Kevin de Haan
EdgeRunner 20B: Military Task Parity with GPT-5 while Running on the Edge
Oct 30, 2025Abstract:We present EdgeRunner 20B, a fine-tuned version of gpt-oss-20b optimized for military tasks. EdgeRunner 20B was trained on 1.6M high-quality records curated from military documentation and websites. We also present four new tests sets: (a) combat arms, (b) combat medic, (c) cyber operations, and (d) mil-bench-5k (general military knowledge). On these military test sets, EdgeRunner 20B matches or exceeds GPT-5 task performance with 95%+ statistical significance, except for the high reasoning setting on the combat medic test set and the low reasoning setting on the mil-bench-5k test set. Versus gpt-oss-20b, there is no statistically-significant regression on general-purpose benchmarks like ARC-C, GPQA Diamond, GSM8k, IFEval, MMLU Pro, or TruthfulQA, except for GSM8k in the low reasoning setting. We also present analyses on hyperparameter settings, cost, and throughput. These findings show that small, locally-hosted models are ideal solutions for data-sensitive operations such as in the military domain, allowing for deployment in air-gapped edge devices.
Label-free evaluation of lung and heart transplant biopsies using virtual staining
Sep 09, 2024



Abstract:Organ transplantation serves as the primary therapeutic strategy for end-stage organ failures. However, allograft rejection is a common complication of organ transplantation. Histological assessment is essential for the timely detection and diagnosis of transplant rejection and remains the gold standard. Nevertheless, the traditional histochemical staining process is time-consuming, costly, and labor-intensive. Here, we present a panel of virtual staining neural networks for lung and heart transplant biopsies, which digitally convert autofluorescence microscopic images of label-free tissue sections into their brightfield histologically stained counterparts, bypassing the traditional histochemical staining process. Specifically, we virtually generated Hematoxylin and Eosin (H&E), Masson's Trichrome (MT), and Elastic Verhoeff-Van Gieson (EVG) stains for label-free transplant lung tissue, along with H&E and MT stains for label-free transplant heart tissue. Subsequent blind evaluations conducted by three board-certified pathologists have confirmed that the virtual staining networks consistently produce high-quality histology images with high color uniformity, closely resembling their well-stained histochemical counterparts across various tissue features. The use of virtually stained images for the evaluation of transplant biopsies achieved comparable diagnostic outcomes to those obtained via traditional histochemical staining, with a concordance rate of 82.4% for lung samples and 91.7% for heart samples. Moreover, virtual staining models create multiple stains from the same autofluorescence input, eliminating structural mismatches observed between adjacent sections stained in the traditional workflow, while also saving tissue, expert time, and staining costs.
Virtual Gram staining of label-free bacteria using darkfield microscopy and deep learning
Jul 17, 2024



Abstract:Gram staining has been one of the most frequently used staining protocols in microbiology for over a century, utilized across various fields, including diagnostics, food safety, and environmental monitoring. Its manual procedures make it vulnerable to staining errors and artifacts due to, e.g., operator inexperience and chemical variations. Here, we introduce virtual Gram staining of label-free bacteria using a trained deep neural network that digitally transforms darkfield images of unstained bacteria into their Gram-stained equivalents matching brightfield image contrast. After a one-time training effort, the virtual Gram staining model processes an axial stack of darkfield microscopy images of label-free bacteria (never seen before) to rapidly generate Gram staining, bypassing several chemical steps involved in the conventional staining process. We demonstrated the success of the virtual Gram staining workflow on label-free bacteria samples containing Escherichia coli and Listeria innocua by quantifying the staining accuracy of the virtual Gram staining model and comparing the chromatic and morphological features of the virtually stained bacteria against their chemically stained counterparts. This virtual bacteria staining framework effectively bypasses the traditional Gram staining protocol and its challenges, including stain standardization, operator errors, and sensitivity to chemical variations.
Virtual histological staining of unlabeled autopsy tissue
Aug 02, 2023Abstract:Histological examination is a crucial step in an autopsy; however, the traditional histochemical staining of post-mortem samples faces multiple challenges, including the inferior staining quality due to autolysis caused by delayed fixation of cadaver tissue, as well as the resource-intensive nature of chemical staining procedures covering large tissue areas, which demand substantial labor, cost, and time. These challenges can become more pronounced during global health crises when the availability of histopathology services is limited, resulting in further delays in tissue fixation and more severe staining artifacts. Here, we report the first demonstration of virtual staining of autopsy tissue and show that a trained neural network can rapidly transform autofluorescence images of label-free autopsy tissue sections into brightfield equivalent images that match hematoxylin and eosin (H&E) stained versions of the same samples, eliminating autolysis-induced severe staining artifacts inherent in traditional histochemical staining of autopsied tissue. Our virtual H&E model was trained using >0.7 TB of image data and a data-efficient collaboration scheme that integrates the virtual staining network with an image registration network. The trained model effectively accentuated nuclear, cytoplasmic and extracellular features in new autopsy tissue samples that experienced severe autolysis, such as COVID-19 samples never seen before, where the traditional histochemical staining failed to provide consistent staining quality. This virtual autopsy staining technique can also be extended to necrotic tissue, and can rapidly and cost-effectively generate artifact-free H&E stains despite severe autolysis and cell death, also reducing labor, cost and infrastructure requirements associated with the standard histochemical staining.
Roadmap on Deep Learning for Microscopy
Mar 07, 2023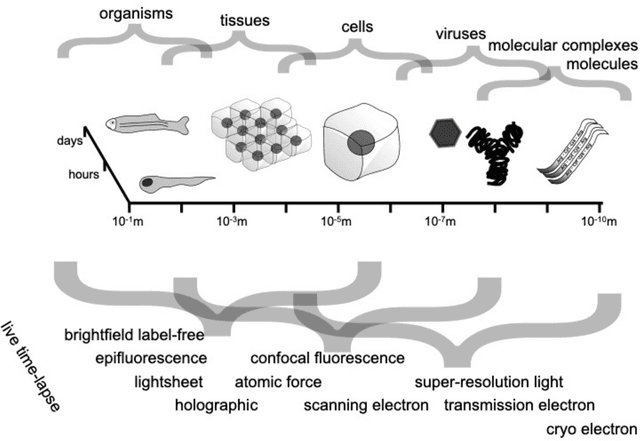
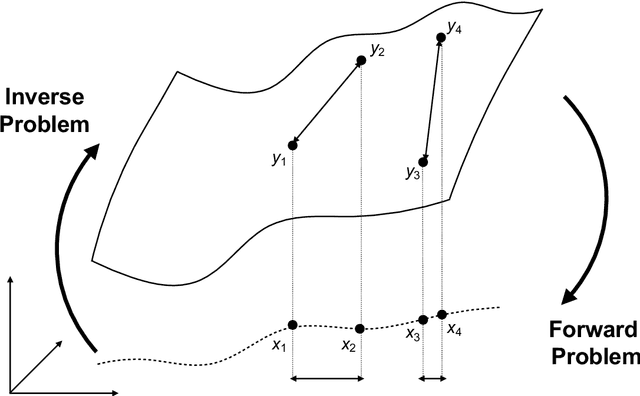
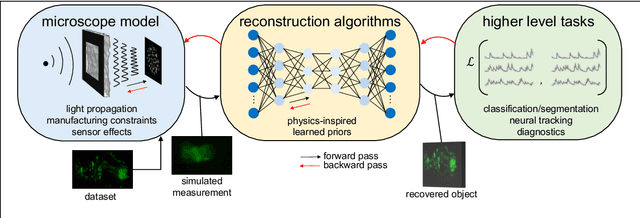
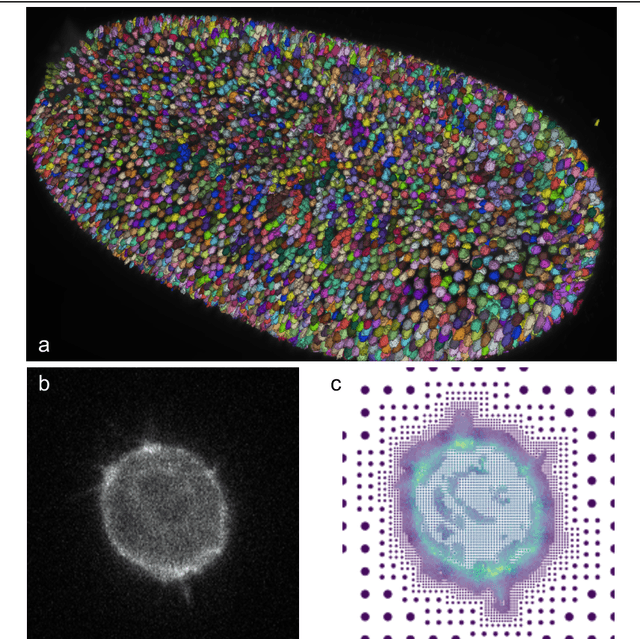
Abstract:Through digital imaging, microscopy has evolved from primarily being a means for visual observation of life at the micro- and nano-scale, to a quantitative tool with ever-increasing resolution and throughput. Artificial intelligence, deep neural networks, and machine learning are all niche terms describing computational methods that have gained a pivotal role in microscopy-based research over the past decade. This Roadmap is written collectively by prominent researchers and encompasses selected aspects of how machine learning is applied to microscopy image data, with the aim of gaining scientific knowledge by improved image quality, automated detection, segmentation, classification and tracking of objects, and efficient merging of information from multiple imaging modalities. We aim to give the reader an overview of the key developments and an understanding of possibilities and limitations of machine learning for microscopy. It will be of interest to a wide cross-disciplinary audience in the physical sciences and life sciences.
Virtual stain transfer in histology via cascaded deep neural networks
Jul 14, 2022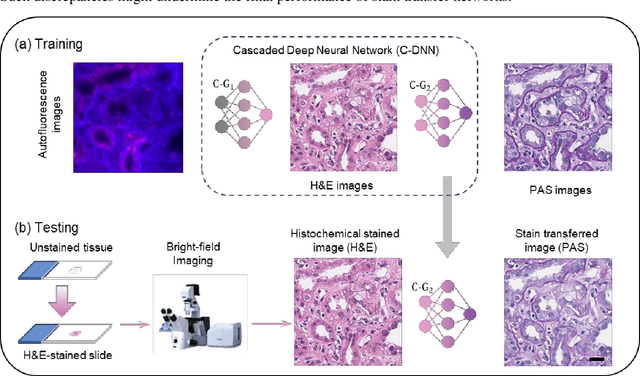
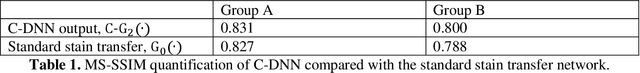
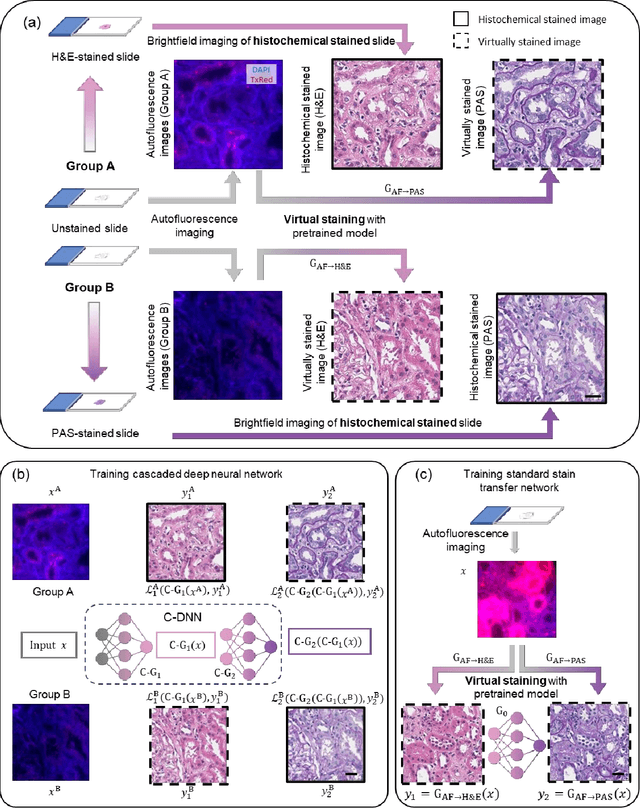
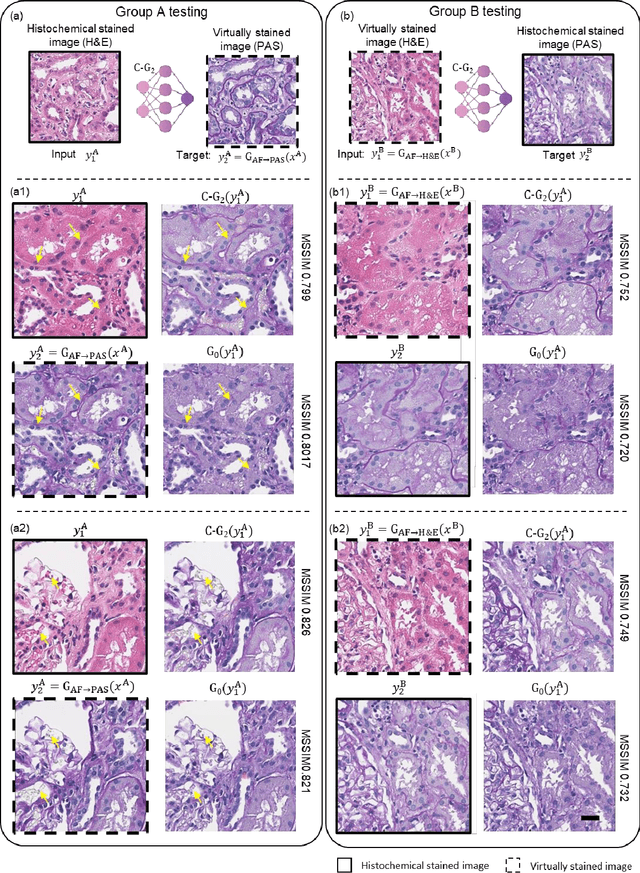
Abstract:Pathological diagnosis relies on the visual inspection of histologically stained thin tissue specimens, where different types of stains are applied to bring contrast to and highlight various desired histological features. However, the destructive histochemical staining procedures are usually irreversible, making it very difficult to obtain multiple stains on the same tissue section. Here, we demonstrate a virtual stain transfer framework via a cascaded deep neural network (C-DNN) to digitally transform hematoxylin and eosin (H&E) stained tissue images into other types of histological stains. Unlike a single neural network structure which only takes one stain type as input to digitally output images of another stain type, C-DNN first uses virtual staining to transform autofluorescence microscopy images into H&E and then performs stain transfer from H&E to the domain of the other stain in a cascaded manner. This cascaded structure in the training phase allows the model to directly exploit histochemically stained image data on both H&E and the target special stain of interest. This advantage alleviates the challenge of paired data acquisition and improves the image quality and color accuracy of the virtual stain transfer from H&E to another stain. We validated the superior performance of this C-DNN approach using kidney needle core biopsy tissue sections and successfully transferred the H&E-stained tissue images into virtual PAS (periodic acid-Schiff) stain. This method provides high-quality virtual images of special stains using existing, histochemically stained slides and creates new opportunities in digital pathology by performing highly accurate stain-to-stain transformations.
Virtual staining of defocused autofluorescence images of unlabeled tissue using deep neural networks
Jul 06, 2022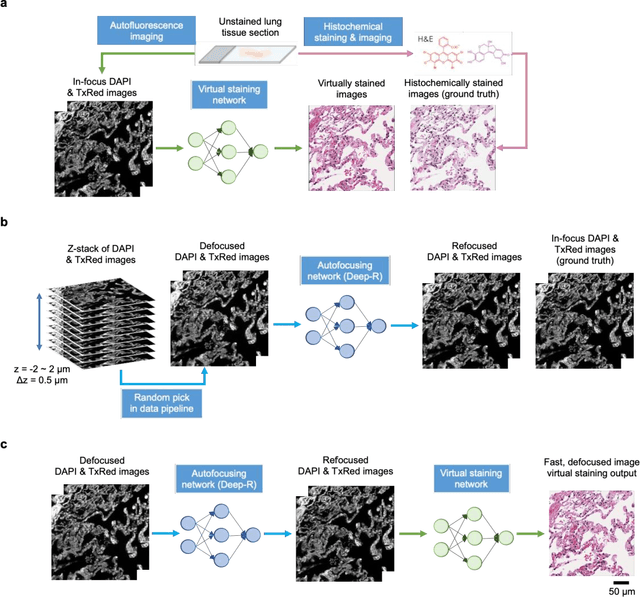
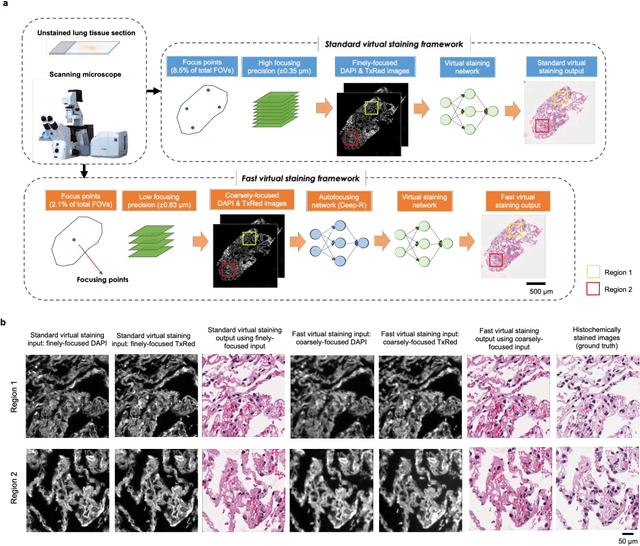
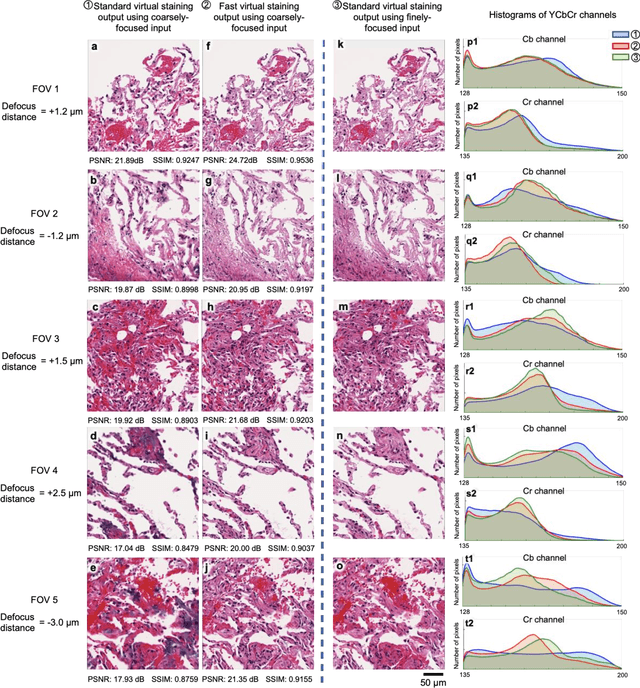
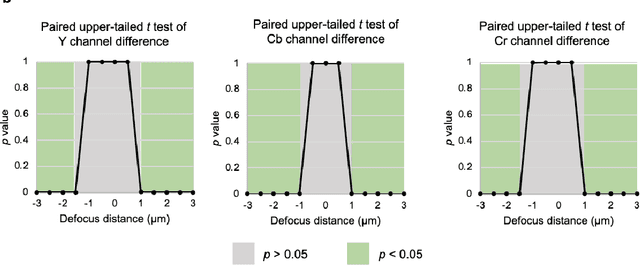
Abstract:Deep learning-based virtual staining was developed to introduce image contrast to label-free tissue sections, digitally matching the histological staining, which is time-consuming, labor-intensive, and destructive to tissue. Standard virtual staining requires high autofocusing precision during the whole slide imaging of label-free tissue, which consumes a significant portion of the total imaging time and can lead to tissue photodamage. Here, we introduce a fast virtual staining framework that can stain defocused autofluorescence images of unlabeled tissue, achieving equivalent performance to virtual staining of in-focus label-free images, also saving significant imaging time by lowering the microscope's autofocusing precision. This framework incorporates a virtual-autofocusing neural network to digitally refocus the defocused images and then transforms the refocused images into virtually stained images using a successive network. These cascaded networks form a collaborative inference scheme: the virtual staining model regularizes the virtual-autofocusing network through a style loss during the training. To demonstrate the efficacy of this framework, we trained and blindly tested these networks using human lung tissue. Using 4x fewer focus points with 2x lower focusing precision, we successfully transformed the coarsely-focused autofluorescence images into high-quality virtually stained H&E images, matching the standard virtual staining framework that used finely-focused autofluorescence input images. Without sacrificing the staining quality, this framework decreases the total image acquisition time needed for virtual staining of a label-free whole-slide image (WSI) by ~32%, together with a ~89% decrease in the autofocusing time, and has the potential to eliminate the laborious and costly histochemical staining process in pathology.
Label-free virtual HER2 immunohistochemical staining of breast tissue using deep learning
Dec 08, 2021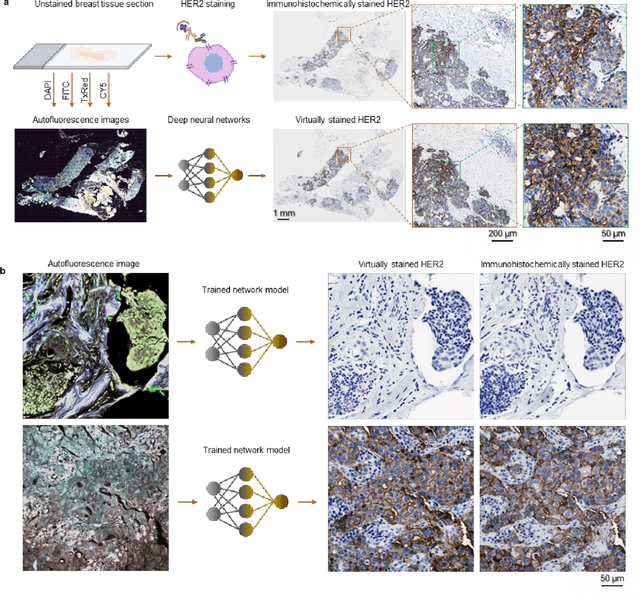
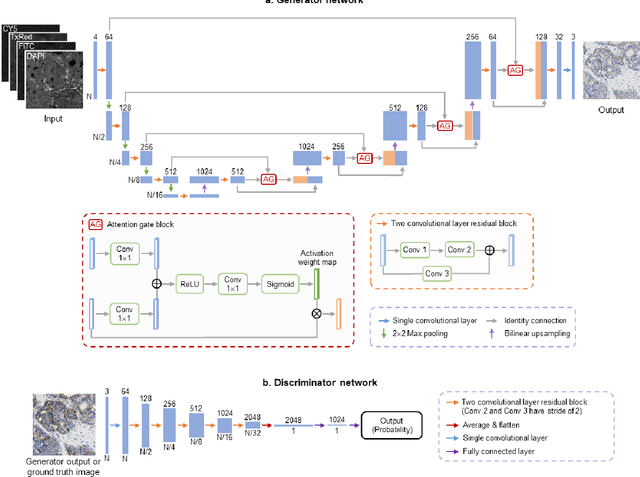
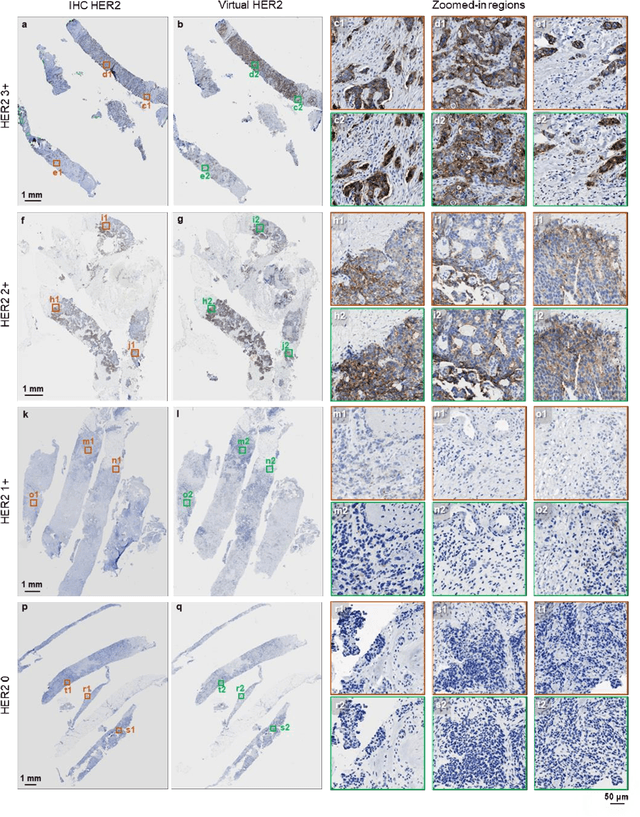
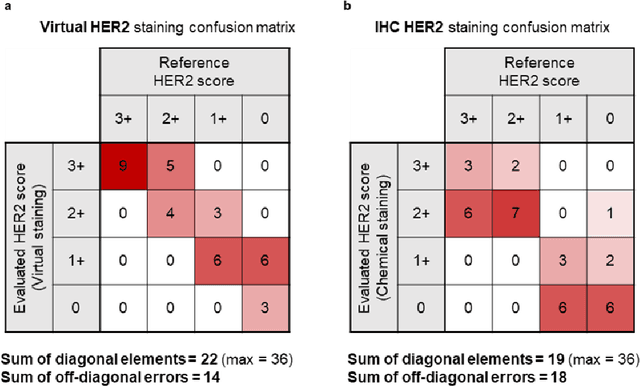
Abstract:The immunohistochemical (IHC) staining of the human epidermal growth factor receptor 2 (HER2) biomarker is widely practiced in breast tissue analysis, preclinical studies and diagnostic decisions, guiding cancer treatment and investigation of pathogenesis. HER2 staining demands laborious tissue treatment and chemical processing performed by a histotechnologist, which typically takes one day to prepare in a laboratory, increasing analysis time and associated costs. Here, we describe a deep learning-based virtual HER2 IHC staining method using a conditional generative adversarial network that is trained to rapidly transform autofluorescence microscopic images of unlabeled/label-free breast tissue sections into bright-field equivalent microscopic images, matching the standard HER2 IHC staining that is chemically performed on the same tissue sections. The efficacy of this virtual HER2 staining framework was demonstrated by quantitative analysis, in which three board-certified breast pathologists blindly graded the HER2 scores of virtually stained and immunohistochemically stained HER2 whole slide images (WSIs) to reveal that the HER2 scores determined by inspecting virtual IHC images are as accurate as their immunohistochemically stained counterparts. A second quantitative blinded study performed by the same diagnosticians further revealed that the virtually stained HER2 images exhibit a comparable staining quality in the level of nuclear detail, membrane clearness, and absence of staining artifacts with respect to their immunohistochemically stained counterparts. This virtual HER2 staining framework bypasses the costly, laborious, and time-consuming IHC staining procedures in laboratory, and can be extended to other types of biomarkers to accelerate the IHC tissue staining used in life sciences and biomedical workflow.
Deep learning-based transformation of the H&E stain into special stains improves kidney disease diagnosis
Aug 20, 2020
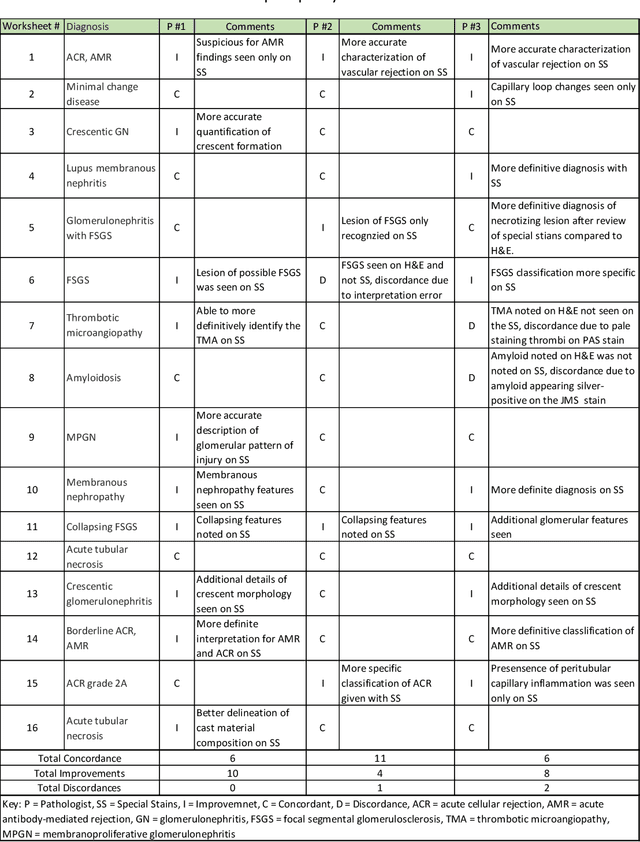


Abstract:Pathology is practiced by visual inspection of histochemically stained slides. Most commonly, the hematoxylin and eosin (H&E) stain is used in the diagnostic workflow and it is the gold standard for cancer diagnosis. However, in many cases, especially for non-neoplastic diseases, additional "special stains" are used to provide different levels of contrast and color to tissue components and allow pathologists to get a clearer diagnostic picture. In this study, we demonstrate the utility of supervised learning-based computational stain transformation from H&E to different special stains (Masson's Trichrome, periodic acid-Schiff and Jones silver stain) using tissue sections from kidney needle core biopsies. Based on evaluation by three renal pathologists, followed by adjudication by a fourth renal pathologist, we show that the generation of virtual special stains from existing H&E images improves the diagnosis in several non-neoplastic kidney diseases, sampled from 16 unique subjects. Adjudication of N=48 diagnoses from the three pathologists revealed that the virtually generated special stains yielded 22 improvements (45.8%), 23 concordances (47.9%) and 3 discordances (6.3%), when compared against the use of H&E stained tissue only. As the virtual transformation of H&E images into special stains can be achieved in less than 1 min per patient core specimen slide, this stain-to-stain transformation framework can improve the quality of the preliminary diagnosis when additional special stains are needed, along with significant savings in time and cost, reducing the burden on healthcare system and patients.
Deep learning-based holographic polarization microscopy
Jul 01, 2020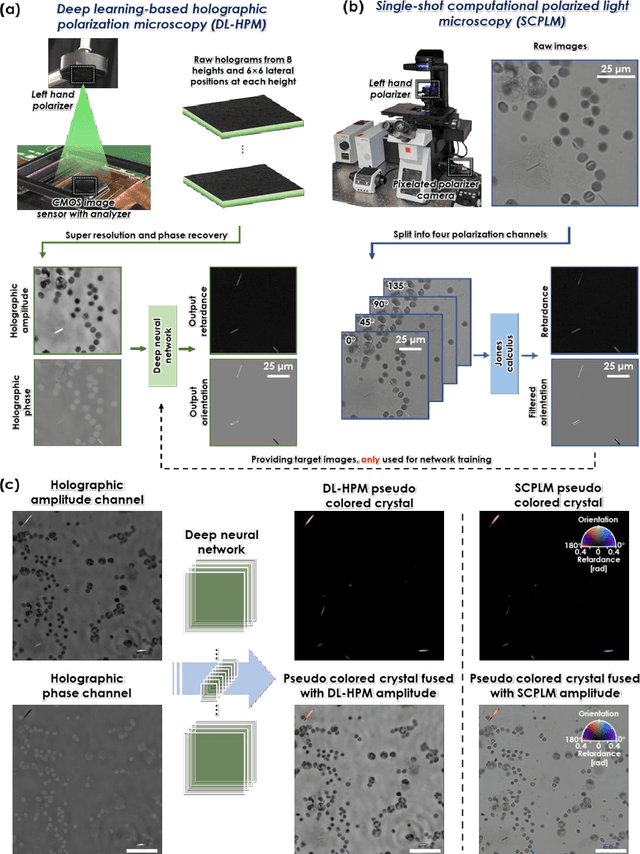
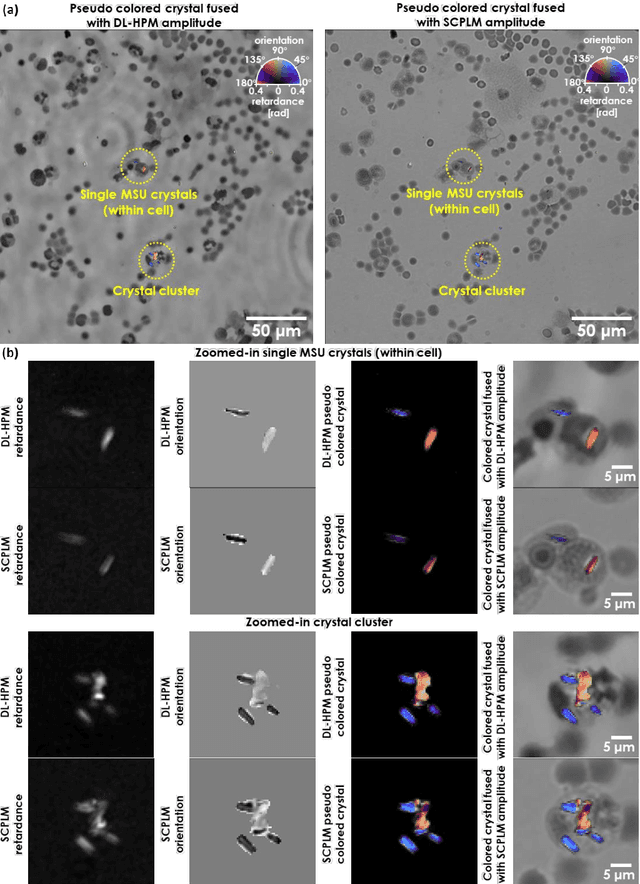
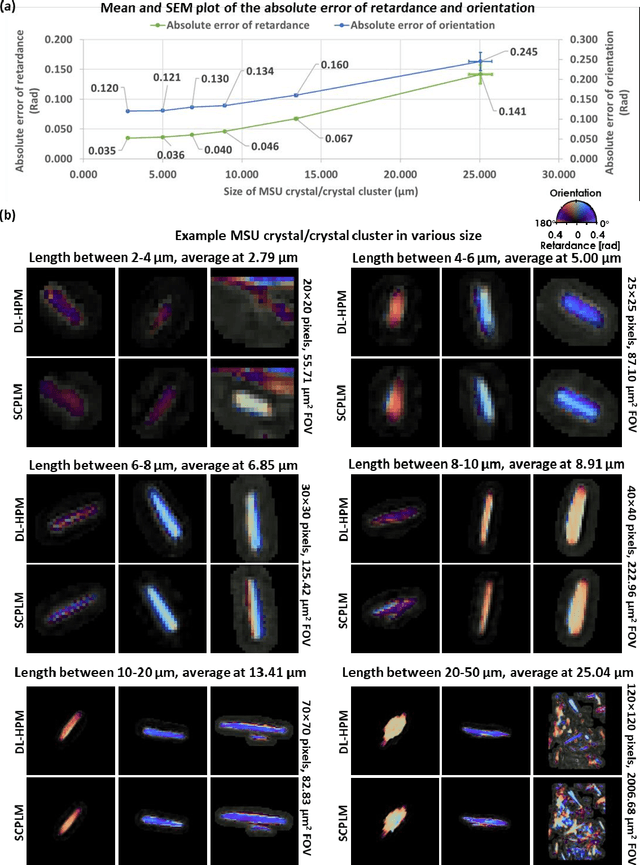
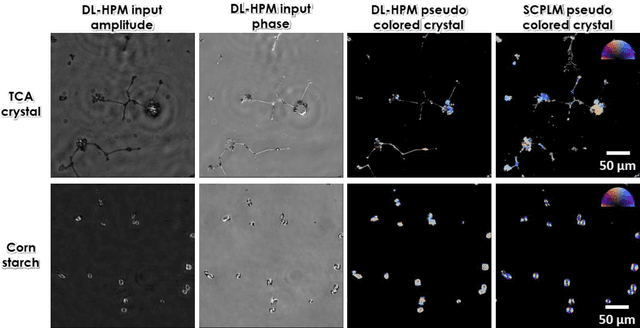
Abstract:Polarized light microscopy provides high contrast to birefringent specimen and is widely used as a diagnostic tool in pathology. However, polarization microscopy systems typically operate by analyzing images collected from two or more light paths in different states of polarization, which lead to relatively complex optical designs, high system costs or experienced technicians being required. Here, we present a deep learning-based holographic polarization microscope that is capable of obtaining quantitative birefringence retardance and orientation information of specimen from a phase recovered hologram, while only requiring the addition of one polarizer/analyzer pair to an existing holographic imaging system. Using a deep neural network, the reconstructed holographic images from a single state of polarization can be transformed into images equivalent to those captured using a single-shot computational polarized light microscope (SCPLM). Our analysis shows that a trained deep neural network can extract the birefringence information using both the sample specific morphological features as well as the holographic amplitude and phase distribution. To demonstrate the efficacy of this method, we tested it by imaging various birefringent samples including e.g., monosodium urate (MSU) and triamcinolone acetonide (TCA) crystals. Our method achieves similar results to SCPLM both qualitatively and quantitatively, and due to its simpler optical design and significantly larger field-of-view, this method has the potential to expand the access to polarization microscopy and its use for medical diagnosis in resource limited settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge