Yanbo Feng
Fair Canonical Correlation Analysis
Sep 27, 2023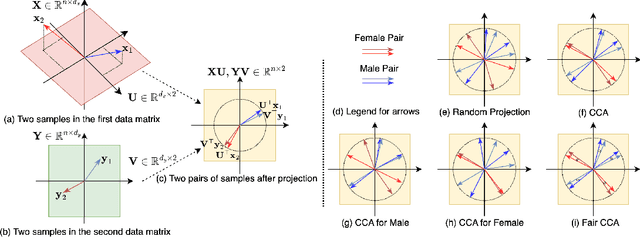
Abstract:This paper investigates fairness and bias in Canonical Correlation Analysis (CCA), a widely used statistical technique for examining the relationship between two sets of variables. We present a framework that alleviates unfairness by minimizing the correlation disparity error associated with protected attributes. Our approach enables CCA to learn global projection matrices from all data points while ensuring that these matrices yield comparable correlation levels to group-specific projection matrices. Experimental evaluation on both synthetic and real-world datasets demonstrates the efficacy of our method in reducing correlation disparity error without compromising CCA accuracy.
The ACROBAT 2022 Challenge: Automatic Registration Of Breast Cancer Tissue
May 29, 2023Abstract:The alignment of tissue between histopathological whole-slide-images (WSI) is crucial for research and clinical applications. Advances in computing, deep learning, and availability of large WSI datasets have revolutionised WSI analysis. Therefore, the current state-of-the-art in WSI registration is unclear. To address this, we conducted the ACROBAT challenge, based on the largest WSI registration dataset to date, including 4,212 WSIs from 1,152 breast cancer patients. The challenge objective was to align WSIs of tissue that was stained with routine diagnostic immunohistochemistry to its H&E-stained counterpart. We compare the performance of eight WSI registration algorithms, including an investigation of the impact of different WSI properties and clinical covariates. We find that conceptually distinct WSI registration methods can lead to highly accurate registration performances and identify covariates that impact performances across methods. These results establish the current state-of-the-art in WSI registration and guide researchers in selecting and developing methods.
ACROBAT -- a multi-stain breast cancer histological whole-slide-image data set from routine diagnostics for computational pathology
Nov 24, 2022Abstract:The analysis of FFPE tissue sections stained with haematoxylin and eosin (H&E) or immunohistochemistry (IHC) is an essential part of the pathologic assessment of surgically resected breast cancer specimens. IHC staining has been broadly adopted into diagnostic guidelines and routine workflows to manually assess status and scoring of several established biomarkers, including ER, PGR, HER2 and KI67. However, this is a task that can also be facilitated by computational pathology image analysis methods. The research in computational pathology has recently made numerous substantial advances, often based on publicly available whole slide image (WSI) data sets. However, the field is still considerably limited by the sparsity of public data sets. In particular, there are no large, high quality publicly available data sets with WSIs of matching IHC and H&E-stained tissue sections. Here, we publish the currently largest publicly available data set of WSIs of tissue sections from surgical resection specimens from female primary breast cancer patients with matched WSIs of corresponding H&E and IHC-stained tissue, consisting of 4,212 WSIs from 1,153 patients. The primary purpose of the data set was to facilitate the ACROBAT WSI registration challenge, aiming at accurately aligning H&E and IHC images. For research in the area of image registration, automatic quantitative feedback on registration algorithm performance remains available through the ACROBAT challenge website, based on more than 37,000 manually annotated landmark pairs from 13 annotators. Beyond registration, this data set has the potential to enable many different avenues of computational pathology research, including stain-guided learning, virtual staining, unsupervised pre-training, artefact detection and stain-independent models.
A deep learning based multiscale approach to segment cancer area in liver whole slide image
Jul 25, 2020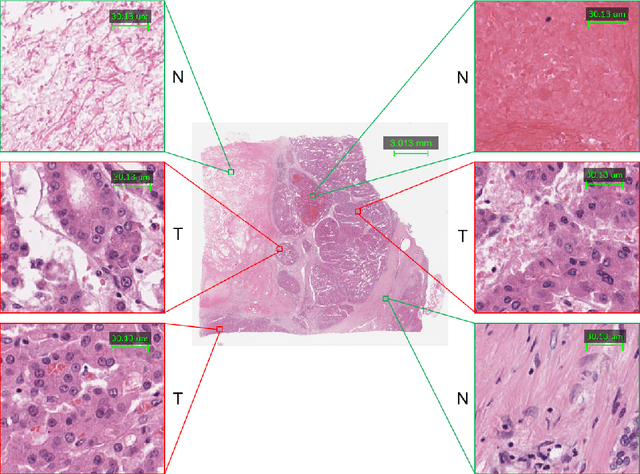
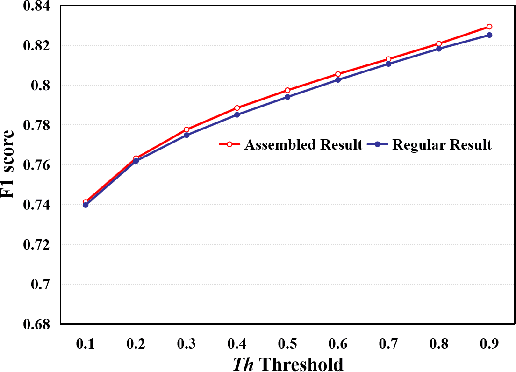
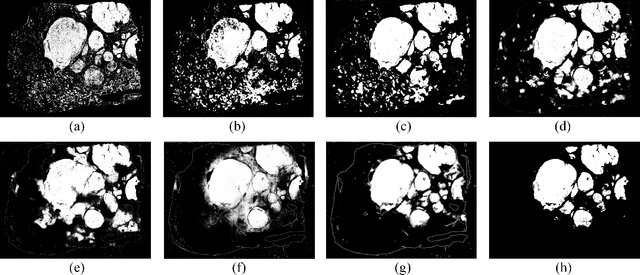
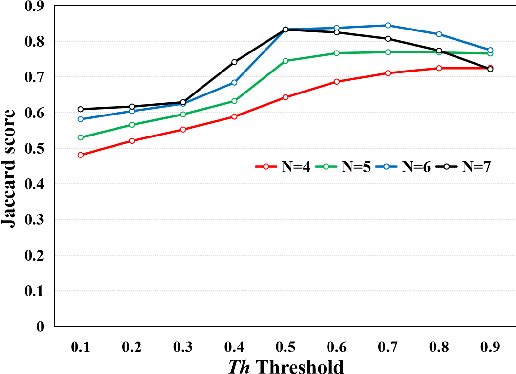
Abstract:This paper addresses the problem of liver cancer segmentation in Whole Slide Image (WSI). We propose a multi-scale image processing method based on automatic end-to-end deep neural network algorithm for segmentation of cancer area. A seven-levels gaussian pyramid representation of the histopathological image was built to provide the texture information in different scales. In this work, several neural architectures were compared using the original image level for the training procedure. The proposed method is based on U-Net applied to seven levels of various resolutions (pyramidal subsumpling). The predictions in different levels are combined through a voting mechanism. The final segmentation result is generated at the original image level. Partial color normalization and weighted overlapping method were applied in preprocessing and prediction separately. The results show the effectiveness of the proposed multi-scales approach achieving better scores compared to the state-of-the-art.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge