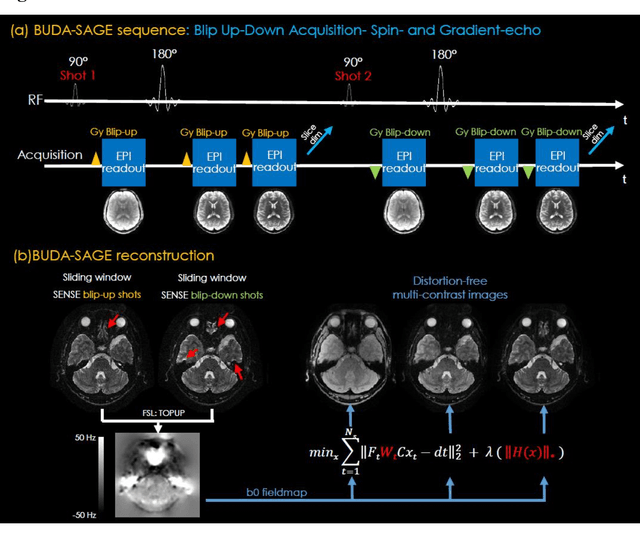
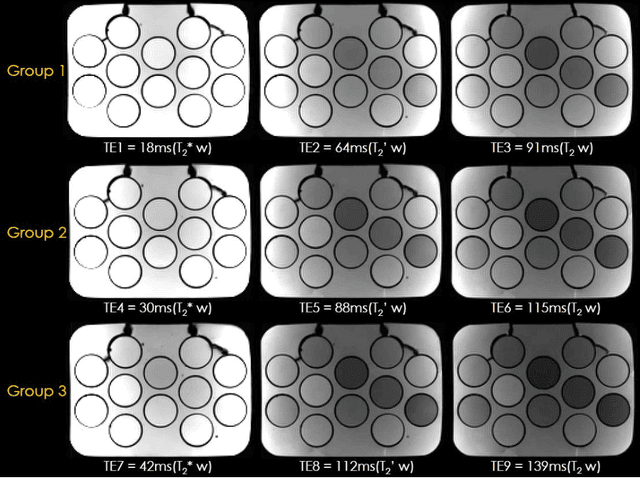
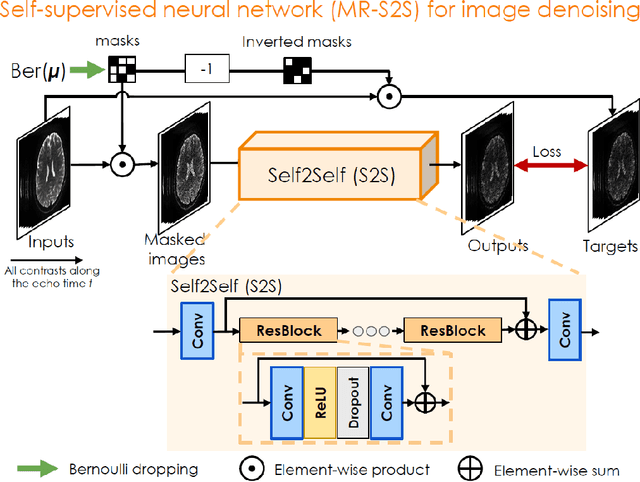

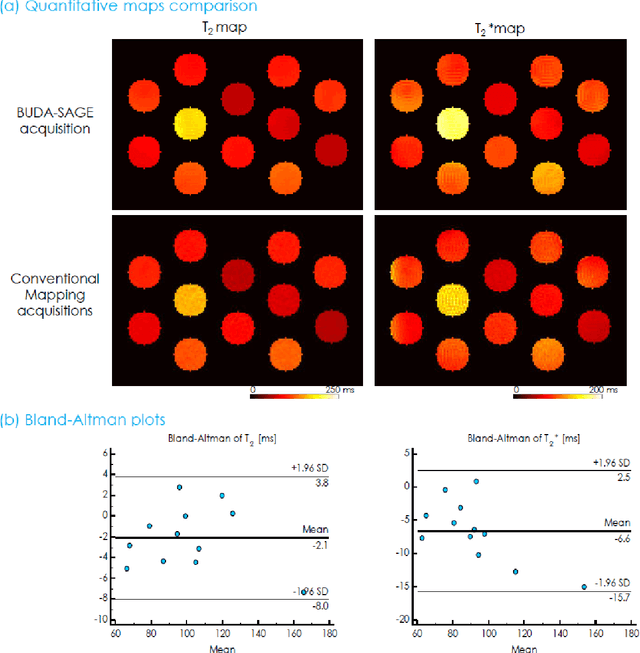
Abstract:To rapidly obtain high resolution T2, T2* and quantitative susceptibility mapping (QSM) source separation maps with whole-brain coverage and high geometric fidelity. We propose Blip Up-Down Acquisition for Spin And Gradient Echo imaging (BUDA-SAGE), an efficient echo-planar imaging (EPI) sequence for quantitative mapping. The acquisition includes multiple T2*-, T2'- and T2-weighted contrasts. We alternate the phase-encoding polarities across the interleaved shots in this multi-shot navigator-free acquisition. A field map estimated from interim reconstructions was incorporated into the joint multi-shot EPI reconstruction with a structured low rank constraint to eliminate geometric distortion. A self-supervised MR-Self2Self (MR-S2S) neural network (NN) was utilized to perform denoising after BUDA reconstruction to boost SNR. Employing Slider encoding allowed us to reach 1 mm isotropic resolution by performing super-resolution reconstruction on BUDA-SAGE volumes acquired with 2 mm slice thickness. Quantitative T2 and T2* maps were obtained using Bloch dictionary matching on the reconstructed echoes. QSM was estimated using nonlinear dipole inversion (NDI) on the gradient echoes. Starting from the estimated R2 and R2* maps, R2' information was derived and used in source separation QSM reconstruction, which provided additional para- and dia-magnetic susceptibility maps. In vivo results demonstrate the ability of BUDA-SAGE to provide whole-brain, distortion-free, high-resolution multi-contrast images and quantitative T2 and T2* maps, as well as yielding para- and dia-magnetic susceptibility maps. Derived quantitative maps showed comparable values to conventional mapping methods in phantom and in vivo measurements. BUDA-SAGE acquisition with self-supervised denoising and Slider encoding enabled rapid, distortion-free, whole-brain T2, T2* mapping at 1 mm3 isotropic resolution in 90 seconds.
Accelerated MRI Reconstruction with Separable and Enhanced Low-Rank Hankel Regularization
Jul 24, 2021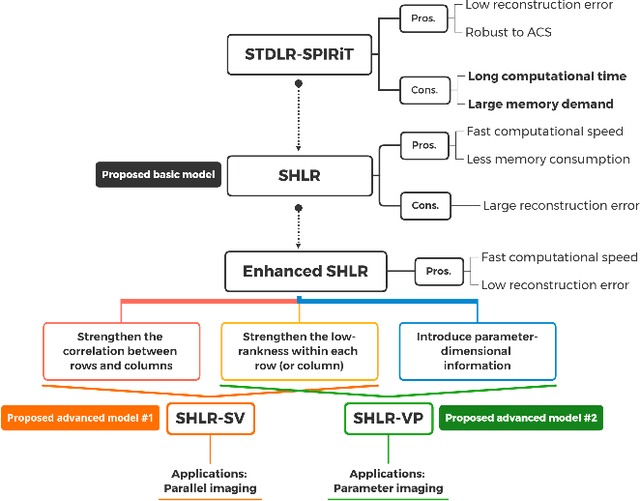
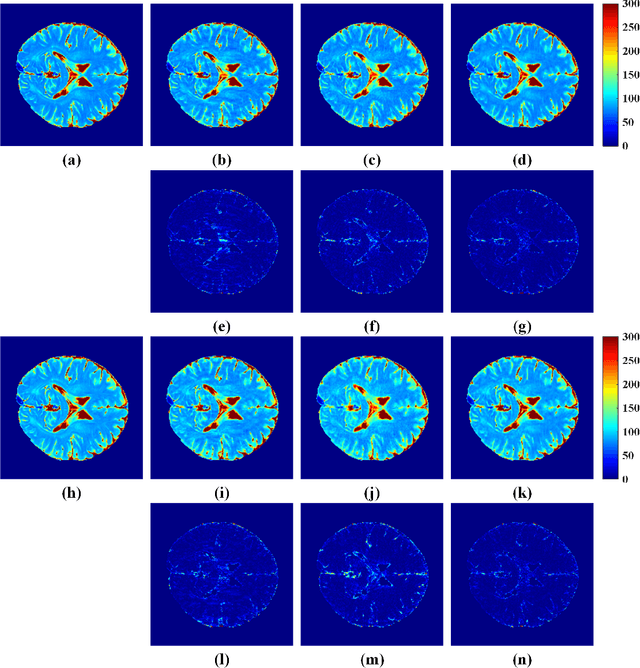
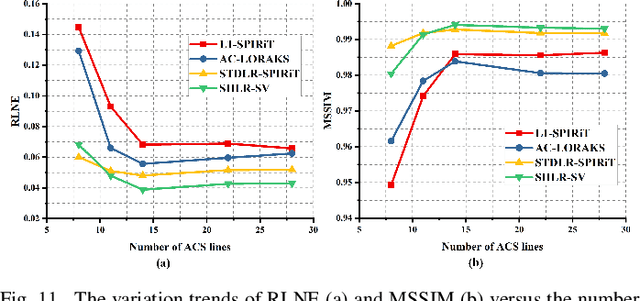
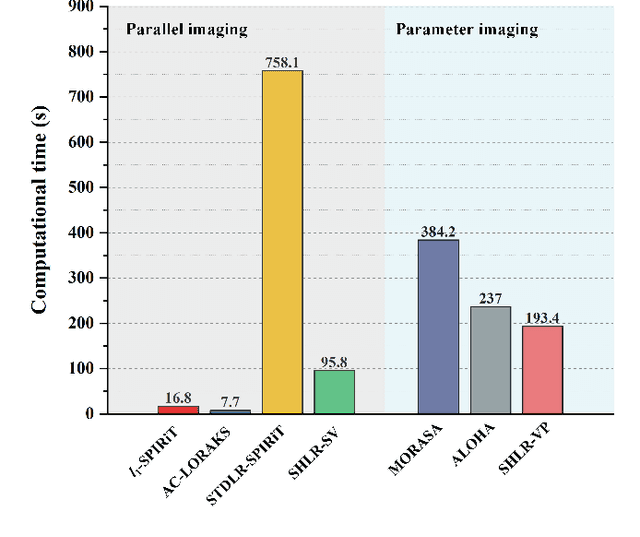
Abstract:The combination of the sparse sampling and the low-rank structured matrix reconstruction has shown promising performance, enabling a significant reduction of the magnetic resonance imaging data acquisition time. However, the low-rank structured approaches demand considerable memory consumption and are time-consuming due to a noticeable number of matrix operations performed on the huge-size block Hankel-like matrix. In this work, we proposed a novel framework to utilize the low-rank property but meanwhile to achieve faster reconstructions and promising results. The framework allows us to enforce the low-rankness of Hankel matrices constructing from 1D vectors instead of 2D matrices from 1D vectors and thus avoid the construction of huge block Hankel matrix for 2D k-space matrices. Moreover, under this framework, we can easily incorporate other information, such as the smooth phase of the image and the low-rankness in the parameter dimension, to further improve the image quality. We built and validated two models for parallel and parameter magnetic resonance imaging experiments, respectively. Our retrospective in-vivo results indicate that the proposed approaches enable faster reconstructions than the state-of-the-art approaches, e.g., about 8x faster than STDLRSPIRiT, and faithful removal of undersampling artifacts.
A Multi-resolution Model for Histopathology Image Classification and Localization with Multiple Instance Learning
Nov 05, 2020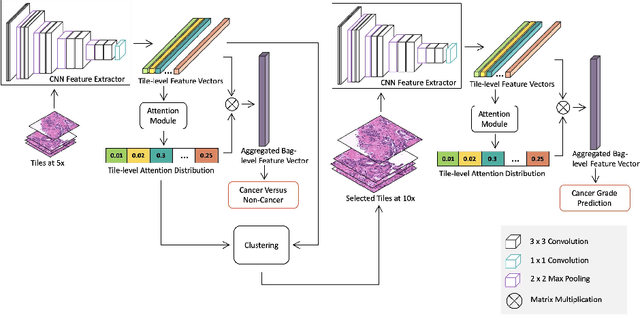
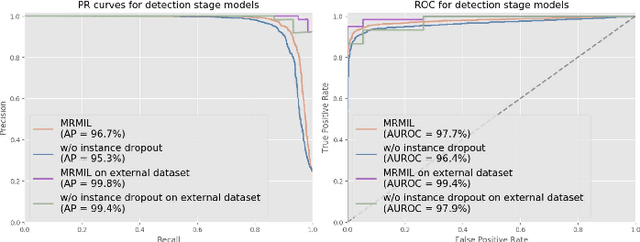
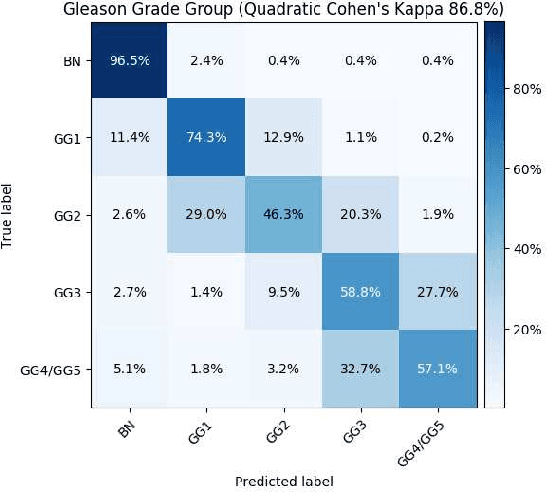
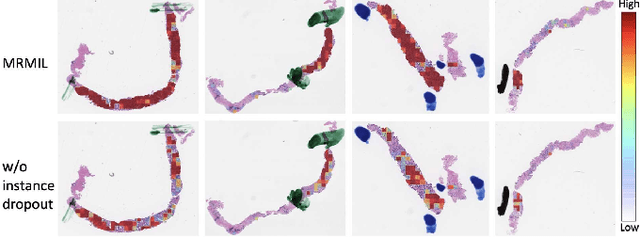
Abstract:Histopathological images provide rich information for disease diagnosis. Large numbers of histopathological images have been digitized into high resolution whole slide images, opening opportunities in developing computational image analysis tools to reduce pathologists' workload and potentially improve inter- and intra- observer agreement. Most previous work on whole slide image analysis has focused on classification or segmentation of small pre-selected regions-of-interest, which requires fine-grained annotation and is non-trivial to extend for large-scale whole slide analysis. In this paper, we proposed a multi-resolution multiple instance learning model that leverages saliency maps to detect suspicious regions for fine-grained grade prediction. Instead of relying on expensive region- or pixel-level annotations, our model can be trained end-to-end with only slide-level labels. The model is developed on a large-scale prostate biopsy dataset containing 20,229 slides from 830 patients. The model achieved 92.7% accuracy, 81.8% Cohen's Kappa for benign, low grade (i.e. Grade group 1) and high grade (i.e. Grade group >= 2) prediction, an area under the receiver operating characteristic curve (AUROC) of 98.2% and an average precision (AP) of 97.4% for differentiating malignant and benign slides. The model obtained an AUROC of 99.4% and an AP of 99.8% for cancer detection on an external dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge