Borjan Gagoski
MIMOSA: Multi-parametric Imaging using Multiple-echoes with Optimized Simultaneous Acquisition for highly-efficient quantitative MRI
Aug 13, 2025



Abstract:Purpose: To develop a new sequence, MIMOSA, for highly-efficient T1, T2, T2*, proton density (PD), and source separation quantitative susceptibility mapping (QSM). Methods: MIMOSA was developed based on 3D-quantification using an interleaved Look-Locker acquisition sequence with T2 preparation pulse (3D-QALAS) by combining 3D turbo Fast Low Angle Shot (FLASH) and multi-echo gradient echo acquisition modules with a spiral-like Cartesian trajectory to facilitate highly-efficient acquisition. Simulations were performed to optimize the sequence. Multi-contrast/-slice zero-shot self-supervised learning algorithm was employed for reconstruction. The accuracy of quantitative mapping was assessed by comparing MIMOSA with 3D-QALAS and reference techniques in both ISMRM/NIST phantom and in-vivo experiments. MIMOSA's acceleration capability was assessed at R = 3.3, 6.5, and 11.8 in in-vivo experiments, with repeatability assessed through scan-rescan studies. Beyond the 3T experiments, mesoscale quantitative mapping was performed at 750 um isotropic resolution at 7T. Results: Simulations demonstrated that MIMOSA achieved improved parameter estimation accuracy compared to 3D-QALAS. Phantom experiments indicated that MIMOSA exhibited better agreement with the reference techniques than 3D-QALAS. In-vivo experiments demonstrated that an acceleration factor of up to R = 11.8-fold can be achieved while preserving parameter estimation accuracy, with intra-class correlation coefficients of 0.998 (T1), 0.973 (T2), 0.947 (T2*), 0.992 (QSM), 0.987 (paramagnetic susceptibility), and 0.977 (diamagnetic susceptibility) in scan-rescan studies. Whole-brain T1, T2, T2*, PD, source separation QSM were obtained with 1 mm isotropic resolution in 3 min at 3T and 750 um isotropic resolution in 13 min at 7T. Conclusion: MIMOSA demonstrated potential for highly-efficient multi-parametric mapping.
SE-Equivariant and Noise-Invariant 3D Motion Tracking in Medical Images
Dec 21, 2023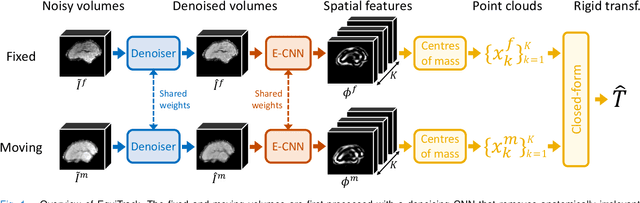
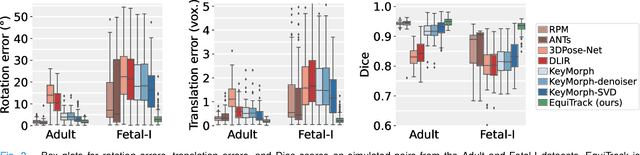
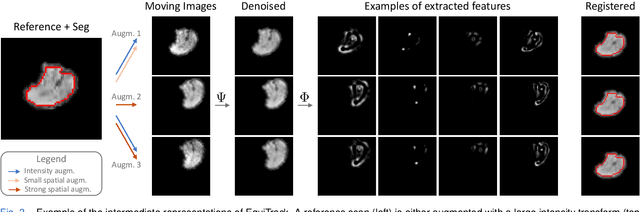
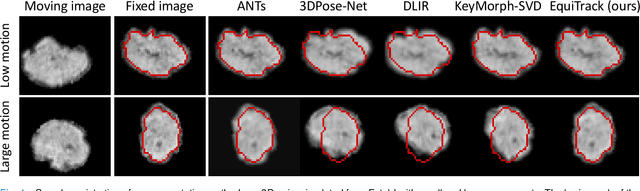
Abstract:Rigid motion tracking is paramount in many medical imaging applications where movements need to be detected, corrected, or accounted for. Modern strategies rely on convolutional neural networks (CNN) and pose this problem as rigid registration. Yet, CNNs do not exploit natural symmetries in this task, as they are equivariant to translations (their outputs shift with their inputs) but not to rotations. Here we propose EquiTrack, the first method that uses recent steerable SE(3)-equivariant CNNs (E-CNN) for motion tracking. While steerable E-CNNs can extract corresponding features across different poses, testing them on noisy medical images reveals that they do not have enough learning capacity to learn noise invariance. Thus, we introduce a hybrid architecture that pairs a denoiser with an E-CNN to decouple the processing of anatomically irrelevant intensity features from the extraction of equivariant spatial features. Rigid transforms are then estimated in closed-form. EquiTrack outperforms state-of-the-art learning and optimisation methods for motion tracking in adult brain MRI and fetal MRI time series. Our code is available at github.com/BBillot/equitrack.
Zero-DeepSub: Zero-Shot Deep Subspace Reconstruction for Rapid Multiparametric Quantitative MRI Using 3D-QALAS
Jul 04, 2023Abstract:Purpose: To develop and evaluate methods for 1) reconstructing 3D-quantification using an interleaved Look-Locker acquisition sequence with T2 preparation pulse (3D-QALAS) time-series images using a low-rank subspace method, which enables accurate and rapid T1 and T2 mapping, and 2) improving the fidelity of subspace QALAS by combining scan-specific deep-learning-based reconstruction and subspace modeling. Methods: A low-rank subspace method for 3D-QALAS (i.e., subspace QALAS) and zero-shot deep-learning subspace method (i.e., Zero-DeepSub) were proposed for rapid and high fidelity T1 and T2 mapping and time-resolved imaging using 3D-QALAS. Using an ISMRM/NIST system phantom, the accuracy of the T1 and T2 maps estimated using the proposed methods was evaluated by comparing them with reference techniques. The reconstruction performance of the proposed subspace QALAS using Zero-DeepSub was evaluated in vivo and compared with conventional QALAS at high reduction factors of up to 9-fold. Results: Phantom experiments showed that subspace QALAS had good linearity with respect to the reference methods while reducing biases compared to conventional QALAS, especially for T2 maps. Moreover, in vivo results demonstrated that subspace QALAS had better g-factor maps and could reduce voxel blurring, noise, and artifacts compared to conventional QALAS and showed robust performance at up to 9-fold acceleration with Zero-DeepSub, which enabled whole-brain T1, T2, and PD mapping at 1 mm isotropic resolution within 2 min of scan time. Conclusion: The proposed subspace QALAS along with Zero-DeepSub enabled high fidelity and rapid whole-brain multiparametric quantification and time-resolved imaging.
SSL-QALAS: Self-Supervised Learning for Rapid Multiparameter Estimation in Quantitative MRI Using 3D-QALAS
Feb 28, 2023Abstract:Purpose: To develop and evaluate a method for rapid estimation of multiparametric T1, T2, proton density (PD), and inversion efficiency (IE) maps from 3D-quantification using an interleaved Look-Locker acquisition sequence with T2 preparation pulse (3D-QALAS) measurements using self-supervised learning (SSL) without the need for an external dictionary. Methods: A SSL-based QALAS mapping method (SSL-QALAS) was developed for rapid and dictionary-free estimation of multiparametric maps from 3D-QALAS measurements. The accuracy of the reconstructed quantitative maps using dictionary matching and SSL-QALAS was evaluated by comparing the estimated T1 and T2 values with those obtained from the reference methods on an ISMRM/NIST phantom. The SSL-QALAS and the dictionary matching methods were also compared in vivo, and generalizability was evaluated by comparing the scan-specific, pre-trained, and transfer learning models. Results: Phantom experiments showed that both the dictionary matching and SSL-QALAS methods produced T1 and T2 estimates that had a strong linear agreement with the reference values in the ISMRM/NIST phantom. Further, SSL-QALAS showed similar performance with dictionary matching in reconstructing the T1, T2, PD, and IE maps on in vivo data. Rapid reconstruction of multiparametric maps was enabled by inferring the data using a pre-trained SSL-QALAS model within 10 s. Fast scan-specific tuning was also demonstrated by fine-tuning the pre-trained model with the target subject's data within 15 min. Conclusion: The proposed SSL-QALAS method enabled rapid reconstruction of multiparametric maps from 3D-QALAS measurements without an external dictionary or labeled ground-truth training data.
Time-efficient, High Resolution 3T Whole Brain Quantitative Relaxometry using 3D-QALAS with Wave-CAIPI Readouts
Nov 08, 2022

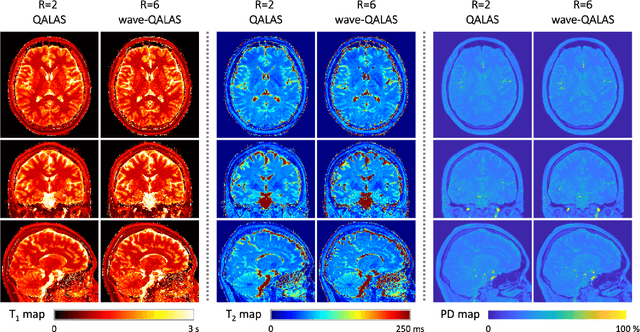

Abstract:Purpose: Volumetric, high resolution, quantitative mapping of brain tissues relaxation properties is hindered by long acquisition times and SNR challenges. This study, for the first time, combines the time efficient wave-CAIPI readouts into the 3D-QALAS acquisition scheme, enabling full brain quantitative T1, T2 and PD maps at 1.15 isotropic voxels in only 3 minutes. Methods: Wave-CAIPI readouts were embedded in the standard 3d-QALAS encoding scheme, enabling full brain quantitative parameter maps (T1, T2 and PD) at acceleration factors of R=3x2 with minimum SNR loss due to g-factor penalties. The quantitative maps using the accelerated protocol were quantitatively compared against those obtained from conventional 3D-QALAS sequence using GRAPPA acceleration of R=2 in the ISMRM NIST phantom, and ten healthy volunteers. To show the feasibility of the proposed methods in clinical settings, the accelerated wave-CAIPI 3D-QALAS sequence was also employed in pediatric patients undergoing clinical MRI examinations. Results: When tested in both the ISMRM/NIST phantom and 7 healthy volunteers, the quantitative maps using the accelerated protocol showed excellent agreement against those obtained from conventional 3D-QALAS at R=2. Conclusion: 3D-QALAS enhanced with wave-CAIPI readouts enables time-efficient, full brain quantitative T1, T2 and PD mapping at 1.15 in 3 minutes at R=3x2 acceleration. When tested on the NIST phantom and 7 healthy volunteers, the quantitative maps obtained from the accelerated wave-CAIPI 3D-QALAS protocol showed very similar values to those obtained from the standard 3D-QALAS (R=2) protocol, alluding to the robustness and reliability of the proposed methods. This study also shows that the accelerated protocol can be effectively employed in pediatric patient populations, making high-quality high-resolution full brain quantitative imaging feasible in clinical settings.
Wave-Encoded Model-based Deep Learning for Highly Accelerated Imaging with Joint Reconstruction
Feb 06, 2022Abstract:Purpose: To propose a wave-encoded model-based deep learning (wave-MoDL) strategy for highly accelerated 3D imaging and joint multi-contrast image reconstruction, and further extend this to enable rapid quantitative imaging using an interleaved look-locker acquisition sequence with T2 preparation pulse (3D-QALAS). Method: Recently introduced MoDL technique successfully incorporates convolutional neural network (CNN)-based regularizers into physics-based parallel imaging reconstruction using a small number of network parameters. Wave-CAIPI is an emerging parallel imaging method that accelerates the imaging speed by employing sinusoidal gradients in the phase- and slice-encoding directions during the readout to take better advantage of 3D coil sensitivity profiles. In wave-MoDL, we propose to combine the wave-encoding strategy with unrolled network constraints to accelerate the acquisition speed while enforcing wave-encoded data consistency. We further extend wave-MoDL to reconstruct multi-contrast data with controlled aliasing in parallel imaging (CAIPI) sampling patterns to leverage similarity between multiple images to improve the reconstruction quality. Result: Wave-MoDL enables a 47-second MPRAGE acquisition at 1 mm resolution at 16-fold acceleration. For quantitative imaging, wave-MoDL permits a 2-minute acquisition for T1, T2, and proton density mapping at 1 mm resolution at 12-fold acceleration, from which contrast weighted images can be synthesized as well. Conclusion: Wave-MoDL allows rapid MR acquisition and high-fidelity image reconstruction and may facilitate clinical and neuroscientific applications by incorporating unrolled neural networks into wave-CAIPI reconstruction.
Rapid head-pose detection for automated slice prescription of fetal-brain MRI
Oct 08, 2021Abstract:In fetal-brain MRI, head-pose changes between prescription and acquisition present a challenge to obtaining the standard sagittal, coronal and axial views essential to clinical assessment. As motion limits acquisitions to thick slices that preclude retrospective resampling, technologists repeat ~55-second stack-of-slices scans (HASTE) with incrementally reoriented field of view numerous times, deducing the head pose from previous stacks. To address this inefficient workflow, we propose a robust head-pose detection algorithm using full-uterus scout scans (EPI) which take ~5 seconds to acquire. Our ~2-second procedure automatically locates the fetal brain and eyes, which we derive from maximally stable extremal regions (MSERs). The success rate of the method exceeds 94% in the third trimester, outperforming a trained technologist by up to 20%. The pipeline may be used to automatically orient the anatomical sequence, removing the need to estimate the head pose from 2D views and reducing delays during which motion can occur.
* 19 pages, 10 figures, 2 tables, fetal MRI, head-pose detection, MSER, scan automation, scan prescription, slice positioning, final published version
Semi-Supervised Learning for Fetal Brain MRI Quality Assessment with ROI consistency
Jun 23, 2020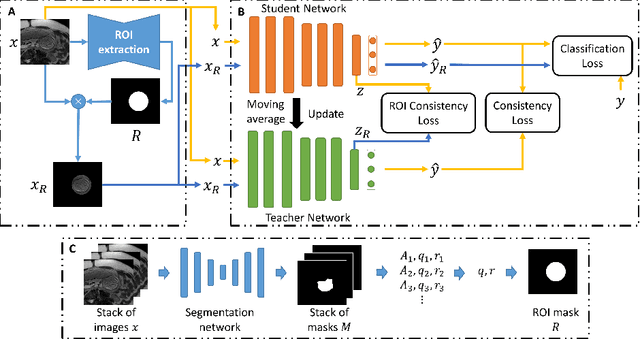
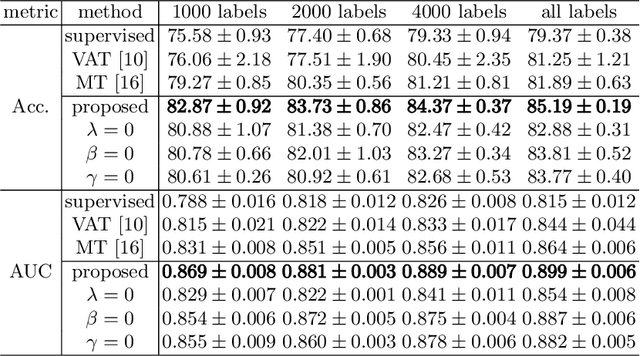
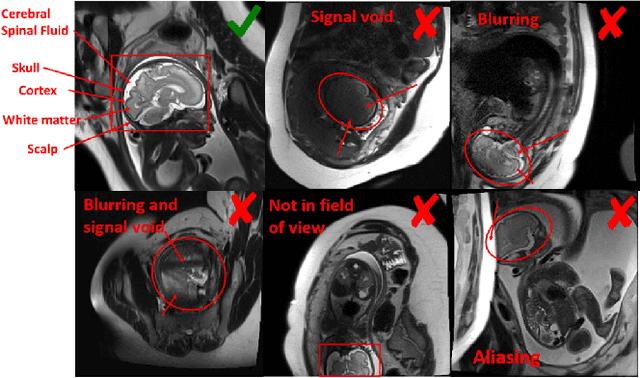
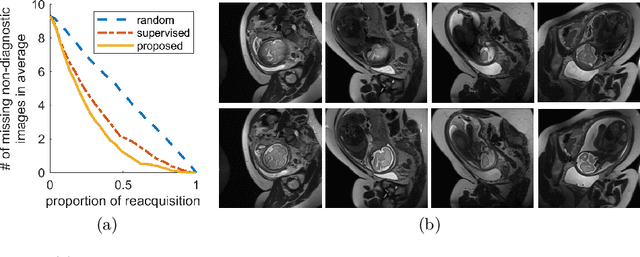
Abstract:Fetal brain MRI is useful for diagnosing brain abnormalities but is challenged by fetal motion. The current protocol for T2-weighted fetal brain MRI is not robust to motion so image volumes are degraded by inter- and intra- slice motion artifacts. Besides, manual annotation for fetal MR image quality assessment are usually time-consuming. Therefore, in this work, a semi-supervised deep learning method that detects slices with artifacts during the brain volume scan is proposed. Our method is based on the mean teacher model, where we not only enforce consistency between student and teacher models on the whole image, but also adopt an ROI consistency loss to guide the network to focus on the brain region. The proposed method is evaluated on a fetal brain MR dataset with 11,223 labeled images and more than 200,000 unlabeled images. Results show that compared with supervised learning, the proposed method can improve model accuracy by about 6\% and outperform other state-of-the-art semi-supervised learning methods. The proposed method is also implemented and evaluated on an MR scanner, which demonstrates the feasibility of online image quality assessment and image reacquisition during fetal MR scans.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge