Jay DeYoung
Understanding Usage and Engagement in AI-Powered Scientific Research Tools: The Asta Interaction Dataset
Feb 26, 2026Abstract:AI-powered scientific research tools are rapidly being integrated into research workflows, yet the field lacks a clear lens into how researchers use these systems in real-world settings. We present and analyze the Asta Interaction Dataset, a large-scale resource comprising over 200,000 user queries and interaction logs from two deployed tools (a literature discovery interface and a scientific question-answering interface) within an LLM-powered retrieval-augmented generation platform. Using this dataset, we characterize query patterns, engagement behaviors, and how usage evolves with experience. We find that users submit longer and more complex queries than in traditional search, and treat the system as a collaborative research partner, delegating tasks such as drafting content and identifying research gaps. Users treat generated responses as persistent artifacts, revisiting and navigating among outputs and cited evidence in non-linear ways. With experience, users issue more targeted queries and engage more deeply with supporting citations, although keyword-style queries persist even among experienced users. We release the anonymized dataset and analysis with a new query intent taxonomy to inform future designs of real-world AI research assistants and to support realistic evaluation.
PreScience: A Benchmark for Forecasting Scientific Contributions
Feb 24, 2026Abstract:Can AI systems trained on the scientific record up to a fixed point in time forecast the scientific advances that follow? Such a capability could help researchers identify collaborators and impactful research directions, and anticipate which problems and methods will become central next. We introduce PreScience -- a scientific forecasting benchmark that decomposes the research process into four interdependent generative tasks: collaborator prediction, prior work selection, contribution generation, and impact prediction. PreScience is a carefully curated dataset of 98K recent AI-related research papers, featuring disambiguated author identities, temporally aligned scholarly metadata, and a structured graph of companion author publication histories and citations spanning 502K total papers. We develop baselines and evaluations for each task, including LACERScore, a novel LLM-based measure of contribution similarity that outperforms previous metrics and approximates inter-annotator agreement. We find substantial headroom remains in each task -- e.g. in contribution generation, frontier LLMs achieve only moderate similarity to the ground-truth (GPT-5, averages 5.6 on a 1-10 scale). When composed into a 12-month end-to-end simulation of scientific production, the resulting synthetic corpus is systematically less diverse and less novel than human-authored research from the same period.
Automated Metrics for Medical Multi-Document Summarization Disagree with Human Evaluations
May 23, 2023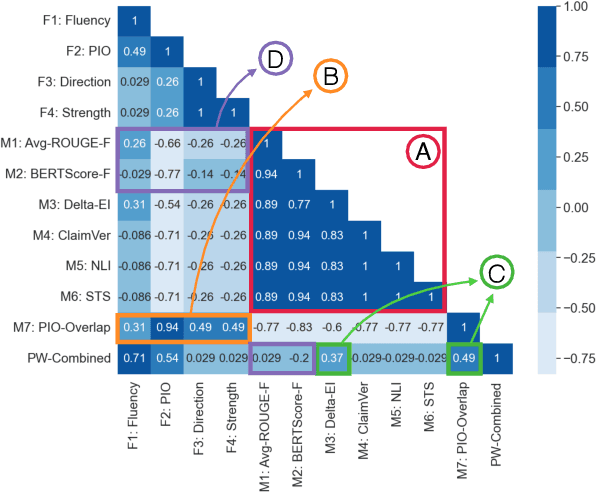

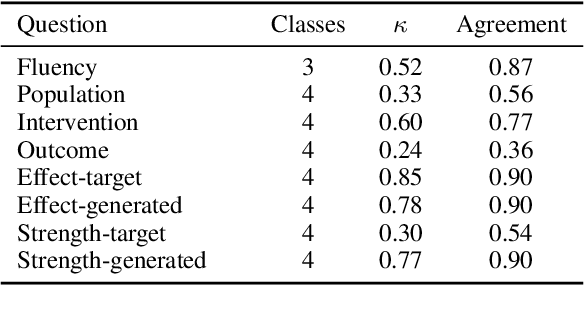

Abstract:Evaluating multi-document summarization (MDS) quality is difficult. This is especially true in the case of MDS for biomedical literature reviews, where models must synthesize contradicting evidence reported across different documents. Prior work has shown that rather than performing the task, models may exploit shortcuts that are difficult to detect using standard n-gram similarity metrics such as ROUGE. Better automated evaluation metrics are needed, but few resources exist to assess metrics when they are proposed. Therefore, we introduce a dataset of human-assessed summary quality facets and pairwise preferences to encourage and support the development of better automated evaluation methods for literature review MDS. We take advantage of community submissions to the Multi-document Summarization for Literature Review (MSLR) shared task to compile a diverse and representative sample of generated summaries. We analyze how automated summarization evaluation metrics correlate with lexical features of generated summaries, to other automated metrics including several we propose in this work, and to aspects of human-assessed summary quality. We find that not only do automated metrics fail to capture aspects of quality as assessed by humans, in many cases the system rankings produced by these metrics are anti-correlated with rankings according to human annotators.
Jointly Extracting Interventions, Outcomes, and Findings from RCT Reports with LLMs
May 05, 2023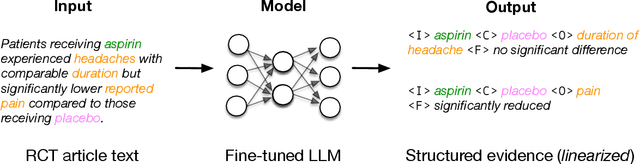
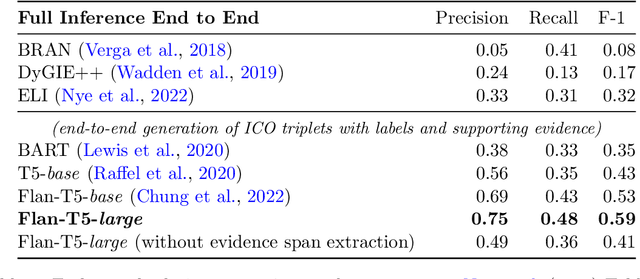
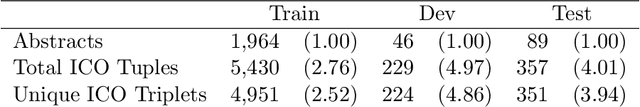
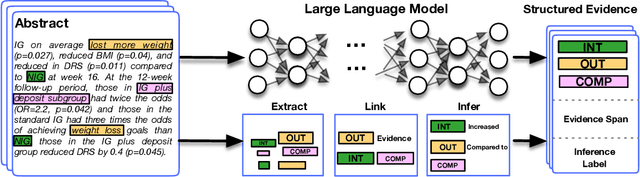

Abstract:Results from Randomized Controlled Trials (RCTs) establish the comparative effectiveness of interventions, and are in turn critical inputs for evidence-based care. However, results from RCTs are presented in (often unstructured) natural language articles describing the design, execution, and outcomes of trials; clinicians must manually extract findings pertaining to interventions and outcomes of interest from such articles. This onerous manual process has motivated work on (semi-)automating extraction of structured evidence from trial reports. In this work we propose and evaluate a text-to-text model built on instruction-tuned Large Language Models (LLMs) to jointly extract Interventions, Outcomes, and Comparators (ICO elements) from clinical abstracts, and infer the associated results reported. Manual (expert) and automated evaluations indicate that framing evidence extraction as a conditional generation task and fine-tuning LLMs for this purpose realizes considerable ($\sim$20 point absolute F1 score) gains over the previous SOTA. We perform ablations and error analyses to assess aspects that contribute to model performance, and to highlight potential directions for further improvements. We apply our model to a collection of published RCTs through mid-2022, and release a searchable database of structured findings (anonymously for now): bit.ly/joint-relations-extraction-mlhc
Do Multi-Document Summarization Models Synthesize?
Jan 31, 2023Abstract:Multi-document summarization entails producing concise synopses of collections of inputs. For some applications, the synopsis should accurately \emph{synthesize} inputs with respect to a key property or aspect. For example, a synopsis of film reviews all written about a particular movie should reflect the average critic consensus. As a more consequential example, consider narrative summaries that accompany biomedical \emph{systematic reviews} of clinical trial results. These narratives should fairly summarize the potentially conflicting results from individual trials. In this paper we ask: To what extent do modern multi-document summarization models implicitly perform this type of synthesis? To assess this we perform a suite of experiments that probe the degree to which conditional generation models trained for summarization using standard methods yield outputs that appropriately synthesize inputs. We find that existing models do partially perform synthesis, but do so imperfectly. In particular, they are over-sensitive to changes in input ordering and under-sensitive to changes in input compositions (e.g., the ratio of positive to negative movie reviews). We propose a simple, general method for improving model synthesis capabilities by generating an explicitly diverse set of candidate outputs, and then selecting from these the string best aligned with the expected aggregate measure for the inputs, or \emph{abstaining} when the model produces no good candidate. This approach improves model synthesis performance. We hope highlighting the need for synthesis (in some summarization settings), motivates further research into multi-document summarization methods and learning objectives that explicitly account for the need to synthesize.
Entity Anchored ICD Coding
Aug 15, 2022
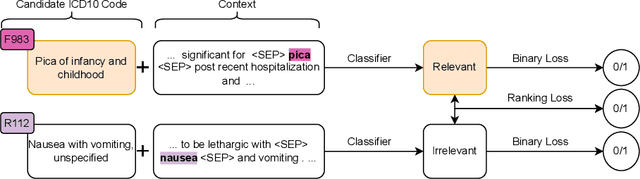
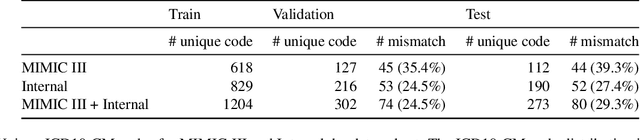
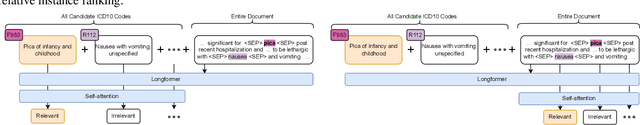
Abstract:Medical coding is a complex task, requiring assignment of a subset of over 72,000 ICD codes to a patient's notes. Modern natural language processing approaches to these tasks have been challenged by the length of the input and size of the output space. We limit our model inputs to a small window around medical entities found in our documents. From those local contexts, we build contextualized representations of both ICD codes and entities, and aggregate over these representations to form document-level predictions. In contrast to existing methods which use a representation fixed either in size or by codes seen in training, we represent ICD codes by encoding the code description with local context. We discuss metrics appropriate to deploying coding systems in practice. We show that our approach is superior to existing methods in both standard and deployable measures, including performance on rare and unseen codes.
MS2: Multi-Document Summarization of Medical Studies
Apr 15, 2021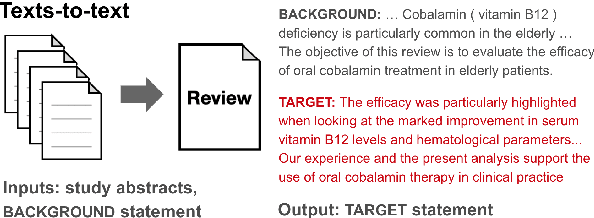
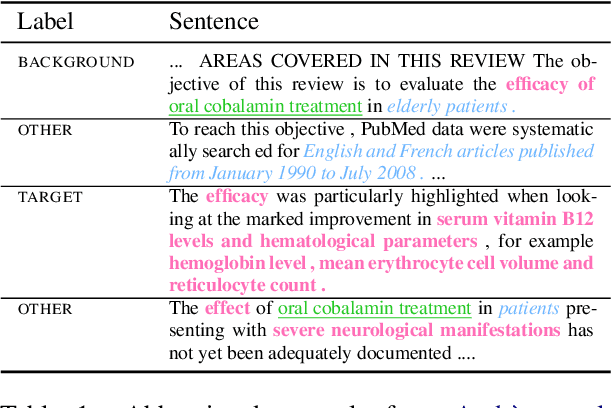



Abstract:To assess the effectiveness of any medical intervention, researchers must conduct a time-intensive and highly manual literature review. NLP systems can help to automate or assist in parts of this expensive process. In support of this goal, we release MS^2 (Multi-Document Summarization of Medical Studies), a dataset of over 470k documents and 20k summaries derived from the scientific literature. This dataset facilitates the development of systems that can assess and aggregate contradictory evidence across multiple studies, and is the first large-scale, publicly available multi-document summarization dataset in the biomedical domain. We experiment with a summarization system based on BART, with promising early results. We formulate our summarization inputs and targets in both free text and structured forms and modify a recently proposed metric to assess the quality of our system's generated summaries. Data and models are available at https://github.com/allenai/ms2
Understanding Clinical Trial Reports: Extracting Medical Entities and Their Relations
Oct 08, 2020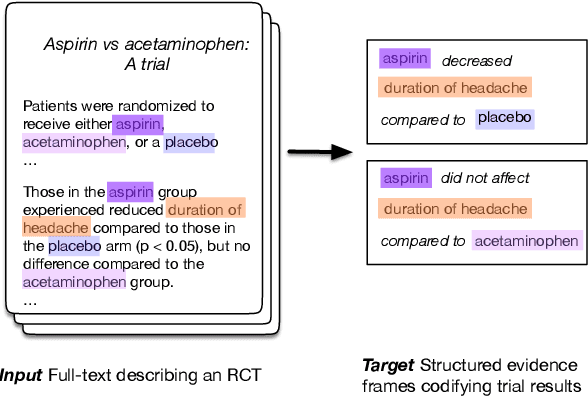
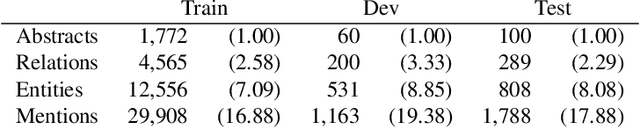
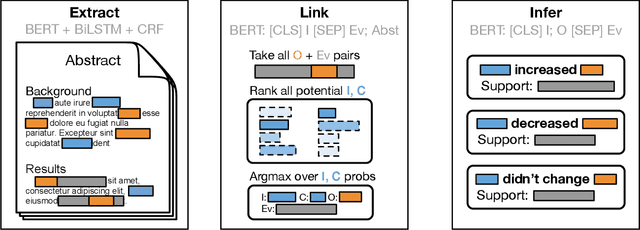

Abstract:The best evidence concerning comparative treatment effectiveness comes from clinical trials, the results of which are reported in unstructured articles. Medical experts must manually extract information from articles to inform decision-making, which is time-consuming and expensive. Here we consider the end-to-end task of both (a) extracting treatments and outcomes from full-text articles describing clinical trials (entity identification) and, (b) inferring the reported results for the former with respect to the latter (relation extraction). We introduce new data for this task, and evaluate models that have recently achieved state-of-the-art results on similar tasks in Natural Language Processing. We then propose a new method motivated by how trial results are typically presented that outperforms these purely data-driven baselines. Finally, we run a fielded evaluation of the model with a non-profit seeking to identify existing drugs that might be re-purposed for cancer, showing the potential utility of end-to-end evidence extraction systems.
Evidence Inference 2.0: More Data, Better Models
May 14, 2020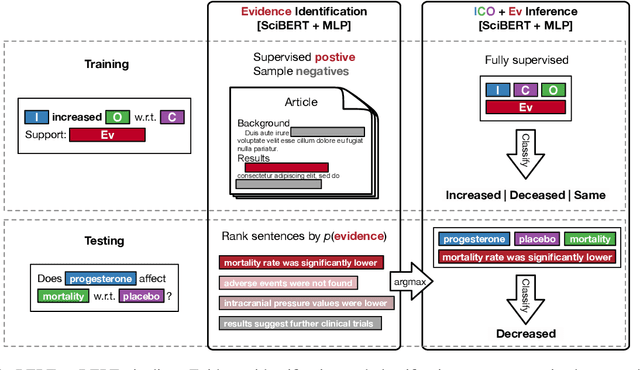
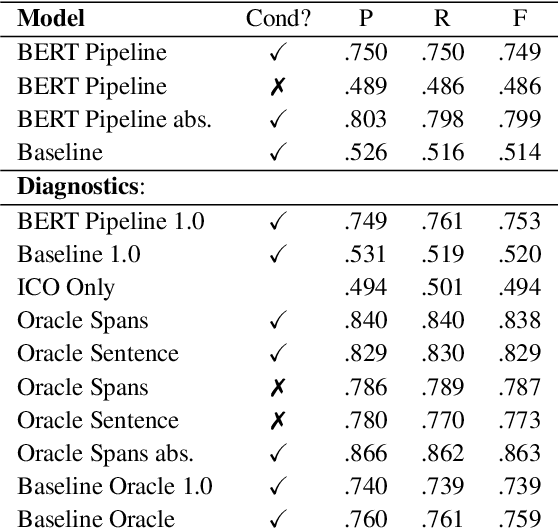
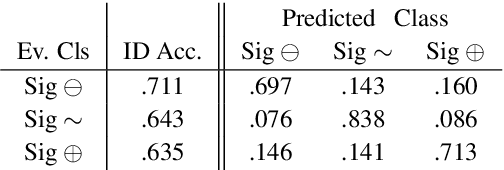
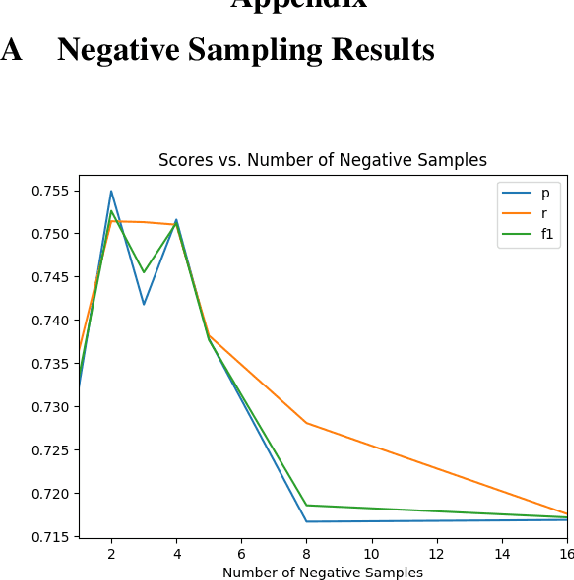
Abstract:How do we most effectively treat a disease or condition? Ideally, we could consult a database of evidence gleaned from clinical trials to answer such questions. Unfortunately, no such database exists; clinical trial results are instead disseminated primarily via lengthy natural language articles. Perusing all such articles would be prohibitively time-consuming for healthcare practitioners; they instead tend to depend on manually compiled systematic reviews of medical literature to inform care. NLP may speed this process up, and eventually facilitate immediate consult of published evidence. The Evidence Inference dataset was recently released to facilitate research toward this end. This task entails inferring the comparative performance of two treatments, with respect to a given outcome, from a particular article (describing a clinical trial) and identifying supporting evidence. For instance: Does this article report that chemotherapy performed better than surgery for five-year survival rates of operable cancers? In this paper, we collect additional annotations to expand the Evidence Inference dataset by 25\%, provide stronger baseline models, systematically inspect the errors that these make, and probe dataset quality. We also release an abstract only (as opposed to full-texts) version of the task for rapid model prototyping. The updated corpus, documentation, and code for new baselines and evaluations are available at http://evidence-inference.ebm-nlp.com/.
ERASER: A Benchmark to Evaluate Rationalized NLP Models
Nov 08, 2019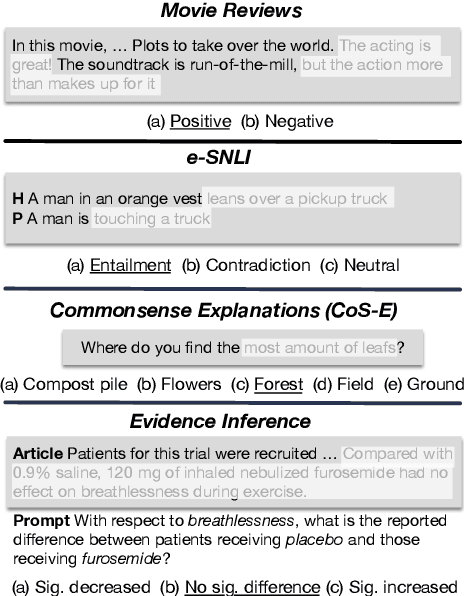
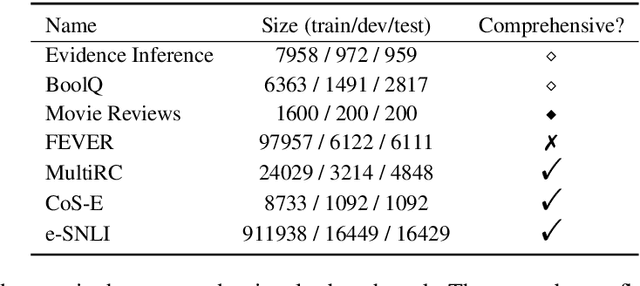
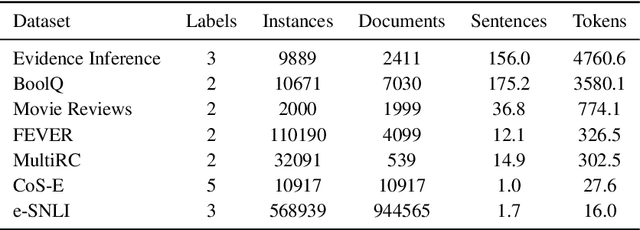
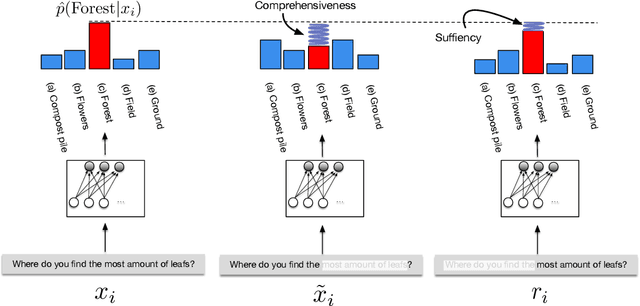
Abstract:State-of-the-art models in NLP are now predominantly based on deep neural networks that are generally opaque in terms of how they come to specific predictions. This limitation has led to increased interest in designing more interpretable deep models for NLP that can reveal the `reasoning' underlying model outputs. But work in this direction has been conducted on different datasets and tasks with correspondingly unique aims and metrics; this makes it difficult to track progress. We propose the Evaluating Rationales And Simple English Reasoning (ERASER) benchmark to advance research on interpretable models in NLP. This benchmark comprises multiple datasets and tasks for which human annotations of "rationales" (supporting evidence) have been collected. We propose several metrics that aim to capture how well the rationales provided by models align with human rationales, and also how faithful these rationales are (i.e., the degree to which provided rationales influenced the corresponding predictions). Our hope is that releasing this benchmark facilitates progress on designing more interpretable NLP systems. The benchmark, code, and documentation are available at: www.eraserbenchmark.com .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge