Yuxuan Qi
Deep Learning-based Detection of Bacterial Swarm Motion Using a Single Image
Oct 19, 2024
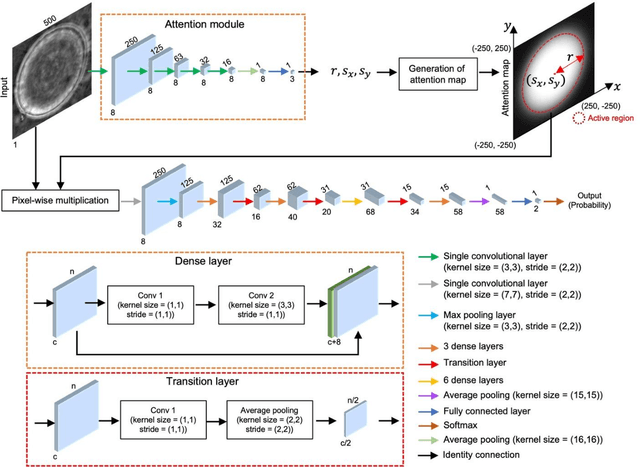
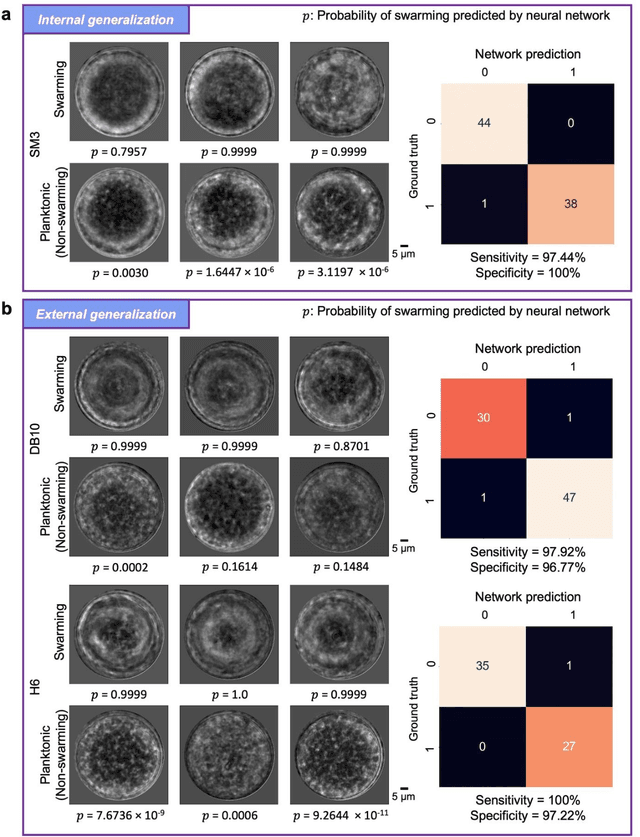
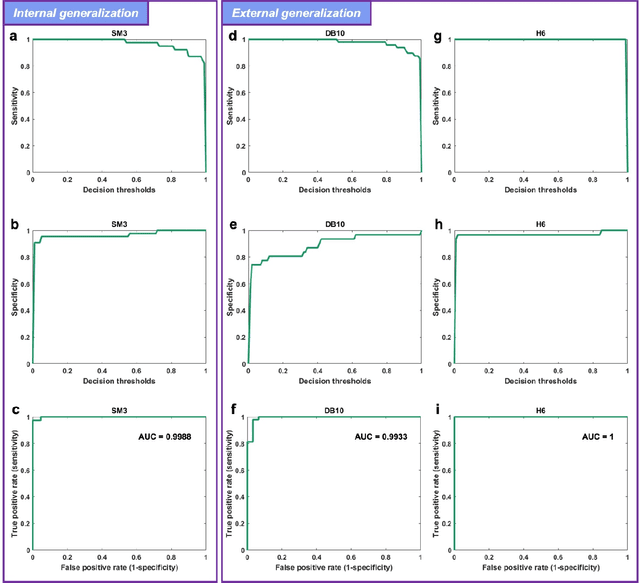
Abstract:Distinguishing between swarming and swimming, the two principal forms of bacterial movement, holds significant conceptual and clinical relevance. This is because bacteria that exhibit swarming capabilities often possess unique properties crucial to the pathogenesis of infectious diseases and may also have therapeutic potential. Here, we report a deep learning-based swarming classifier that rapidly and autonomously predicts swarming probability using a single blurry image. Compared with traditional video-based, manually-processed approaches, our method is particularly suited for high-throughput environments and provides objective, quantitative assessments of swarming probability. The swarming classifier demonstrated in our work was trained on Enterobacter sp. SM3 and showed good performance when blindly tested on new swarming (positive) and swimming (negative) test images of SM3, achieving a sensitivity of 97.44% and a specificity of 100%. Furthermore, this classifier demonstrated robust external generalization capabilities when applied to unseen bacterial species, such as Serratia marcescens DB10 and Citrobacter koseri H6. It blindly achieved a sensitivity of 97.92% and a specificity of 96.77% for DB10, and a sensitivity of 100% and a specificity of 97.22% for H6. This competitive performance indicates the potential to adapt our approach for diagnostic applications through portable devices or even smartphones. This adaptation would facilitate rapid, objective, on-site screening for bacterial swarming motility, potentially enhancing the early detection and treatment assessment of various diseases, including inflammatory bowel diseases (IBD) and urinary tract infections (UTI).
Label-free evaluation of lung and heart transplant biopsies using virtual staining
Sep 09, 2024



Abstract:Organ transplantation serves as the primary therapeutic strategy for end-stage organ failures. However, allograft rejection is a common complication of organ transplantation. Histological assessment is essential for the timely detection and diagnosis of transplant rejection and remains the gold standard. Nevertheless, the traditional histochemical staining process is time-consuming, costly, and labor-intensive. Here, we present a panel of virtual staining neural networks for lung and heart transplant biopsies, which digitally convert autofluorescence microscopic images of label-free tissue sections into their brightfield histologically stained counterparts, bypassing the traditional histochemical staining process. Specifically, we virtually generated Hematoxylin and Eosin (H&E), Masson's Trichrome (MT), and Elastic Verhoeff-Van Gieson (EVG) stains for label-free transplant lung tissue, along with H&E and MT stains for label-free transplant heart tissue. Subsequent blind evaluations conducted by three board-certified pathologists have confirmed that the virtual staining networks consistently produce high-quality histology images with high color uniformity, closely resembling their well-stained histochemical counterparts across various tissue features. The use of virtually stained images for the evaluation of transplant biopsies achieved comparable diagnostic outcomes to those obtained via traditional histochemical staining, with a concordance rate of 82.4% for lung samples and 91.7% for heart samples. Moreover, virtual staining models create multiple stains from the same autofluorescence input, eliminating structural mismatches observed between adjacent sections stained in the traditional workflow, while also saving tissue, expert time, and staining costs.
Multi-modal Evidential Fusion Network for Trusted PET/CT Tumor Segmentation
Jun 26, 2024Abstract:Accurate segmentation of tumors in PET/CT images is important in computer-aided diagnosis and treatment of cancer. The key issue of such a segmentation problem lies in the effective integration of complementary information from PET and CT images. However, the quality of PET and CT images varies widely in clinical settings, which leads to uncertainty in the modality information extracted by networks. To take the uncertainty into account in multi-modal information fusion, this paper proposes a novel Multi-modal Evidential Fusion Network (MEFN) comprising a Cross-Modal Feature Learning (CFL) module and a Multi-modal Trusted Fusion (MTF) module. The CFL module reduces the domain gap upon modality conversion and highlights common tumor features, thereby alleviating the needs of the segmentation module to handle modality specificity. The MTF module utilizes mutual attention mechanisms and an uncertainty calibrator to fuse modality features based on modality uncertainty and then fuse the segmentation results under the guidance of Dempster-Shafer Theory. Besides, a new uncertainty perceptual loss is introduced to force the model focusing on uncertain features and hence improve its ability to extract trusted modality information. Extensive comparative experiments are conducted on two publicly available PET/CT datasets to evaluate the performance of our proposed method whose results demonstrate that our MEFN significantly outperforms state-of-the-art methods with improvements of 2.15% and 3.23% in DSC scores on the AutoPET dataset and the Hecktor dataset, respectively. More importantly, our model can provide radiologists with credible uncertainty of the segmentation results for their decision in accepting or rejecting the automatic segmentation results, which is particularly important for clinical applications. Our code will be available at https://github.com/QPaws/MEFN.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge