Ines Machado
ST-Prompt Guided Histological Hypergraph Learning for Spatial Gene Expression Prediction
Mar 21, 2025
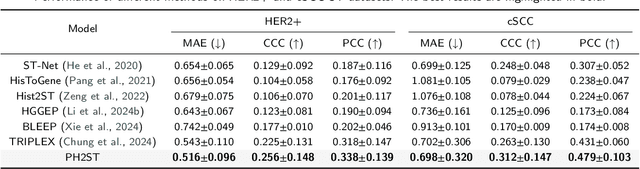


Abstract:Spatial Transcriptomics (ST) reveals the spatial distribution of gene expression in tissues, offering critical insights into biological processes and disease mechanisms. However, predicting ST from H\&E-stained histology images is challenging due to the heterogeneous relationship between histomorphology and gene expression, which arises from substantial variability across different patients and tissue sections. A more practical and valuable approach is to utilize ST data from a few local regions to predict the spatial transcriptomic landscape across the remaining regions in H&E slides. In response, we propose PHG2ST, an ST-prompt guided histological hypergraph learning framework, which leverages sparse ST signals as prompts to guide histological hypergraph learning for global spatial gene expression prediction. Our framework fuses histological hypergraph representations at multiple scales through a masked ST-prompt encoding mechanism, improving robustness and generalizability. Benchmark evaluations on two public ST datasets demonstrate that PHG2ST outperforms the existing state-of-the-art methods and closely aligns with the ground truth. These results underscore the potential of leveraging sparse local ST data for scalable and cost-effective spatial gene expression mapping in real-world biomedical applications.
General Vision Encoder Features as Guidance in Medical Image Registration
Jul 18, 2024Abstract:General vision encoders like DINOv2 and SAM have recently transformed computer vision. Even though they are trained on natural images, such encoder models have excelled in medical imaging, e.g., in classification, segmentation, and registration. However, no in-depth comparison of different state-of-the-art general vision encoders for medical registration is available. In this work, we investigate how well general vision encoder features can be used in the dissimilarity metrics for medical image registration. We explore two encoders that were trained on natural images as well as one that was fine-tuned on medical data. We apply the features within the well-established B-spline FFD registration framework. In extensive experiments on cardiac cine MRI data, we find that using features as additional guidance for conventional metrics improves the registration quality. The code is available at github.com/compai-lab/2024-miccai-koegl.
GPU optimization of the 3D Scale-invariant Feature Transform Algorithm and a Novel BRIEF-inspired 3D Fast Descriptor
Dec 19, 2021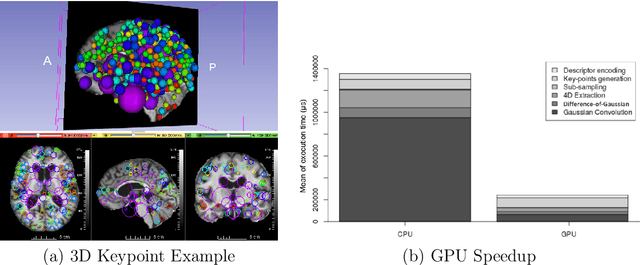
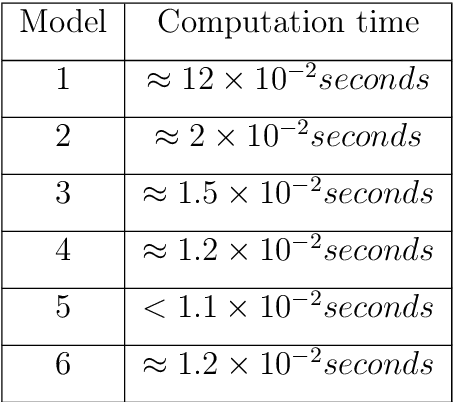
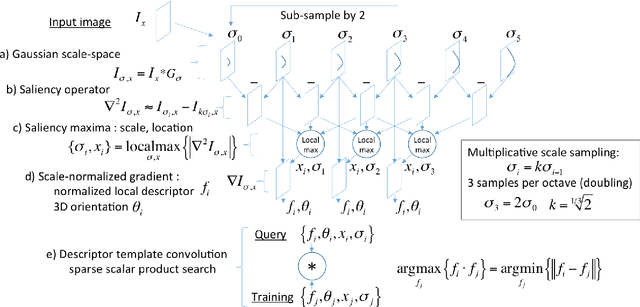
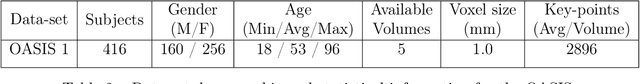
Abstract:This work details a highly efficient implementation of the 3D scale-invariant feature transform (SIFT) algorithm, for the purpose of machine learning from large sets of volumetric medical image data. The primary operations of the 3D SIFT code are implemented on a graphics processing unit (GPU), including convolution, sub-sampling, and 4D peak detection from scale-space pyramids. The performance improvements are quantified in keypoint detection and image-to-image matching experiments, using 3D MRI human brain volumes of different people. Computationally efficient 3D keypoint descriptors are proposed based on the Binary Robust Independent Elementary Feature (BRIEF) code, including a novel descriptor we call Ranked Robust Independent Elementary Features (RRIEF), and compared to the original 3D SIFT-Rank method\citep{toews2013efficient}. The GPU implementation affords a speedup of approximately 7X beyond an optimised CPU implementation, where computation time is reduced from 1.4 seconds to 0.2 seconds for 3D volumes of size (145, 174, 145) voxels with approximately 3000 keypoints. Notable speedups include the convolution operation (20X), 4D peak detection (3X), sub-sampling (3X), and difference-of-Gaussian pyramid construction (2X). Efficient descriptors offer a speedup of 2X and a memory savings of 6X compared to standard SIFT-Rank descriptors, at a cost of reduced numbers of keypoint correspondences, revealing a trade-off between computational efficiency and algorithmic performance. The speedups gained by our implementation will allow for a more efficient analysis on larger data sets. Our optimized GPU implementation of the 3D SIFT-Rank extractor is available at https://github.com/CarluerJB/3D_SIFT_CUDA.
The Impact of Domain Shift on Left and Right Ventricle Segmentation in Short Axis Cardiac MR Images
Sep 22, 2021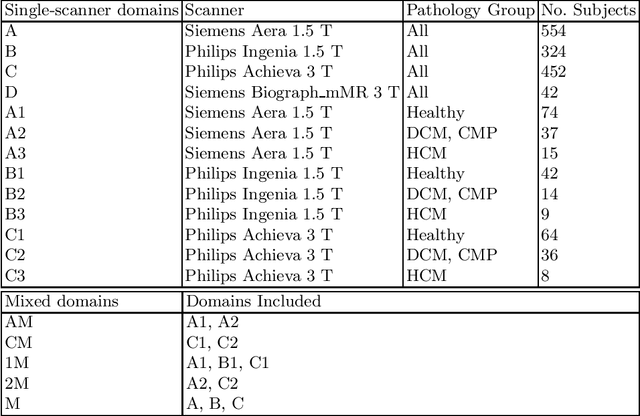
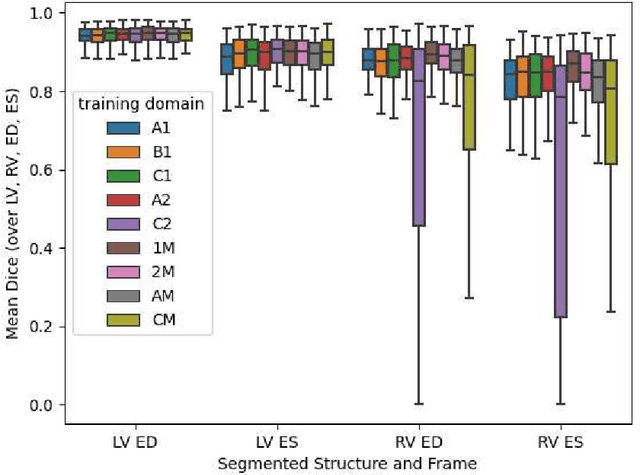
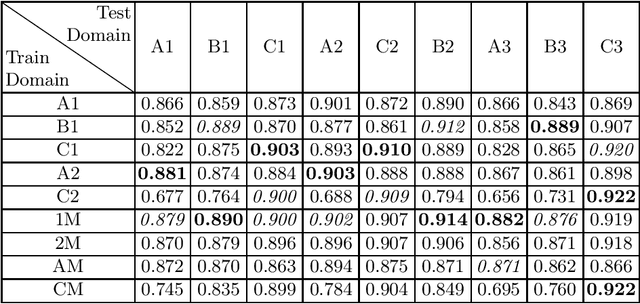

Abstract:Domain shift refers to the difference in the data distribution of two datasets, normally between the training set and the test set for machine learning algorithms. Domain shift is a serious problem for generalization of machine learning models and it is well-established that a domain shift between the training and test sets may cause a drastic drop in the model's performance. In medical imaging, there can be many sources of domain shift such as different scanners or scan protocols, different pathologies in the patient population, anatomical differences in the patient population (e.g. men vs women) etc. Therefore, in order to train models that have good generalization performance, it is important to be aware of the domain shift problem, its potential causes and to devise ways to address it. In this paper, we study the effect of domain shift on left and right ventricle blood pool segmentation in short axis cardiac MR images. Our dataset contains short axis images from 4 different MR scanners and 3 different pathology groups. The training is performed with nnUNet. The results show that scanner differences cause a greater drop in performance compared to changing the pathology group, and that the impact of domain shift is greater on right ventricle segmentation compared to left ventricle segmentation. Increasing the number of training subjects increased cross-scanner performance more than in-scanner performance at small training set sizes, but this difference in improvement decreased with larger training set sizes. Training models using data from multiple scanners improved cross-domain performance.
Quality-aware Cine Cardiac MRI Reconstruction and Analysis from Undersampled k-space Data
Sep 16, 2021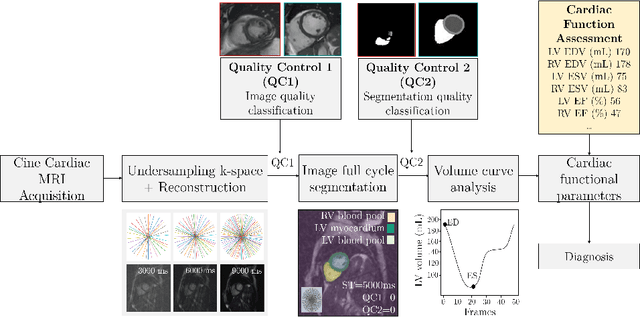
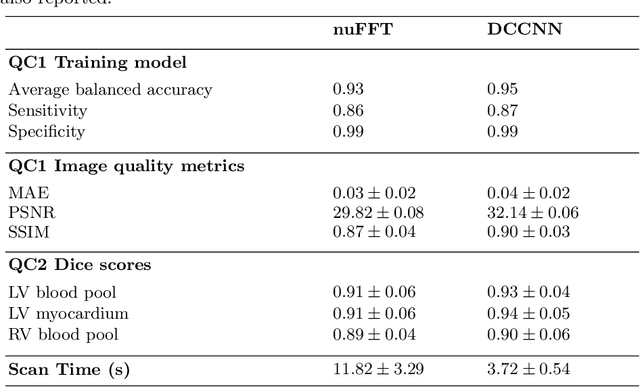
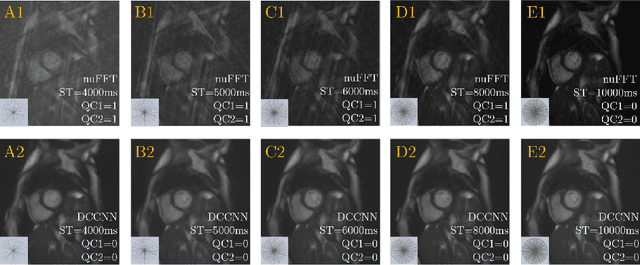
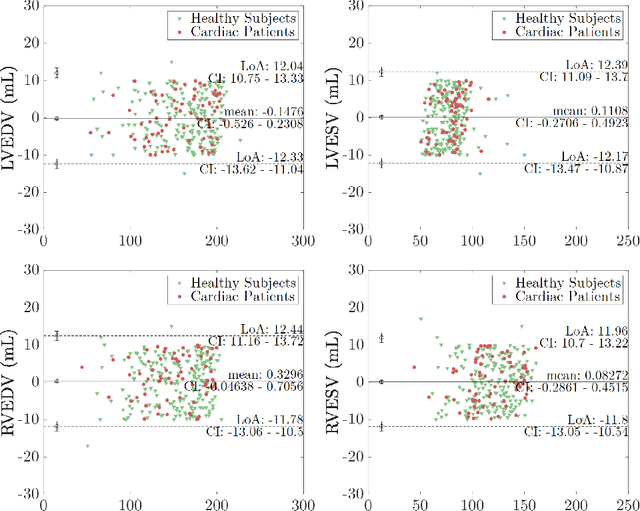
Abstract:Cine cardiac MRI is routinely acquired for the assessment of cardiac health, but the imaging process is slow and typically requires several breath-holds to acquire sufficient k-space profiles to ensure good image quality. Several undersampling-based reconstruction techniques have been proposed during the last decades to speed up cine cardiac MRI acquisition. However, the undersampling factor is commonly fixed to conservative values before acquisition to ensure diagnostic image quality, potentially leading to unnecessarily long scan times. In this paper, we propose an end-to-end quality-aware cine short-axis cardiac MRI framework that combines image acquisition and reconstruction with downstream tasks such as segmentation, volume curve analysis and estimation of cardiac functional parameters. The goal is to reduce scan time by acquiring only a fraction of k-space data to enable the reconstruction of images that can pass quality control checks and produce reliable estimates of cardiac functional parameters. The framework consists of a deep learning model for the reconstruction of 2D+t cardiac cine MRI images from undersampled data, an image quality-control step to detect good quality reconstructions, followed by a deep learning model for bi-ventricular segmentation, a quality-control step to detect good quality segmentations and automated calculation of cardiac functional parameters. To demonstrate the feasibility of the proposed approach, we perform simulations using a cohort of selected participants from the UK Biobank (n=270), 200 healthy subjects and 70 patients with cardiomyopathies. Our results show that we can produce quality-controlled images in a scan time reduced from 12 to 4 seconds per slice, enabling reliable estimates of cardiac functional parameters such as ejection fraction within 5% mean absolute error.
A Feature-Driven Active Framework for Ultrasound-Based Brain Shift Compensation
Mar 20, 2018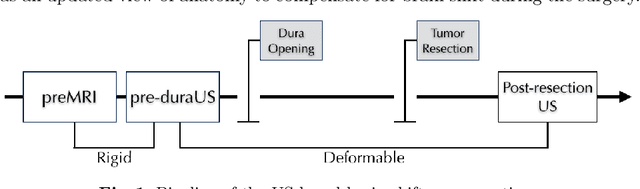
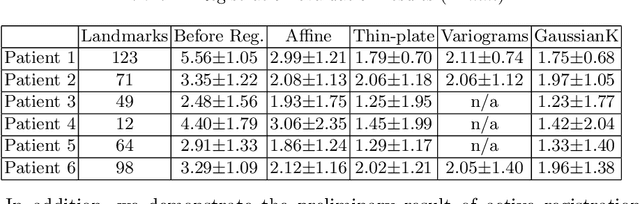
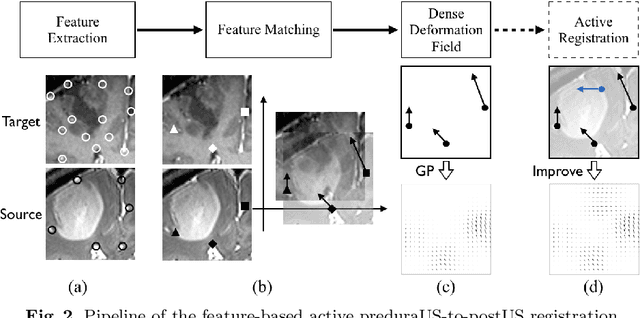
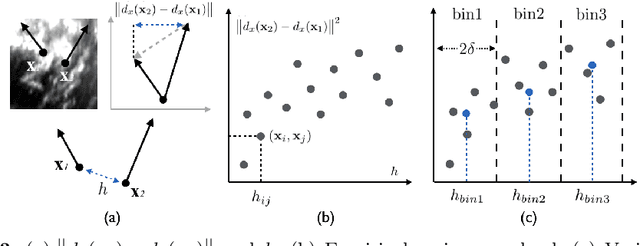
Abstract:A reliable Ultrasound (US)-to-US registration method to compensate for brain shift would substantially improve Image-Guided Neurological Surgery. Developing such a registration method is very challenging, due to factors such as missing correspondence in images, the complexity of brain pathology and the demand for fast computation. We propose a novel feature-driven active framework. Here, landmarks and their displacement are first estimated from a pair of US images using corresponding local image features. Subsequently, a Gaussian Process (GP) model is used to interpolate a dense deformation field from the sparse landmarks. Kernels of the GP are estimated by using variograms and a discrete grid search method. If necessary, the user can actively add new landmarks based on the image context and visualization of the uncertainty measure provided by the GP to further improve the result. We retrospectively demonstrate our registration framework as a robust and accurate brain shift compensation solution on clinical data acquired during neurosurgery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge