Jonas Ammeling
SWAN -- Enabling Fast and Mobile Histopathology Image Annotation through Swipeable Interfaces
Nov 11, 2025Abstract:The annotation of large scale histopathology image datasets remains a major bottleneck in developing robust deep learning models for clinically relevant tasks, such as mitotic figure classification. Folder-based annotation workflows are usually slow, fatiguing, and difficult to scale. To address these challenges, we introduce SWipeable ANnotations (SWAN), an open-source, MIT-licensed web application that enables intuitive image patch classification using a swiping gesture. SWAN supports both desktop and mobile platforms, offers real-time metadata capture, and allows flexible mapping of swipe gestures to class labels. In a pilot study with four pathologists annotating 600 mitotic figure image patches, we compared SWAN against a traditional folder-sorting workflow. SWAN enabled rapid annotations with pairwise percent agreement ranging from 86.52% to 93.68% (Cohen's Kappa = 0.61-0.80), while for the folder-based method, the pairwise percent agreement ranged from 86.98% to 91.32% (Cohen's Kappa = 0.63-0.75) for the task of classifying atypical versus normal mitotic figures, demonstrating high consistency between annotators and comparable performance. Participants rated the tool as highly usable and appreciated the ability to annotate on mobile devices. These results suggest that SWAN can accelerate image annotation while maintaining annotation quality, offering a scalable and user-friendly alternative to conventional workflows.
Effortless Vision-Language Model Specialization in Histopathology without Annotation
Aug 11, 2025Abstract:Recent advances in Vision-Language Models (VLMs) in histopathology, such as CONCH and QuiltNet, have demonstrated impressive zero-shot classification capabilities across various tasks. However, their general-purpose design may lead to suboptimal performance in specific downstream applications. While supervised fine-tuning methods address this issue, they require manually labeled samples for adaptation. This paper investigates annotation-free adaptation of VLMs through continued pretraining on domain- and task-relevant image-caption pairs extracted from existing databases. Our experiments on two VLMs, CONCH and QuiltNet, across three downstream tasks reveal that these pairs substantially enhance both zero-shot and few-shot performance. Notably, with larger training sizes, continued pretraining matches the performance of few-shot methods while eliminating manual labeling. Its effectiveness, task-agnostic design, and annotation-free workflow make it a promising pathway for adapting VLMs to new histopathology tasks. Code is available at https://github.com/DeepMicroscopy/Annotation-free-VLM-specialization.
Benchmarking Deep Learning and Vision Foundation Models for Atypical vs. Normal Mitosis Classification with Cross-Dataset Evaluation
Jun 26, 2025Abstract:Atypical mitoses mark a deviation in the cell division process that can be an independent prognostically relevant marker for tumor malignancy. However, their identification remains challenging due to low prevalence, at times subtle morphological differences from normal mitoses, low inter-rater agreement among pathologists, and class imbalance in datasets. Building on the Atypical Mitosis dataset for Breast Cancer (AMi-Br), this study presents a comprehensive benchmark comparing deep learning approaches for automated atypical mitotic figure (AMF) classification, including baseline models, foundation models with linear probing, and foundation models fine-tuned with low-rank adaptation (LoRA). For rigorous evaluation, we further introduce two new hold-out AMF datasets - AtNorM-Br, a dataset of mitoses from the The TCGA breast cancer cohort, and AtNorM-MD, a multi-domain dataset of mitoses from the MIDOG++ training set. We found average balanced accuracy values of up to 0.8135, 0.7696, and 0.7705 on the in-domain AMi-Br and the out-of-domain AtNorm-Br and AtNorM-MD datasets, respectively, with the results being particularly good for LoRA-based adaptation of the Virchow-line of foundation models. Our work shows that atypical mitosis classification, while being a challenging problem, can be effectively addressed through the use of recent advances in transfer learning and model fine-tuning techniques. We make available all code and data used in this paper in this github repository: https://github.com/DeepMicroscopy/AMi-Br_Benchmark.
A Histologic Dataset of Normal and Atypical Mitotic Figures on Human Breast Cancer (AMi-Br)
Jan 08, 2025Abstract:Assessment of the density of mitotic figures (MFs) in histologic tumor sections is an important prognostic marker for many tumor types, including breast cancer. Recently, it has been reported in multiple works that the quantity of MFs with an atypical morphology (atypical MFs, AMFs) might be an independent prognostic criterion for breast cancer. AMFs are an indicator of mutations in the genes regulating the cell cycle and can lead to aberrant chromosome constitution (aneuploidy) of the tumor cells. To facilitate further research on this topic using pattern recognition, we present the first ever publicly available dataset of atypical and normal MFs (AMi-Br). For this, we utilized two of the most popular MF datasets (MIDOG 2021 and TUPAC) and subclassified all MFs using a three expert majority vote. Our final dataset consists of 3,720 MFs, split into 832 AMFs (22.4%) and 2,888 normal MFs (77.6%) across all 223 tumor cases in the combined set. We provide baseline classification experiments to investigate the consistency of the dataset, using a Monte Carlo cross-validation and different strategies to combat class imbalance. We found an averaged balanced accuracy of up to 0.806 when using a patch-level data set split, and up to 0.713 when using a patient-level split.
Is Self-Supervision Enough? Benchmarking Foundation Models Against End-to-End Training for Mitotic Figure Classification
Dec 09, 2024


Abstract:Foundation models (FMs), i.e., models trained on a vast amount of typically unlabeled data, have become popular and available recently for the domain of histopathology. The key idea is to extract semantically rich vectors from any input patch, allowing for the use of simple subsequent classification networks potentially reducing the required amounts of labeled data, and increasing domain robustness. In this work, we investigate to which degree this also holds for mitotic figure classification. Utilizing two popular public mitotic figure datasets, we compared linear probing of five publicly available FMs against models trained on ImageNet and a simple ResNet50 end-to-end-trained baseline. We found that the end-to-end-trained baseline outperformed all FM-based classifiers, regardless of the amount of data provided. Additionally, we did not observe the FM-based classifiers to be more robust against domain shifts, rendering both of the above assumptions incorrect.
On the Value of PHH3 for Mitotic Figure Detection on H&E-stained Images
Jun 28, 2024Abstract:The count of mitotic figures (MFs) observed in hematoxylin and eosin (H&E)-stained slides is an important prognostic marker as it is a measure for tumor cell proliferation. However, the identification of MFs has a known low inter-rater agreement. Deep learning algorithms can standardize this task, but they require large amounts of annotated data for training and validation. Furthermore, label noise introduced during the annotation process may impede the algorithm's performance. Unlike H&E, the mitosis-specific antibody phospho-histone H3 (PHH3) specifically highlights MFs. Counting MFs on slides stained against PHH3 leads to higher agreement among raters and has therefore recently been used as a ground truth for the annotation of MFs in H&E. However, as PHH3 facilitates the recognition of cells indistinguishable from H&E stain alone, the use of this ground truth could potentially introduce noise into the H&E-related dataset, impacting model performance. This study analyzes the impact of PHH3-assisted MF annotation on inter-rater reliability and object level agreement through an extensive multi-rater experiment. We found that the annotators' object-level agreement increased when using PHH3-assisted labeling. Subsequently, MF detectors were evaluated on the resulting datasets to investigate the influence of PHH3-assisted labeling on the models' performance. Additionally, a novel dual-stain MF detector was developed to investigate the interpretation-shift of PHH3-assisted labels used in H&E, which clearly outperformed single-stain detectors. However, the PHH3-assisted labels did not have a positive effect on solely H&E-based models. The high performance of our dual-input detector reveals an information mismatch between the H&E and PHH3-stained images as the cause of this effect.
Model-based Cleaning of the QUILT-1M Pathology Dataset for Text-Conditional Image Synthesis
Apr 11, 2024Abstract:The QUILT-1M dataset is the first openly available dataset containing images harvested from various online sources. While it provides a huge data variety, the image quality and composition is highly heterogeneous, impacting its utility for text-conditional image synthesis. We propose an automatic pipeline that provides predictions of the most common impurities within the images, e.g., visibility of narrators, desktop environment and pathology software, or text within the image. Additionally, we propose to use semantic alignment filtering of the image-text pairs. Our findings demonstrate that by rigorously filtering the dataset, there is a substantial enhancement of image fidelity in text-to-image tasks.
Re-identification from histopathology images
Mar 19, 2024



Abstract:In numerous studies, deep learning algorithms have proven their potential for the analysis of histopathology images, for example, for revealing the subtypes of tumors or the primary origin of metastases. These models require large datasets for training, which must be anonymized to prevent possible patient identity leaks. This study demonstrates that even relatively simple deep learning algorithms can re-identify patients in large histopathology datasets with substantial accuracy. We evaluated our algorithms on two TCIA datasets including lung squamous cell carcinoma (LSCC) and lung adenocarcinoma (LUAD). We also demonstrate the algorithm's performance on an in-house dataset of meningioma tissue. We predicted the source patient of a slide with F1 scores of 50.16 % and 52.30 % on the LSCC and LUAD datasets, respectively, and with 62.31 % on our meningioma dataset. Based on our findings, we formulated a risk assessment scheme to estimate the risk to the patient's privacy prior to publication.
Rethinking U-net Skip Connections for Biomedical Image Segmentation
Feb 13, 2024Abstract:The U-net architecture has significantly impacted deep learning-based segmentation of medical images. Through the integration of long-range skip connections, it facilitated the preservation of high-resolution features. Out-of-distribution data can, however, substantially impede the performance of neural networks. Previous works showed that the trained network layers differ in their susceptibility to this domain shift, e.g., shallow layers are more affected than deeper layers. In this work, we investigate the implications of this observation of layer sensitivity to domain shifts of U-net-style segmentation networks. By copying features of shallow layers to corresponding decoder blocks, these bear the risk of re-introducing domain-specific information. We used a synthetic dataset to model different levels of data distribution shifts and evaluated the impact on downstream segmentation performance. We quantified the inherent domain susceptibility of each network layer, using the Hellinger distance. These experiments confirmed the higher domain susceptibility of earlier network layers. When gradually removing skip connections, a decrease in domain susceptibility of deeper layers could be observed. For downstream segmentation performance, the original U-net outperformed the variant without any skip connections. The best performance, however, was achieved when removing the uppermost skip connection - not only in the presence of domain shifts but also for in-domain test data. We validated our results on three clinical datasets - two histopathology datasets and one magnetic resonance dataset - with performance increases of up to 10% in-domain and 13% cross-domain when removing the uppermost skip connection.
Automated Volume Corrected Mitotic Index Calculation Through Annotation-Free Deep Learning using Immunohistochemistry as Reference Standard
Nov 15, 2023
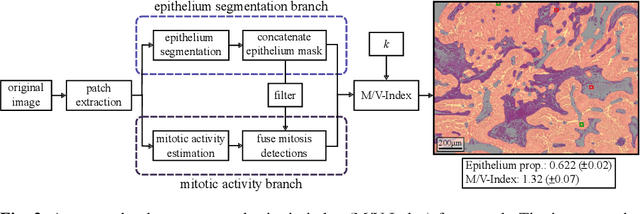
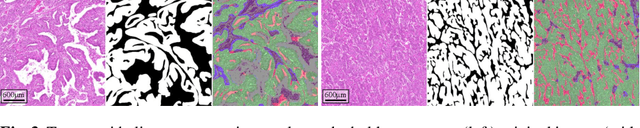
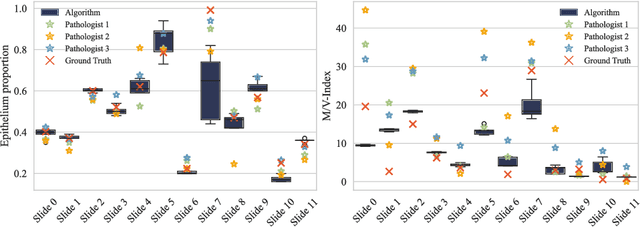
Abstract:The volume-corrected mitotic index (M/V-Index) was shown to provide prognostic value in invasive breast carcinomas. However, despite its prognostic significance, it is not established as the standard method for assessing aggressive biological behaviour, due to the high additional workload associated with determining the epithelial proportion. In this work, we show that using a deep learning pipeline solely trained with an annotation-free, immunohistochemistry-based approach, provides accurate estimations of epithelial segmentation in canine breast carcinomas. We compare our automatic framework with the manually annotated M/V-Index in a study with three board-certified pathologists. Our results indicate that the deep learning-based pipeline shows expert-level performance, while providing time efficiency and reproducibility.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge