Satoshi Kasai
Comparative validation of surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation in endoscopy: Results of the PhaKIR 2024 challenge
Jul 22, 2025Abstract:Reliable recognition and localization of surgical instruments in endoscopic video recordings are foundational for a wide range of applications in computer- and robot-assisted minimally invasive surgery (RAMIS), including surgical training, skill assessment, and autonomous assistance. However, robust performance under real-world conditions remains a significant challenge. Incorporating surgical context - such as the current procedural phase - has emerged as a promising strategy to improve robustness and interpretability. To address these challenges, we organized the Surgical Procedure Phase, Keypoint, and Instrument Recognition (PhaKIR) sub-challenge as part of the Endoscopic Vision (EndoVis) challenge at MICCAI 2024. We introduced a novel, multi-center dataset comprising thirteen full-length laparoscopic cholecystectomy videos collected from three distinct medical institutions, with unified annotations for three interrelated tasks: surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation. Unlike existing datasets, ours enables joint investigation of instrument localization and procedural context within the same data while supporting the integration of temporal information across entire procedures. We report results and findings in accordance with the BIAS guidelines for biomedical image analysis challenges. The PhaKIR sub-challenge advances the field by providing a unique benchmark for developing temporally aware, context-driven methods in RAMIS and offers a high-quality resource to support future research in surgical scene understanding.
crossMoDA Challenge: Evolution of Cross-Modality Domain Adaptation Techniques for Vestibular Schwannoma and Cochlea Segmentation from 2021 to 2023
Jun 13, 2025Abstract:The cross-Modality Domain Adaptation (crossMoDA) challenge series, initiated in 2021 in conjunction with the International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI), focuses on unsupervised cross-modality segmentation, learning from contrast-enhanced T1 (ceT1) and transferring to T2 MRI. The task is an extreme example of domain shift chosen to serve as a meaningful and illustrative benchmark. From a clinical application perspective, it aims to automate Vestibular Schwannoma (VS) and cochlea segmentation on T2 scans for more cost-effective VS management. Over time, the challenge objectives have evolved to enhance its clinical relevance. The challenge evolved from using single-institutional data and basic segmentation in 2021 to incorporating multi-institutional data and Koos grading in 2022, and by 2023, it included heterogeneous routine data and sub-segmentation of intra- and extra-meatal tumour components. In this work, we report the findings of the 2022 and 2023 editions and perform a retrospective analysis of the challenge progression over the years. The observations from the successive challenge contributions indicate that the number of outliers decreases with an expanding dataset. This is notable since the diversity of scanning protocols of the datasets concurrently increased. The winning approach of the 2023 edition reduced the number of outliers on the 2021 and 2022 testing data, demonstrating how increased data heterogeneity can enhance segmentation performance even on homogeneous data. However, the cochlea Dice score declined in 2023, likely due to the added complexity from tumour sub-annotations affecting overall segmentation performance. While progress is still needed for clinically acceptable VS segmentation, the plateauing performance suggests that a more challenging cross-modal task may better serve future benchmarking.
PitVis-2023 Challenge: Workflow Recognition in videos of Endoscopic Pituitary Surgery
Sep 02, 2024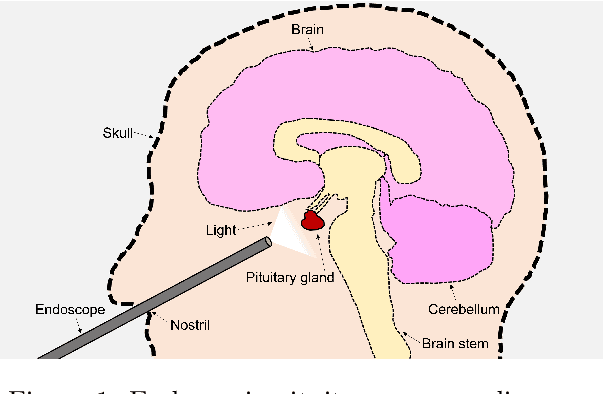
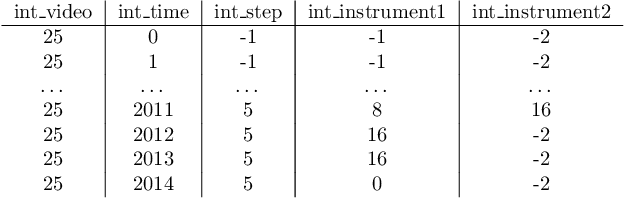

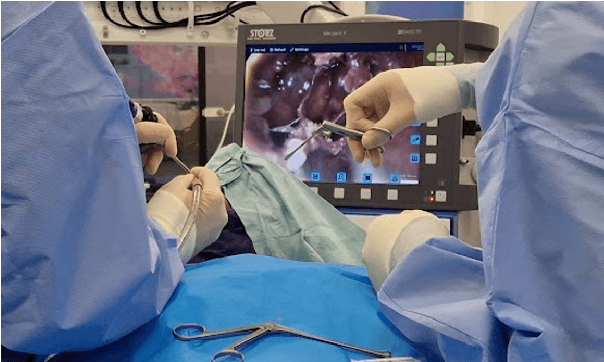
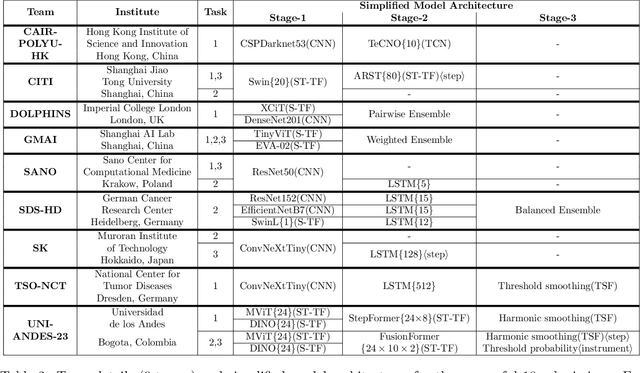
Abstract:The field of computer vision applied to videos of minimally invasive surgery is ever-growing. Workflow recognition pertains to the automated recognition of various aspects of a surgery: including which surgical steps are performed; and which surgical instruments are used. This information can later be used to assist clinicians when learning the surgery; during live surgery; and when writing operation notes. The Pituitary Vision (PitVis) 2023 Challenge tasks the community to step and instrument recognition in videos of endoscopic pituitary surgery. This is a unique task when compared to other minimally invasive surgeries due to the smaller working space, which limits and distorts vision; and higher frequency of instrument and step switching, which requires more precise model predictions. Participants were provided with 25-videos, with results presented at the MICCAI-2023 conference as part of the Endoscopic Vision 2023 Challenge in Vancouver, Canada, on 08-Oct-2023. There were 18-submissions from 9-teams across 6-countries, using a variety of deep learning models. A commonality between the top performing models was incorporating spatio-temporal and multi-task methods, with greater than 50% and 10% macro-F1-score improvement over purely spacial single-task models in step and instrument recognition respectively. The PitVis-2023 Challenge therefore demonstrates state-of-the-art computer vision models in minimally invasive surgery are transferable to a new dataset, with surgery specific techniques used to enhance performance, progressing the field further. Benchmark results are provided in the paper, and the dataset is publicly available at: https://doi.org/10.5522/04/26531686.
LNQ 2023 challenge: Benchmark of weakly-supervised techniques for mediastinal lymph node quantification
Aug 19, 2024



Abstract:Accurate assessment of lymph node size in 3D CT scans is crucial for cancer staging, therapeutic management, and monitoring treatment response. Existing state-of-the-art segmentation frameworks in medical imaging often rely on fully annotated datasets. However, for lymph node segmentation, these datasets are typically small due to the extensive time and expertise required to annotate the numerous lymph nodes in 3D CT scans. Weakly-supervised learning, which leverages incomplete or noisy annotations, has recently gained interest in the medical imaging community as a potential solution. Despite the variety of weakly-supervised techniques proposed, most have been validated only on private datasets or small publicly available datasets. To address this limitation, the Mediastinal Lymph Node Quantification (LNQ) challenge was organized in conjunction with the 26th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2023). This challenge aimed to advance weakly-supervised segmentation methods by providing a new, partially annotated dataset and a robust evaluation framework. A total of 16 teams from 5 countries submitted predictions to the validation leaderboard, and 6 teams from 3 countries participated in the evaluation phase. The results highlighted both the potential and the current limitations of weakly-supervised approaches. On one hand, weakly-supervised approaches obtained relatively good performance with a median Dice score of $61.0\%$. On the other hand, top-ranked teams, with a median Dice score exceeding $70\%$, boosted their performance by leveraging smaller but fully annotated datasets to combine weak supervision and full supervision. This highlights both the promise of weakly-supervised methods and the ongoing need for high-quality, fully annotated data to achieve higher segmentation performance.
SAR-RARP50: Segmentation of surgical instrumentation and Action Recognition on Robot-Assisted Radical Prostatectomy Challenge
Dec 31, 2023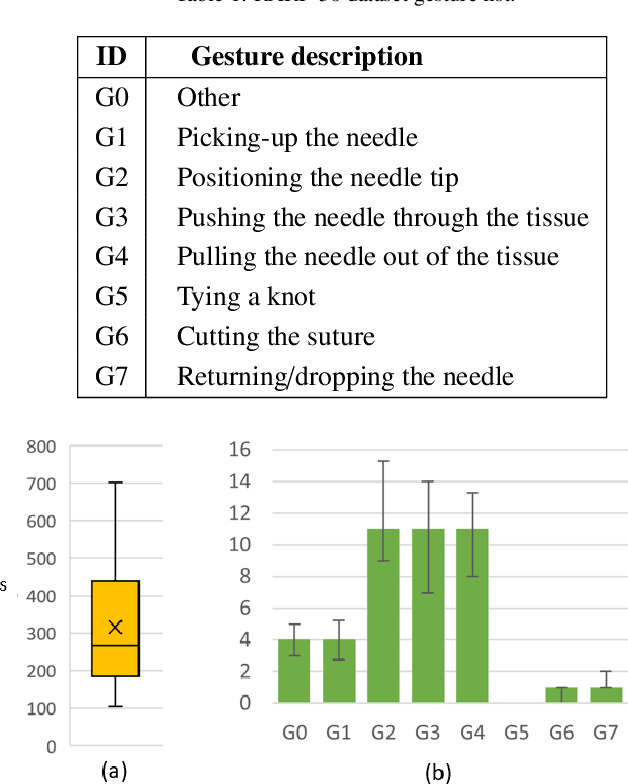
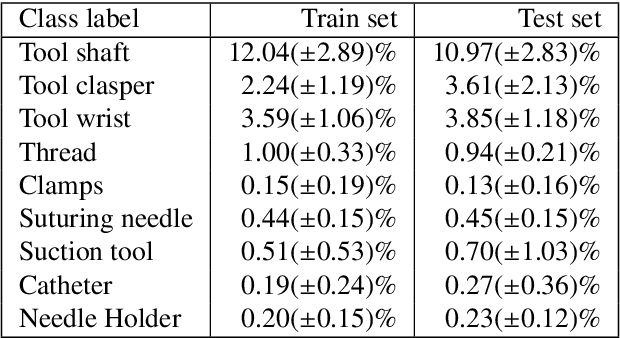
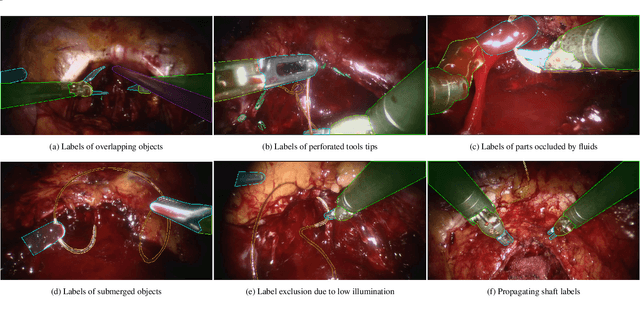
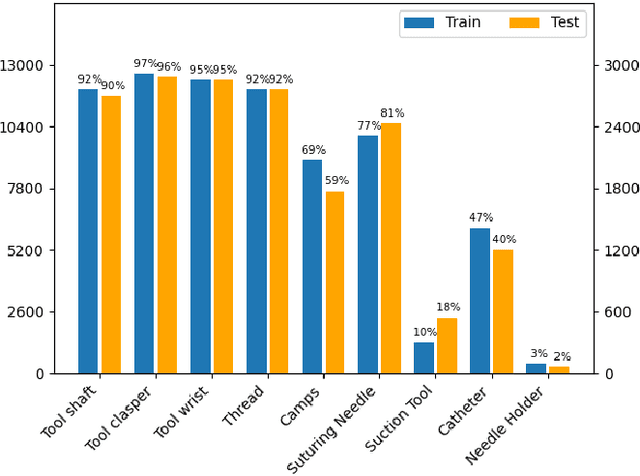
Abstract:Surgical tool segmentation and action recognition are fundamental building blocks in many computer-assisted intervention applications, ranging from surgical skills assessment to decision support systems. Nowadays, learning-based action recognition and segmentation approaches outperform classical methods, relying, however, on large, annotated datasets. Furthermore, action recognition and tool segmentation algorithms are often trained and make predictions in isolation from each other, without exploiting potential cross-task relationships. With the EndoVis 2022 SAR-RARP50 challenge, we release the first multimodal, publicly available, in-vivo, dataset for surgical action recognition and semantic instrumentation segmentation, containing 50 suturing video segments of Robotic Assisted Radical Prostatectomy (RARP). The aim of the challenge is twofold. First, to enable researchers to leverage the scale of the provided dataset and develop robust and highly accurate single-task action recognition and tool segmentation approaches in the surgical domain. Second, to further explore the potential of multitask-based learning approaches and determine their comparative advantage against their single-task counterparts. A total of 12 teams participated in the challenge, contributing 7 action recognition methods, 9 instrument segmentation techniques, and 4 multitask approaches that integrated both action recognition and instrument segmentation.
Domain generalization across tumor types, laboratories, and species -- insights from the 2022 edition of the Mitosis Domain Generalization Challenge
Sep 27, 2023



Abstract:Recognition of mitotic figures in histologic tumor specimens is highly relevant to patient outcome assessment. This task is challenging for algorithms and human experts alike, with deterioration of algorithmic performance under shifts in image representations. Considerable covariate shifts occur when assessment is performed on different tumor types, images are acquired using different digitization devices, or specimens are produced in different laboratories. This observation motivated the inception of the 2022 challenge on MItosis Domain Generalization (MIDOG 2022). The challenge provided annotated histologic tumor images from six different domains and evaluated the algorithmic approaches for mitotic figure detection provided by nine challenge participants on ten independent domains. Ground truth for mitotic figure detection was established in two ways: a three-expert consensus and an independent, immunohistochemistry-assisted set of labels. This work represents an overview of the challenge tasks, the algorithmic strategies employed by the participants, and potential factors contributing to their success. With an $F_1$ score of 0.764 for the top-performing team, we summarize that domain generalization across various tumor domains is possible with today's deep learning-based recognition pipelines. When assessed against the immunohistochemistry-assisted reference standard, all methods resulted in reduced recall scores, but with only minor changes in the order of participants in the ranking.
Synthesizing 3D computed tomography from MRI or CBCT using 2.5D deep neural networks
Aug 23, 2023



Abstract:Deep learning techniques, particularly convolutional neural networks (CNNs), have gained traction for synthetic computed tomography (sCT) generation from Magnetic resonance imaging (MRI), Cone-beam computed tomography (CBCT) and PET. In this report, we introduce a method to syn-thesize CT from MRI or CBCT. Our method is based on multi-slice (2.5D) CNNs. 2.5D CNNs offer distinct advantages over 3D CNNs when dealing with volumetric data. In the experiments, we evaluate the performance of our method for two tasks, MRI-to-sCT and CBCT-to-sCT generation. Target organs for both tasks are brain and pelvis.
The ACROBAT 2022 Challenge: Automatic Registration Of Breast Cancer Tissue
May 29, 2023Abstract:The alignment of tissue between histopathological whole-slide-images (WSI) is crucial for research and clinical applications. Advances in computing, deep learning, and availability of large WSI datasets have revolutionised WSI analysis. Therefore, the current state-of-the-art in WSI registration is unclear. To address this, we conducted the ACROBAT challenge, based on the largest WSI registration dataset to date, including 4,212 WSIs from 1,152 breast cancer patients. The challenge objective was to align WSIs of tissue that was stained with routine diagnostic immunohistochemistry to its H&E-stained counterpart. We compare the performance of eight WSI registration algorithms, including an investigation of the impact of different WSI properties and clinical covariates. We find that conceptually distinct WSI registration methods can lead to highly accurate registration performances and identify covariates that impact performances across methods. These results establish the current state-of-the-art in WSI registration and guide researchers in selecting and developing methods.
Surgical tool classification and localization: results and methods from the MICCAI 2022 SurgToolLoc challenge
May 11, 2023



Abstract:The ability to automatically detect and track surgical instruments in endoscopic videos can enable transformational interventions. Assessing surgical performance and efficiency, identifying skilled tool use and choreography, and planning operational and logistical aspects of OR resources are just a few of the applications that could benefit. Unfortunately, obtaining the annotations needed to train machine learning models to identify and localize surgical tools is a difficult task. Annotating bounding boxes frame-by-frame is tedious and time-consuming, yet large amounts of data with a wide variety of surgical tools and surgeries must be captured for robust training. Moreover, ongoing annotator training is needed to stay up to date with surgical instrument innovation. In robotic-assisted surgery, however, potentially informative data like timestamps of instrument installation and removal can be programmatically harvested. The ability to rely on tool installation data alone would significantly reduce the workload to train robust tool-tracking models. With this motivation in mind we invited the surgical data science community to participate in the challenge, SurgToolLoc 2022. The goal was to leverage tool presence data as weak labels for machine learning models trained to detect tools and localize them in video frames with bounding boxes. We present the results of this challenge along with many of the team's efforts. We conclude by discussing these results in the broader context of machine learning and surgical data science. The training data used for this challenge consisting of 24,695 video clips with tool presence labels is also being released publicly and can be accessed at https://console.cloud.google.com/storage/browser/isi-surgtoolloc-2022.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge