Markus Eckstein
Tissue Classification and Whole-Slide Images Analysis via Modeling of the Tumor Microenvironment and Biological Pathways
Jan 13, 2026Abstract:Automatic integration of whole slide images (WSIs) and gene expression profiles has demonstrated substantial potential in precision clinical diagnosis and cancer progression studies. However, most existing studies focus on individual gene sequences and slide level classification tasks, with limited attention to spatial transcriptomics and patch level applications. To address this limitation, we propose a multimodal network, BioMorphNet, which automatically integrates tissue morphological features and spatial gene expression to support tissue classification and differential gene analysis. For considering morphological features, BioMorphNet constructs a graph to model the relationships between target patches and their neighbors, and adjusts the response strength based on morphological and molecular level similarity, to better characterize the tumor microenvironment. In terms of multimodal interactions, BioMorphNet derives clinical pathway features from spatial transcriptomic data based on a predefined pathway database, serving as a bridge between tissue morphology and gene expression. In addition, a novel learnable pathway module is designed to automatically simulate the biological pathway formation process, providing a complementary representation to existing clinical pathways. Compared with the latest morphology gene multimodal methods, BioMorphNet's average classification metrics improve by 2.67%, 5.48%, and 6.29% for prostate cancer, colorectal cancer, and breast cancer datasets, respectively. BioMorphNet not only classifies tissue categories within WSIs accurately to support tumor localization, but also analyzes differential gene expression between tissue categories based on prediction confidence, contributing to the discovery of potential tumor biomarkers.
Spatial Transcriptomics Expression Prediction from Histopathology Based on Cross-Modal Mask Reconstruction and Contrastive Learning
Jun 10, 2025Abstract:Spatial transcriptomics is a technology that captures gene expression levels at different spatial locations, widely used in tumor microenvironment analysis and molecular profiling of histopathology, providing valuable insights into resolving gene expression and clinical diagnosis of cancer. Due to the high cost of data acquisition, large-scale spatial transcriptomics data remain challenging to obtain. In this study, we develop a contrastive learning-based deep learning method to predict spatially resolved gene expression from whole-slide images. Evaluation across six different disease datasets demonstrates that, compared to existing studies, our method improves Pearson Correlation Coefficient (PCC) in the prediction of highly expressed genes, highly variable genes, and marker genes by 6.27%, 6.11%, and 11.26% respectively. Further analysis indicates that our method preserves gene-gene correlations and applies to datasets with limited samples. Additionally, our method exhibits potential in cancer tissue localization based on biomarker expression.
Domain generalization across tumor types, laboratories, and species -- insights from the 2022 edition of the Mitosis Domain Generalization Challenge
Sep 27, 2023



Abstract:Recognition of mitotic figures in histologic tumor specimens is highly relevant to patient outcome assessment. This task is challenging for algorithms and human experts alike, with deterioration of algorithmic performance under shifts in image representations. Considerable covariate shifts occur when assessment is performed on different tumor types, images are acquired using different digitization devices, or specimens are produced in different laboratories. This observation motivated the inception of the 2022 challenge on MItosis Domain Generalization (MIDOG 2022). The challenge provided annotated histologic tumor images from six different domains and evaluated the algorithmic approaches for mitotic figure detection provided by nine challenge participants on ten independent domains. Ground truth for mitotic figure detection was established in two ways: a three-expert consensus and an independent, immunohistochemistry-assisted set of labels. This work represents an overview of the challenge tasks, the algorithmic strategies employed by the participants, and potential factors contributing to their success. With an $F_1$ score of 0.764 for the top-performing team, we summarize that domain generalization across various tumor domains is possible with today's deep learning-based recognition pipelines. When assessed against the immunohistochemistry-assisted reference standard, all methods resulted in reduced recall scores, but with only minor changes in the order of participants in the ranking.
Fast whole-slide cartography in colon cancer histology using superpixels and CNN classification
Jun 30, 2021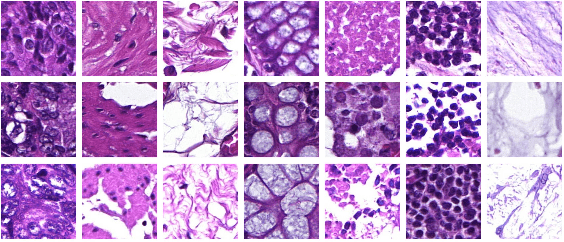
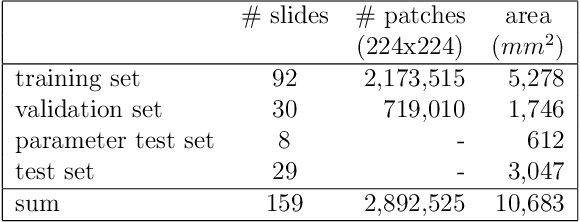
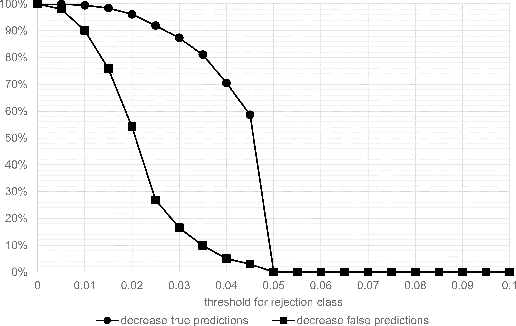
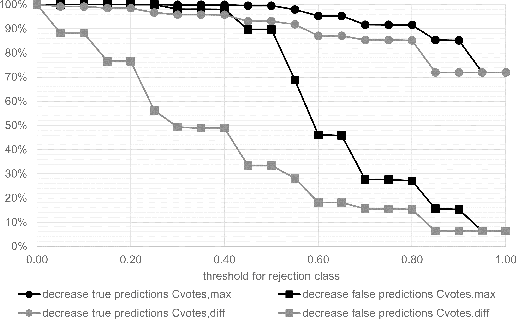
Abstract:Whole-slide-image cartography is the process of automatically detecting and outlining different tissue types in digitized histological specimen. This semantic segmentation provides a basis for many follow-up analyses and can potentially guide subsequent medical decisions. Due to their large size, whole-slide-images typically have to be divided into smaller patches which are then analyzed individually using machine learning-based approaches. Thereby, local dependencies of image regions get lost and since a whole-slide-image comprises many thousands of such patches this process is inherently slow. We propose to subdivide the image into coherent regions prior to classification by grouping visually similar adjacent image pixels into larger segments, i.e. superpixels. Afterwards, only a random subset of patches per superpixel is classified and patch labels are combined into a single superpixel label. The algorithm has been developed and validated on a dataset of 159 hand-annotated whole-slide-images of colon resections and its performance has been compared to a standard patch-based approach. The algorithm shows an average speed-up of 41% on the test data and the overall accuracy is increased from 93.8% to 95.7%. We additionally propose a metric for identifying superpixels with an uncertain classification so they can be excluded from further analysis. Finally, we evaluate two potential medical applications, namely tumor area estimation including tumor invasive margin generation and tumor composition analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge