Christian Marzahl
A bag of tricks for real-time Mitotic Figure detection
Aug 27, 2025Abstract:Mitotic figure (MF) detection in histopathology images is challenging due to large variations in slide scanners, staining protocols, tissue types, and the presence of artifacts. This paper presents a collection of training techniques - a bag of tricks - that enable robust, real-time MF detection across diverse domains. We build on the efficient RTMDet single stage object detector to achieve high inference speed suitable for clinical deployment. Our method addresses scanner variability and tumor heterogeneity via extensive multi-domain training data, balanced sampling, and careful augmentation. Additionally, we employ targeted, hard negative mining on necrotic and debris tissue to reduce false positives. In a grouped 5-fold cross-validation across multiple MF datasets, our model achieves an F1 score between 0.78 and 0.84. On the preliminary test set of the MItosis DOmain Generalization (MIDOG) 2025 challenge, our single-stage RTMDet-S based approach reaches an F1 of 0.81, outperforming larger models and demonstrating adaptability to new, unfamiliar domains. The proposed solution offers a practical trade-off between accuracy and speed, making it attractive for real-world clinical adoption.
On the Value of PHH3 for Mitotic Figure Detection on H&E-stained Images
Jun 28, 2024Abstract:The count of mitotic figures (MFs) observed in hematoxylin and eosin (H&E)-stained slides is an important prognostic marker as it is a measure for tumor cell proliferation. However, the identification of MFs has a known low inter-rater agreement. Deep learning algorithms can standardize this task, but they require large amounts of annotated data for training and validation. Furthermore, label noise introduced during the annotation process may impede the algorithm's performance. Unlike H&E, the mitosis-specific antibody phospho-histone H3 (PHH3) specifically highlights MFs. Counting MFs on slides stained against PHH3 leads to higher agreement among raters and has therefore recently been used as a ground truth for the annotation of MFs in H&E. However, as PHH3 facilitates the recognition of cells indistinguishable from H&E stain alone, the use of this ground truth could potentially introduce noise into the H&E-related dataset, impacting model performance. This study analyzes the impact of PHH3-assisted MF annotation on inter-rater reliability and object level agreement through an extensive multi-rater experiment. We found that the annotators' object-level agreement increased when using PHH3-assisted labeling. Subsequently, MF detectors were evaluated on the resulting datasets to investigate the influence of PHH3-assisted labeling on the models' performance. Additionally, a novel dual-stain MF detector was developed to investigate the interpretation-shift of PHH3-assisted labels used in H&E, which clearly outperformed single-stain detectors. However, the PHH3-assisted labels did not have a positive effect on solely H&E-based models. The high performance of our dual-input detector reveals an information mismatch between the H&E and PHH3-stained images as the cause of this effect.
The ACROBAT 2022 Challenge: Automatic Registration Of Breast Cancer Tissue
May 29, 2023Abstract:The alignment of tissue between histopathological whole-slide-images (WSI) is crucial for research and clinical applications. Advances in computing, deep learning, and availability of large WSI datasets have revolutionised WSI analysis. Therefore, the current state-of-the-art in WSI registration is unclear. To address this, we conducted the ACROBAT challenge, based on the largest WSI registration dataset to date, including 4,212 WSIs from 1,152 breast cancer patients. The challenge objective was to align WSIs of tissue that was stained with routine diagnostic immunohistochemistry to its H&E-stained counterpart. We compare the performance of eight WSI registration algorithms, including an investigation of the impact of different WSI properties and clinical covariates. We find that conceptually distinct WSI registration methods can lead to highly accurate registration performances and identify covariates that impact performances across methods. These results establish the current state-of-the-art in WSI registration and guide researchers in selecting and developing methods.
Mitosis domain generalization in histopathology images -- The MIDOG challenge
Apr 06, 2022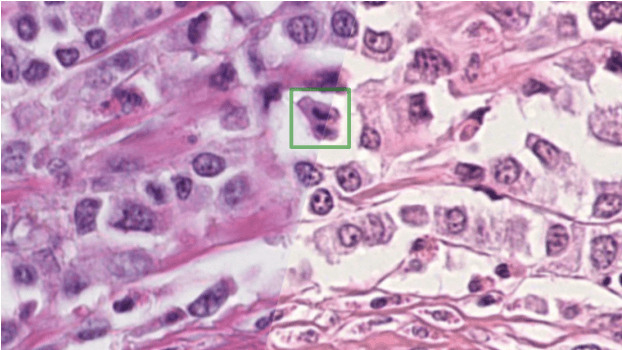
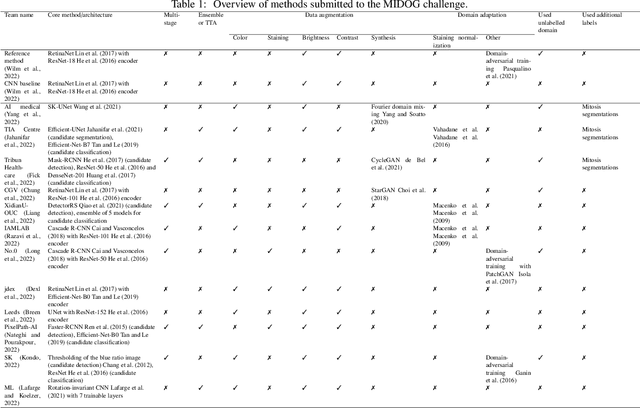
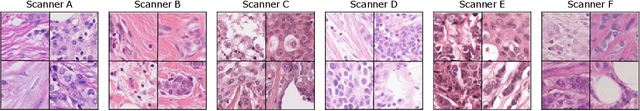
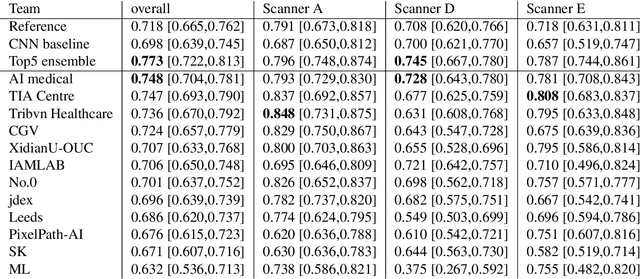
Abstract:The density of mitotic figures within tumor tissue is known to be highly correlated with tumor proliferation and thus is an important marker in tumor grading. Recognition of mitotic figures by pathologists is known to be subject to a strong inter-rater bias, which limits the prognostic value. State-of-the-art deep learning methods can support the expert in this assessment but are known to strongly deteriorate when applied in a different clinical environment than was used for training. One decisive component in the underlying domain shift has been identified as the variability caused by using different whole slide scanners. The goal of the MICCAI MIDOG 2021 challenge has been to propose and evaluate methods that counter this domain shift and derive scanner-agnostic mitosis detection algorithms. The challenge used a training set of 200 cases, split across four scanning systems. As a test set, an additional 100 cases split across four scanning systems, including two previously unseen scanners, were given. The best approaches performed on an expert level, with the winning algorithm yielding an F_1 score of 0.748 (CI95: 0.704-0.781). In this paper, we evaluate and compare the approaches that were submitted to the challenge and identify methodological factors contributing to better performance.
Pan-Tumor CAnine cuTaneous Cancer Histology (CATCH) Dataset
Jan 27, 2022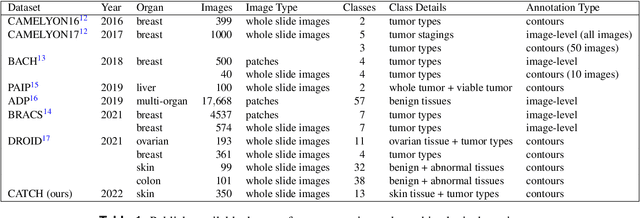
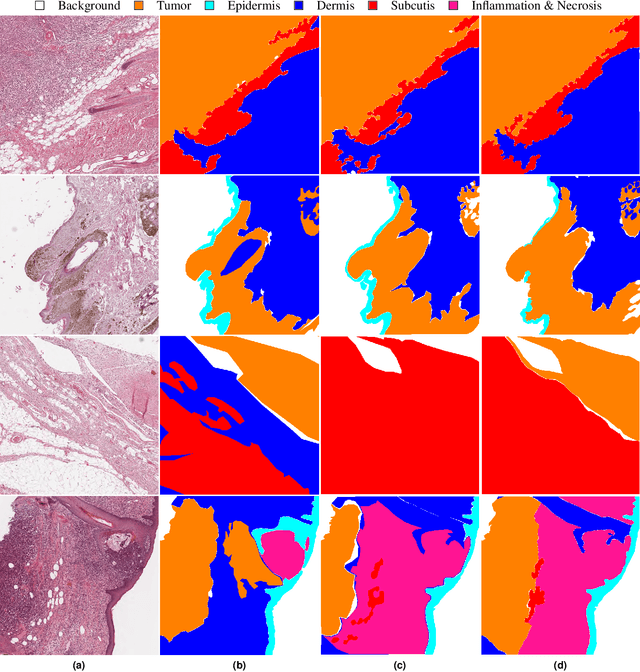
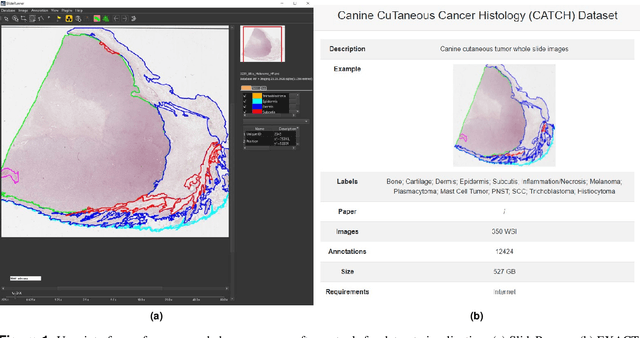
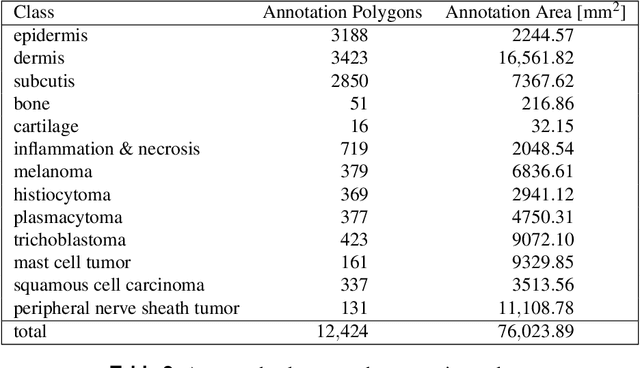

Abstract:Due to morphological similarities, the differentiation of histologic sections of cutaneous tumors into individual subtypes can be challenging. Recently, deep learning-based approaches have proven their potential for supporting pathologists in this regard. However, many of these supervised algorithms require a large amount of annotated data for robust development. We present a publicly available dataset consisting of 350 whole slide images of seven different canine cutaneous tumors complemented by 12,424 polygon annotations for 13 histologic classes including seven cutaneous tumor subtypes. Regarding sample size and annotation extent, this exceeds most publicly available datasets which are oftentimes limited to the tumor area or merely provide patch-level annotations. We validated our model for tissue segmentation, achieving a class-averaged Jaccard coefficient of 0.7047, and 0.9044 for tumor in particular. For tumor subtype classification, we achieve a slide-level accuracy of 0.9857. Since canine cutaneous tumors possess various histologic homologies to human tumors, we believe that the added value of this dataset is not limited to veterinary pathology but extends to more general fields of application.
Superpixel Pre-Segmentation of HER2 Slides for Efficient Annotation
Jan 19, 2022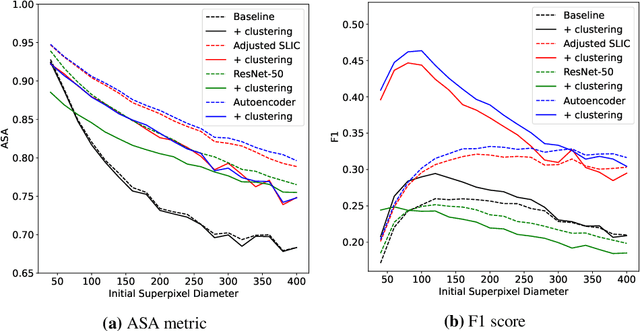
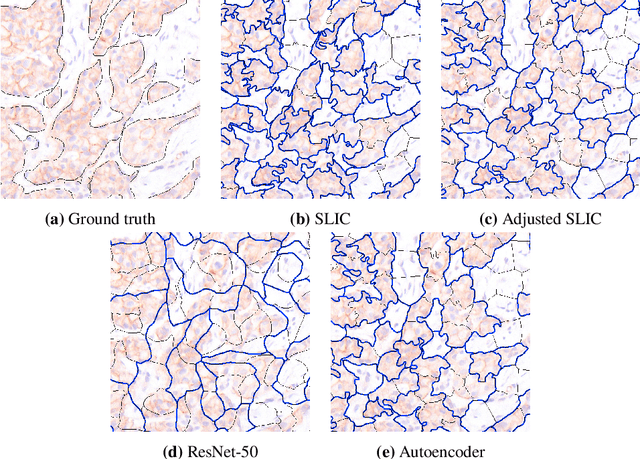
Abstract:Supervised deep learning has shown state-of-the-art performance for medical image segmentation across different applications, including histopathology and cancer research; however, the manual annotation of such data is extremely laborious. In this work, we explore the use of superpixel approaches to compute a pre-segmentation of HER2 stained images for breast cancer diagnosis that facilitates faster manual annotation and correction in a second step. Four methods are compared: Standard Simple Linear Iterative Clustering (SLIC) as a baseline, a domain adapted SLIC, and superpixels based on feature embeddings of a pretrained ResNet-50 and a denoising autoencoder. To tackle oversegmentation, we propose to hierarchically merge superpixels, based on their content in the respective feature space. When evaluating the approaches on fully manually annotated images, we observe that the autoencoder-based superpixels achieve a 23% increase in boundary F1 score compared to the baseline SLIC superpixels. Furthermore, the boundary F1 score increases by 73% when hierarchical clustering is applied on the adapted SLIC and the autoencoder-based superpixels. These evaluations show encouraging first results for a pre-segmentation for efficient manual refinement without the need for an initial set of annotated training data.
First steps on Gamification of Lung Fluid Cells Annotations in the Flower Domain
Nov 05, 2021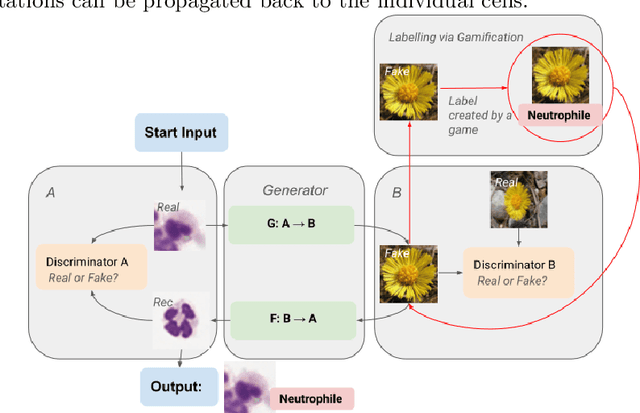

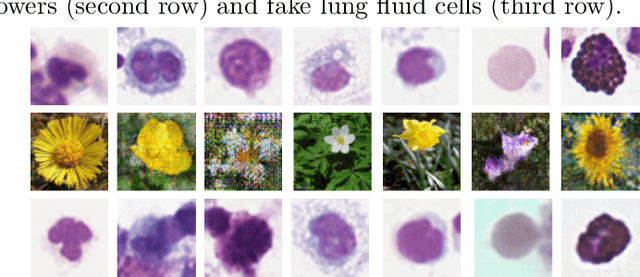
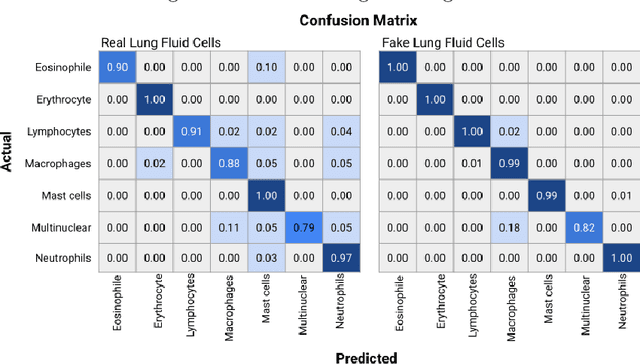
Abstract:Annotating data, especially in the medical domain, requires expert knowledge and a lot of effort. This limits the amount and/or usefulness of available medical data sets for experimentation. Therefore, developing strategies to increase the number of annotations while lowering the needed domain knowledge is of interest. A possible strategy is the use of gamification, that is i.e. transforming the annotation task into a game. We propose an approach to gamify the task of annotating lung fluid cells from pathological whole slide images. As this domain is unknown to non-expert annotators, we transform images of cells detected with a RetinaNet architecture to the domain of flower images. This domain transfer is performed with a CycleGAN architecture for different cell types. In this more assessable domain, non-expert annotators can be (t)asked to annotate different kinds of flowers in a playful setting. In order to provide a proof of concept, this work shows that the domain transfer is possible by evaluating an image classification network trained on real cell images and tested on the cell images generated by the CycleGAN network. The classification network reaches an accuracy of 97.48% and 95.16% on the original lung fluid cells and transformed lung fluid cells, respectively. With this study, we lay the foundation for future research on gamification using CycleGANs.
Inter-Species Cell Detection: Datasets on pulmonary hemosiderophages in equine, human and feline specimens
Aug 19, 2021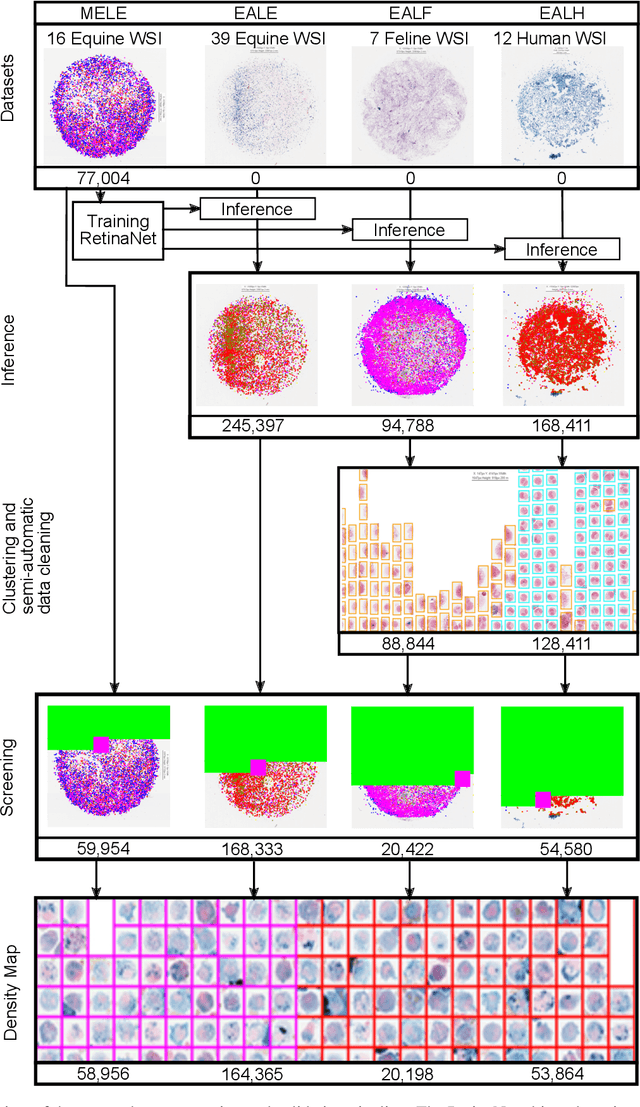
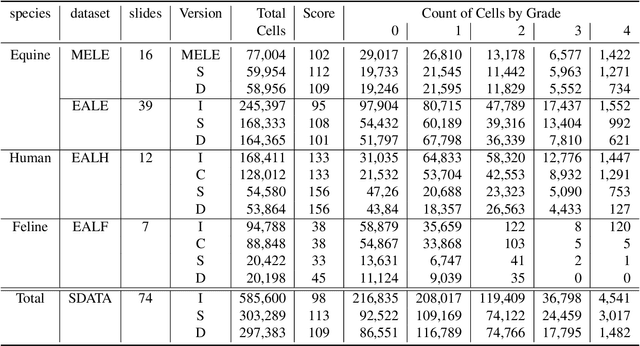
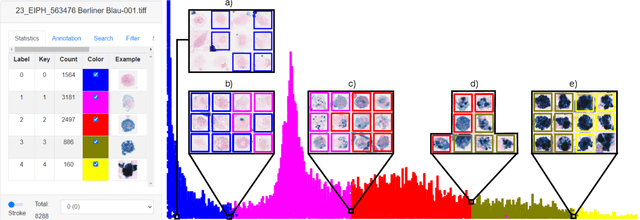
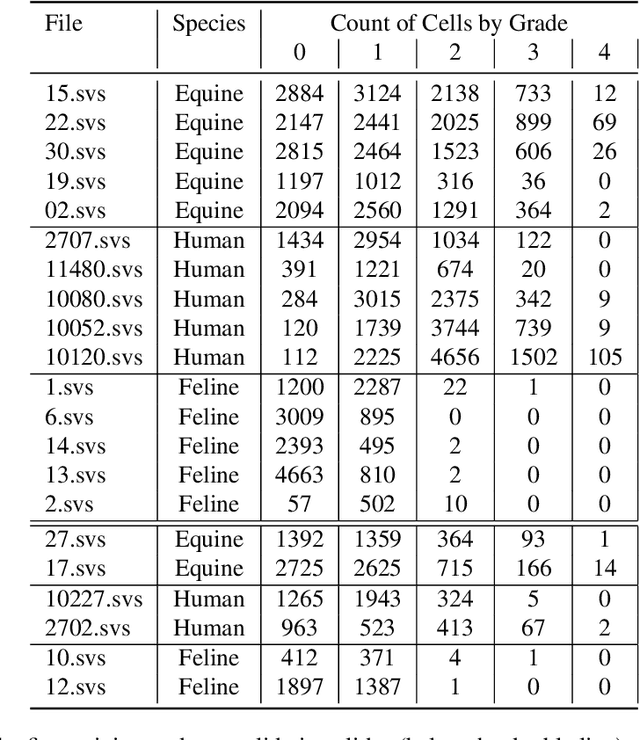
Abstract:Pulmonary hemorrhage (P-Hem) occurs among multiple species and can have various causes. Cytology of bronchoalveolarlavage fluid (BALF) using a 5-tier scoring system of alveolar macrophages based on their hemosiderin content is considered the most sensitive diagnostic method. We introduce a novel, fully annotated multi-species P-Hem dataset which consists of 74 cytology whole slide images (WSIs) with equine, feline and human samples. To create this high-quality and high-quantity dataset, we developed an annotation pipeline combining human expertise with deep learning and data visualisation techniques. We applied a deep learning-based object detection approach trained on 17 expertly annotated equine WSIs, to the remaining 39 equine, 12 human and 7 feline WSIs. The resulting annotations were semi-automatically screened for errors on multiple types of specialised annotation maps and finally reviewed by a trained pathologists. Our dataset contains a total of 297,383 hemosiderophages classified into five grades. It is one of the largest publicly availableWSIs datasets with respect to the number of annotations, the scanned area and the number of species covered.
Learning to be EXACT, Cell Detection for Asthma on Partially Annotated Whole Slide Images
Jan 13, 2021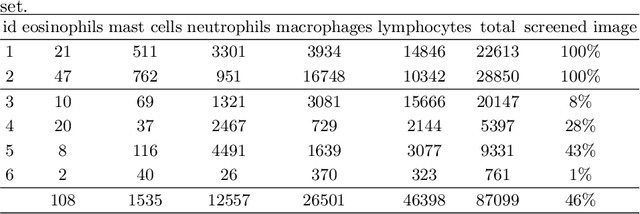
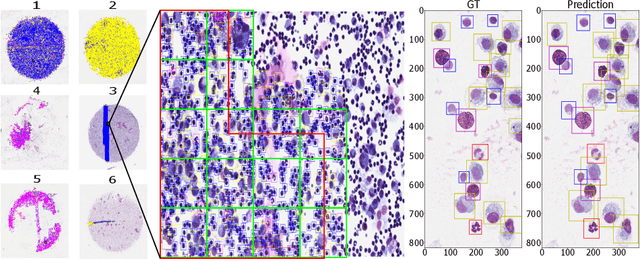
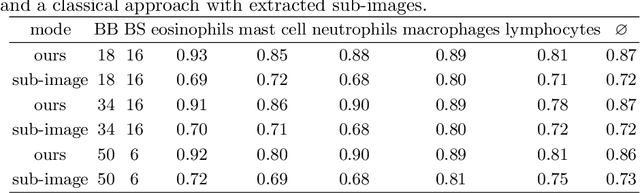
Abstract:Asthma is a chronic inflammatory disorder of the lower respiratory tract and naturally occurs in humans and animals including horses. The annotation of an asthma microscopy whole slide image (WSI) is an extremely labour-intensive task due to the hundreds of thousands of cells per WSI. To overcome the limitation of annotating WSI incompletely, we developed a training pipeline which can train a deep learning-based object detection model with partially annotated WSIs and compensate class imbalances on the fly. With this approach we can freely sample from annotated WSIs areas and are not restricted to fully annotated extracted sub-images of the WSI as with classical approaches. We evaluated our pipeline in a cross-validation setup with a fixed training set using a dataset of six equine WSIs of which four are partially annotated and used for training, and two fully annotated WSI are used for validation and testing. Our WSI-based training approach outperformed classical sub-image-based training methods by up to 15\% $mAP$ and yielded human-like performance when compared to the annotations of ten trained pathologists.
How Many Annotators Do We Need? -- A Study on the Influence of Inter-Observer Variability on the Reliability of Automatic Mitotic Figure Assessment
Jan 08, 2021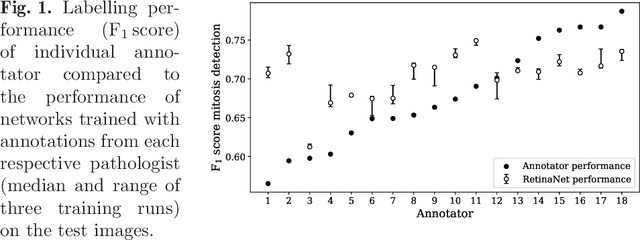
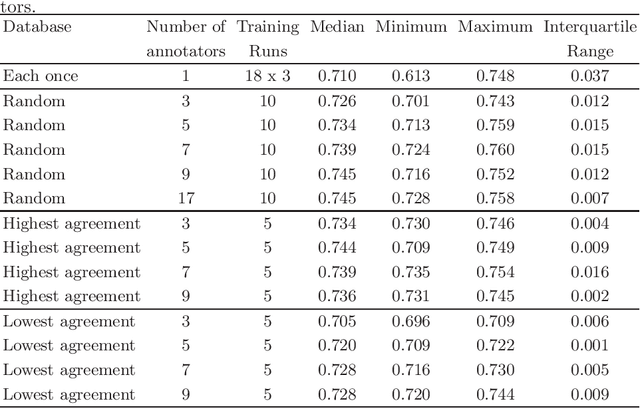
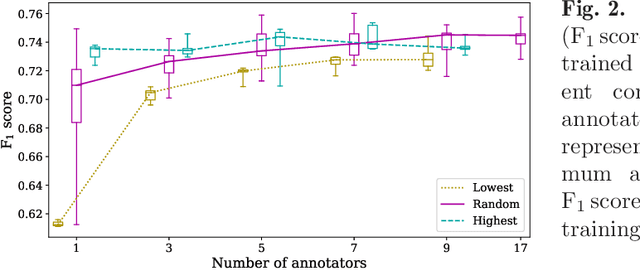
Abstract:Density of mitotic figures in histologic sections is a prognostically relevant characteristic for many tumours. Due to high inter-pathologist variability, deep learning-based algorithms are a promising solution to improve tumour prognostication. Pathologists are the gold standard for database development, however, labelling errors may hamper development of accurate algorithms. In the present work we evaluated the benefit of multi-expert consensus (n = 3, 5, 7, 9, 11) on algorithmic performance. While training with individual databases resulted in highly variable F$_1$ scores, performance was notably increased and more consistent when using the consensus of three annotators. Adding more annotators only resulted in minor improvements. We conclude that databases by few pathologists and high label accuracy may be the best compromise between high algorithmic performance and time investment.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge