Chaoran Cheng
M3: High-fidelity Text-to-Image Generation via Multi-Modal, Multi-Agent and Multi-Round Visual Reasoning
Feb 05, 2026Abstract:Generative models have achieved impressive fidelity in text-to-image synthesis, yet struggle with complex compositional prompts involving multiple constraints. We introduce \textbf{M3 (Multi-Modal, Multi-Agent, Multi-Round)}, a training-free framework that systematically resolves these failures through iterative inference-time refinement. M3 orchestrates off-the-shelf foundation models in a robust multi-agent loop: a Planner decomposes prompts into verifiable checklists, while specialized Checker, Refiner, and Editor agents surgically correct constraints one at a time, with a Verifier ensuring monotonic improvement. Applied to open-source models, M3 achieves remarkable results on the challenging OneIG-EN benchmark, with our Qwen-Image+M3 surpassing commercial flagship systems including Imagen4 (0.515) and Seedream 3.0 (0.530), reaching state-of-the-art performance (0.532 overall). This demonstrates that intelligent multi-agent reasoning can elevate open-source models beyond proprietary alternatives. M3 also substantially improves GenEval compositional metrics, effectively doubling spatial reasoning performance on hardened test sets. As a plug-and-play module compatible with any pre-trained T2I model, M3 establishes a new paradigm for compositional generation without costly retraining.
$α$-Flow: A Unified Framework for Continuous-State Discrete Flow Matching Models
Apr 14, 2025Abstract:Recent efforts have extended the flow-matching framework to discrete generative modeling. One strand of models directly works with the continuous probabilities instead of discrete tokens, which we colloquially refer to as Continuous-State Discrete Flow Matching (CS-DFM). Existing CS-DFM models differ significantly in their representations and geometric assumptions. This work presents a unified framework for CS-DFM models, under which the existing variants can be understood as operating on different $\alpha$-representations of probabilities. Building upon the theory of information geometry, we introduce $\alpha$-Flow, a family of CS-DFM models that adheres to the canonical $\alpha$-geometry of the statistical manifold, and demonstrate its optimality in minimizing the generalized kinetic energy. Theoretically, we show that the flow matching loss for $\alpha$-flow establishes a unified variational bound for the discrete negative log-likelihood. We comprehensively evaluate different instantiations of $\alpha$-flow on various discrete generation domains to demonstrate their effectiveness in discrete generative modeling, including intermediate values whose geometries have never been explored before. $\alpha$-flow significantly outperforms its discrete-state counterpart in image and protein sequence generation and better captures the entropy in language modeling.
Online Reward-Weighted Fine-Tuning of Flow Matching with Wasserstein Regularization
Feb 09, 2025



Abstract:Recent advancements in reinforcement learning (RL) have achieved great success in fine-tuning diffusion-based generative models. However, fine-tuning continuous flow-based generative models to align with arbitrary user-defined reward functions remains challenging, particularly due to issues such as policy collapse from overoptimization and the prohibitively high computational cost of likelihoods in continuous-time flows. In this paper, we propose an easy-to-use and theoretically sound RL fine-tuning method, which we term Online Reward-Weighted Conditional Flow Matching with Wasserstein-2 Regularization (ORW-CFM-W2). Our method integrates RL into the flow matching framework to fine-tune generative models with arbitrary reward functions, without relying on gradients of rewards or filtered datasets. By introducing an online reward-weighting mechanism, our approach guides the model to prioritize high-reward regions in the data manifold. To prevent policy collapse and maintain diversity, we incorporate Wasserstein-2 (W2) distance regularization into our method and derive a tractable upper bound for it in flow matching, effectively balancing exploration and exploitation of policy optimization. We provide theoretical analyses to demonstrate the convergence properties and induced data distributions of our method, establishing connections with traditional RL algorithms featuring Kullback-Leibler (KL) regularization and offering a more comprehensive understanding of the underlying mechanisms and learning behavior of our approach. Extensive experiments on tasks including target image generation, image compression, and text-image alignment demonstrate the effectiveness of our method, where our method achieves optimal policy convergence while allowing controllable trade-offs between reward maximization and diversity preservation.
Hard Constraint Guided Flow Matching for Gradient-Free Generation of PDE Solutions
Dec 02, 2024Abstract:Generative models that satisfy hard constraints are crucial in many scientific and engineering applications where physical laws or system requirements must be strictly respected. However, many existing constrained generative models, especially those developed for computer vision, rely heavily on gradient information, often sparse or computationally expensive in fields like partial differential equations (PDEs). In this work, we introduce a novel framework for adapting pre-trained, unconstrained flow-matching models to satisfy constraints exactly in a zero-shot manner without requiring expensive gradient computations or fine-tuning. Our framework, ECI sampling, alternates between extrapolation (E), correction (C), and interpolation (I) stages during each iterative sampling step of flow matching sampling to ensure accurate integration of constraint information while preserving the validity of the generation. We demonstrate the effectiveness of our approach across various PDE systems, showing that ECI-guided generation strictly adheres to physical constraints and accurately captures complex distribution shifts induced by these constraints. Empirical results demonstrate that our framework consistently outperforms baseline approaches in various zero-shot constrained generation tasks and also achieves competitive results in the regression tasks without additional fine-tuning.
Hotspot-Driven Peptide Design via Multi-Fragment Autoregressive Extension
Nov 26, 2024



Abstract:Peptides, short chains of amino acids, interact with target proteins, making them a unique class of protein-based therapeutics for treating human diseases. Recently, deep generative models have shown great promise in peptide generation. However, several challenges remain in designing effective peptide binders. First, not all residues contribute equally to peptide-target interactions. Second, the generated peptides must adopt valid geometries due to the constraints of peptide bonds. Third, realistic tasks for peptide drug development are still lacking. To address these challenges, we introduce PepHAR, a hot-spot-driven autoregressive generative model for designing peptides targeting specific proteins. Building on the observation that certain hot spot residues have higher interaction potentials, we first use an energy-based density model to fit and sample these key residues. Next, to ensure proper peptide geometry, we autoregressively extend peptide fragments by estimating dihedral angles between residue frames. Finally, we apply an optimization process to iteratively refine fragment assembly, ensuring correct peptide structures. By combining hot spot sampling with fragment-based extension, our approach enables de novo peptide design tailored to a target protein and allows the incorporation of key hot spot residues into peptide scaffolds. Extensive experiments, including peptide design and peptide scaffold generation, demonstrate the strong potential of PepHAR in computational peptide binder design.
Geometric Point Attention Transformer for 3D Shape Reassembly
Nov 26, 2024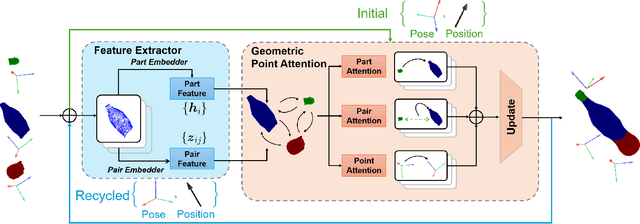

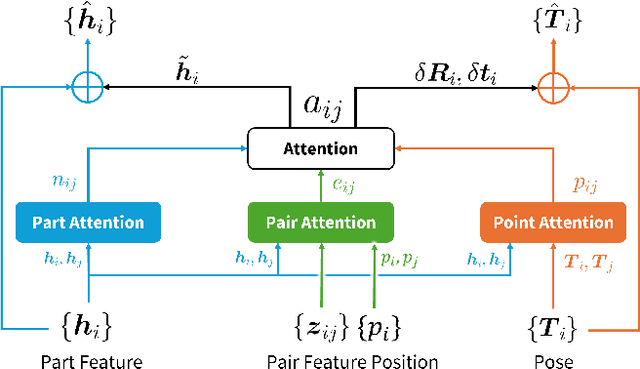

Abstract:Shape assembly, which aims to reassemble separate parts into a complete object, has gained significant interest in recent years. Existing methods primarily rely on networks to predict the poses of individual parts, but often fail to effectively capture the geometric interactions between the parts and their poses. In this paper, we present the Geometric Point Attention Transformer (GPAT), a network specifically designed to address the challenges of reasoning about geometric relationships. In the geometric point attention module, we integrate both global shape information and local pairwise geometric features, along with poses represented as rotation and translation vectors for each part. To enable iterative updates and dynamic reasoning, we introduce a geometric recycling scheme, where each prediction is fed into the next iteration for refinement. We evaluate our model on both the semantic and geometric assembly tasks, showing that it outperforms previous methods in absolute pose estimation, achieving accurate pose predictions and high alignment accuracy.
Training Free Guided Flow Matching with Optimal Control
Oct 23, 2024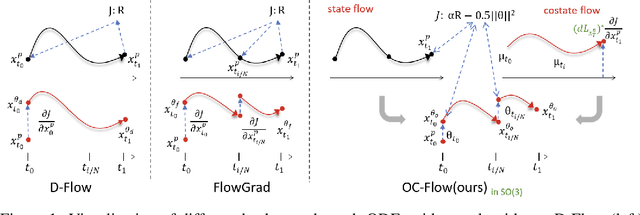
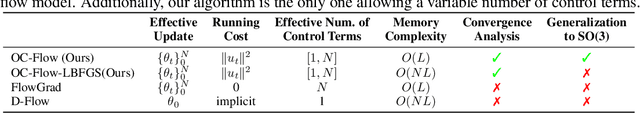


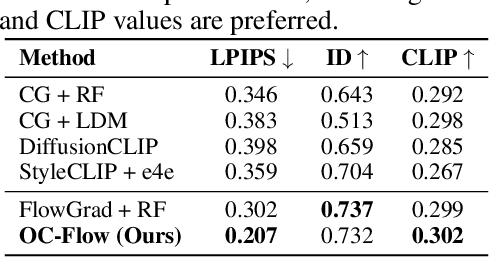
Abstract:Controlled generation with pre-trained Diffusion and Flow Matching models has vast applications. One strategy for guiding ODE-based generative models is through optimizing a target loss $R(x_1)$ while staying close to the prior distribution. Along this line, some recent work showed the effectiveness of guiding flow model by differentiating through its ODE sampling process. Despite the superior performance, the theoretical understanding of this line of methods is still preliminary, leaving space for algorithm improvement. Moreover, existing methods predominately focus on Euclidean data manifold, and there is a compelling need for guided flow methods on complex geometries such as SO(3), which prevails in high-stake scientific applications like protein design. We present OC-Flow, a general and theoretically grounded training-free framework for guided flow matching using optimal control. Building upon advances in optimal control theory, we develop effective and practical algorithms for solving optimal control in guided ODE-based generation and provide a systematic theoretical analysis of the convergence guarantee in both Euclidean and SO(3). We show that existing backprop-through-ODE methods can be interpreted as special cases of Euclidean OC-Flow. OC-Flow achieved superior performance in extensive experiments on text-guided image manipulation, conditional molecule generation, and all-atom peptide design.
Neural P$^3$M: A Long-Range Interaction Modeling Enhancer for Geometric GNNs
Sep 26, 2024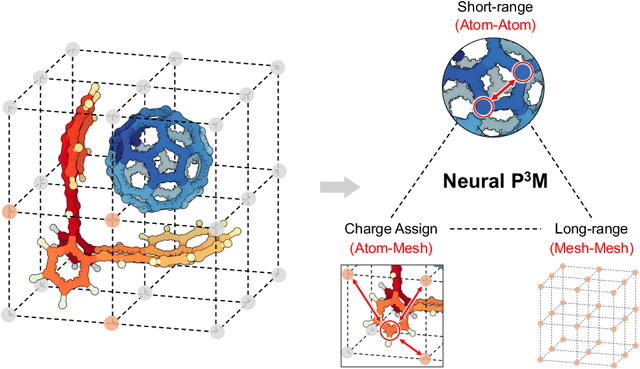
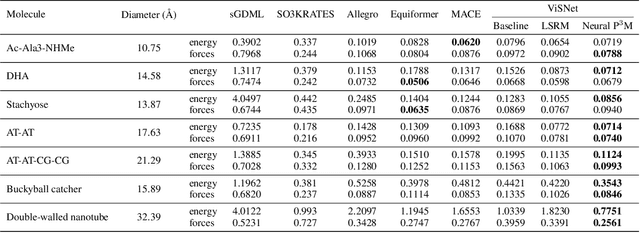
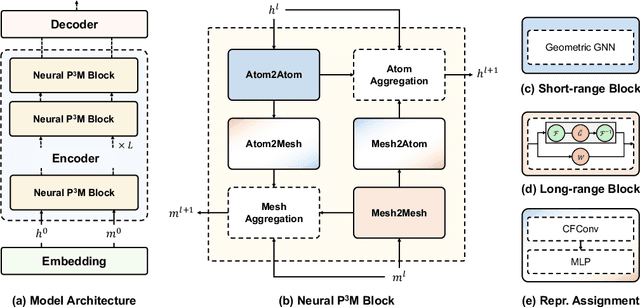
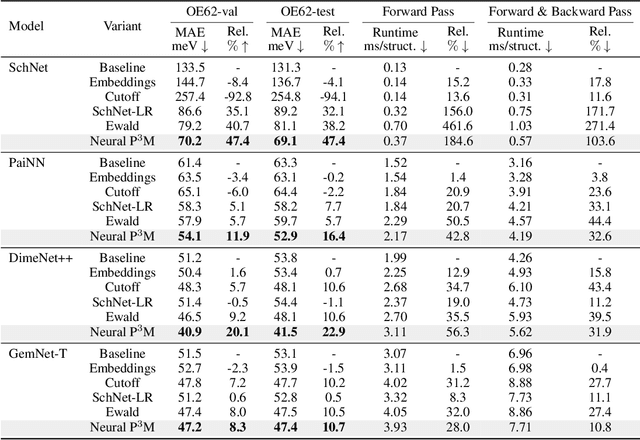
Abstract:Geometric graph neural networks (GNNs) have emerged as powerful tools for modeling molecular geometry. However, they encounter limitations in effectively capturing long-range interactions in large molecular systems. To address this challenge, we introduce Neural P$^3$M, a versatile enhancer of geometric GNNs to expand the scope of their capabilities by incorporating mesh points alongside atoms and reimaging traditional mathematical operations in a trainable manner. Neural P$^3$M exhibits flexibility across a wide range of molecular systems and demonstrates remarkable accuracy in predicting energies and forces, outperforming on benchmarks such as the MD22 dataset. It also achieves an average improvement of 22% on the OE62 dataset while integrating with various architectures.
Full-Atom Peptide Design based on Multi-modal Flow Matching
Jun 02, 2024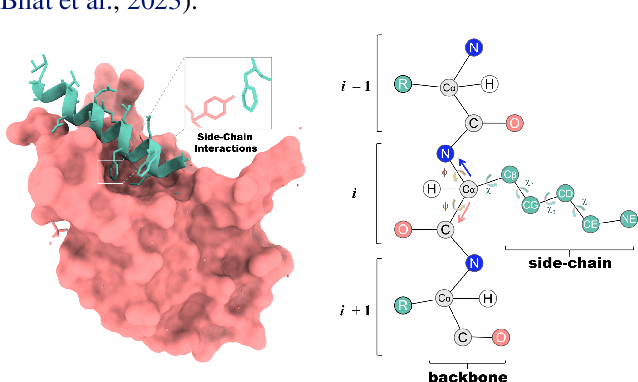
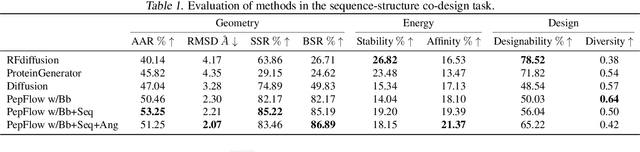
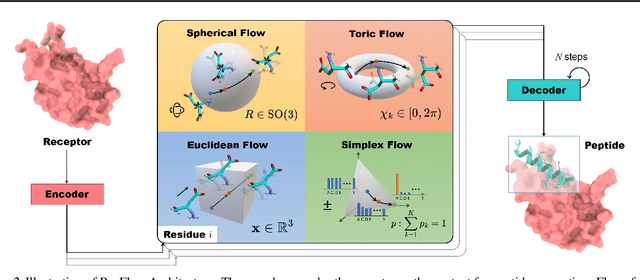
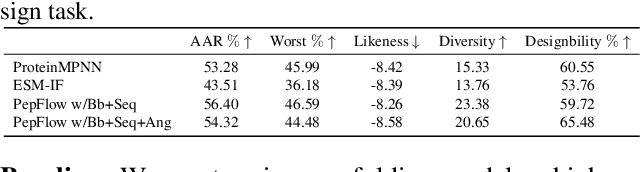
Abstract:Peptides, short chains of amino acid residues, play a vital role in numerous biological processes by interacting with other target molecules, offering substantial potential in drug discovery. In this work, we present PepFlow, the first multi-modal deep generative model grounded in the flow-matching framework for the design of full-atom peptides that target specific protein receptors. Drawing inspiration from the crucial roles of residue backbone orientations and side-chain dynamics in protein-peptide interactions, we characterize the peptide structure using rigid backbone frames within the $\mathrm{SE}(3)$ manifold and side-chain angles on high-dimensional tori. Furthermore, we represent discrete residue types in the peptide sequence as categorical distributions on the probability simplex. By learning the joint distributions of each modality using derived flows and vector fields on corresponding manifolds, our method excels in the fine-grained design of full-atom peptides. Harnessing the multi-modal paradigm, our approach adeptly tackles various tasks such as fix-backbone sequence design and side-chain packing through partial sampling. Through meticulously crafted experiments, we demonstrate that PepFlow exhibits superior performance in comprehensive benchmarks, highlighting its significant potential in computational peptide design and analysis.
Categorical Flow Matching on Statistical Manifolds
May 26, 2024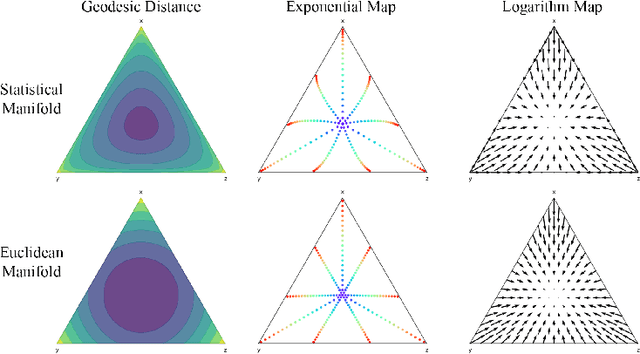
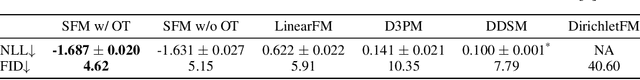
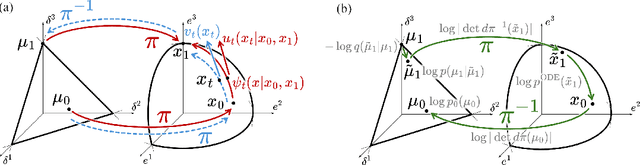
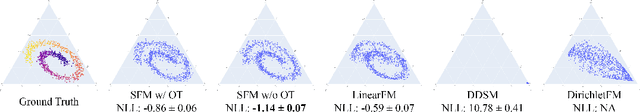
Abstract:We introduce Statistical Flow Matching (SFM), a novel and mathematically rigorous flow-matching framework on the manifold of parameterized probability measures inspired by the results from information geometry. We demonstrate the effectiveness of our method on the discrete generation problem by instantiating SFM on the manifold of categorical distributions whose geometric properties remain unexplored in previous discrete generative models. Utilizing the Fisher information metric, we equip the manifold with a Riemannian structure whose intrinsic geometries are effectively leveraged by following the shortest paths of geodesics. We develop an efficient training and sampling algorithm that overcomes numerical stability issues with a diffeomorphism between manifolds. Our distinctive geometric perspective of statistical manifolds allows us to apply optimal transport during training and interpret SFM as following the steepest direction of the natural gradient. Unlike previous models that rely on variational bounds for likelihood estimation, SFM enjoys the exact likelihood calculation for arbitrary probability measures. We manifest that SFM can learn more complex patterns on the statistical manifold where existing models often fail due to strong prior assumptions. Comprehensive experiments on real-world generative tasks ranging from image, text to biological domains further demonstrate that SFM achieves higher sampling quality and likelihood than other discrete diffusion or flow-based models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge