Iain J. Marshall
Decide less, communicate more: On the construct validity of end-to-end fact-checking in medicine
Jun 25, 2025Abstract:Technological progress has led to concrete advancements in tasks that were regarded as challenging, such as automatic fact-checking. Interest in adopting these systems for public health and medicine has grown due to the high-stakes nature of medical decisions and challenges in critically appraising a vast and diverse medical literature. Evidence-based medicine connects to every individual, and yet the nature of it is highly technical, rendering the medical literacy of majority users inadequate to sufficiently navigate the domain. Such problems with medical communication ripens the ground for end-to-end fact-checking agents: check a claim against current medical literature and return with an evidence-backed verdict. And yet, such systems remain largely unused. To understand this, we present the first study examining how clinical experts verify real claims from social media by synthesizing medical evidence. In searching for this upper-bound, we reveal fundamental challenges in end-to-end fact-checking when applied to medicine: Difficulties connecting claims in the wild to scientific evidence in the form of clinical trials; ambiguities in underspecified claims mixed with mismatched intentions; and inherently subjective veracity labels. We argue that fact-checking should be approached and evaluated as an interactive communication problem, rather than an end-to-end process.
Caught in the Web of Words: Do LLMs Fall for Spin in Medical Literature?
Feb 11, 2025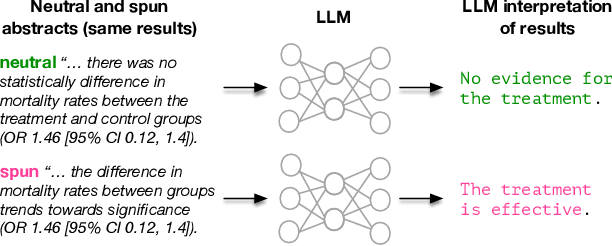
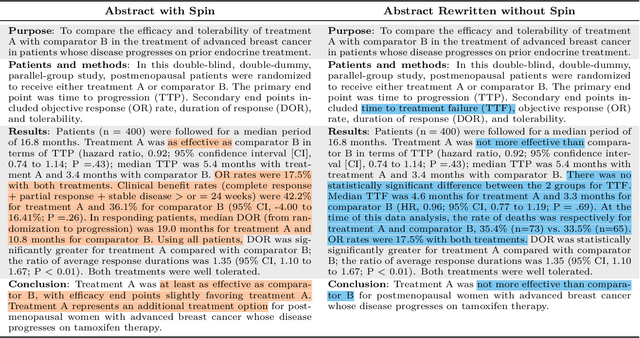
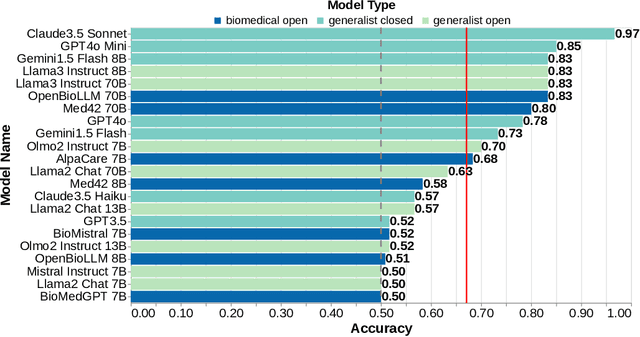
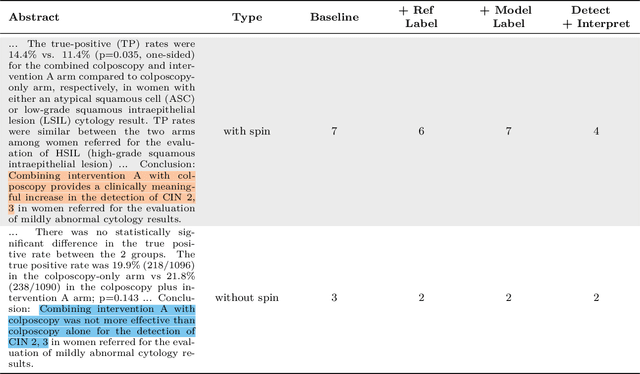
Abstract:Medical research faces well-documented challenges in translating novel treatments into clinical practice. Publishing incentives encourage researchers to present "positive" findings, even when empirical results are equivocal. Consequently, it is well-documented that authors often spin study results, especially in article abstracts. Such spin can influence clinician interpretation of evidence and may affect patient care decisions. In this study, we ask whether the interpretation of trial results offered by Large Language Models (LLMs) is similarly affected by spin. This is important since LLMs are increasingly being used to trawl through and synthesize published medical evidence. We evaluated 22 LLMs and found that they are across the board more susceptible to spin than humans. They might also propagate spin into their outputs: We find evidence, e.g., that LLMs implicitly incorporate spin into plain language summaries that they generate. We also find, however, that LLMs are generally capable of recognizing spin, and can be prompted in a way to mitigate spin's impact on LLM outputs.
RealMedQA: A pilot biomedical question answering dataset containing realistic clinical questions
Aug 16, 2024Abstract:Clinical question answering systems have the potential to provide clinicians with relevant and timely answers to their questions. Nonetheless, despite the advances that have been made, adoption of these systems in clinical settings has been slow. One issue is a lack of question-answering datasets which reflect the real-world needs of health professionals. In this work, we present RealMedQA, a dataset of realistic clinical questions generated by humans and an LLM. We describe the process for generating and verifying the QA pairs and assess several QA models on BioASQ and RealMedQA to assess the relative difficulty of matching answers to questions. We show that the LLM is more cost-efficient for generating "ideal" QA pairs. Additionally, we achieve a lower lexical similarity between questions and answers than BioASQ which provides an additional challenge to the top two QA models, as per the results. We release our code and our dataset publicly to encourage further research.
Automatically Extracting Numerical Results from Randomized Controlled Trials with Large Language Models
May 02, 2024



Abstract:Meta-analyses statistically aggregate the findings of different randomized controlled trials (RCTs) to assess treatment effectiveness. Because this yields robust estimates of treatment effectiveness, results from meta-analyses are considered the strongest form of evidence. However, rigorous evidence syntheses are time-consuming and labor-intensive, requiring manual extraction of data from individual trials to be synthesized. Ideally, language technologies would permit fully automatic meta-analysis, on demand. This requires accurately extracting numerical results from individual trials, which has been beyond the capabilities of natural language processing (NLP) models to date. In this work, we evaluate whether modern large language models (LLMs) can reliably perform this task. We annotate (and release) a modest but granular evaluation dataset of clinical trial reports with numerical findings attached to interventions, comparators, and outcomes. Using this dataset, we evaluate the performance of seven LLMs applied zero-shot for the task of conditionally extracting numerical findings from trial reports. We find that massive LLMs that can accommodate lengthy inputs are tantalizingly close to realizing fully automatic meta-analysis, especially for dichotomous (binary) outcomes (e.g., mortality). However, LLMs -- including ones trained on biomedical texts -- perform poorly when the outcome measures are complex and tallying the results requires inference. This work charts a path toward fully automatic meta-analysis of RCTs via LLMs, while also highlighting the limitations of existing models for this aim.
Appraising the Potential Uses and Harms of LLMs for Medical Systematic Reviews
May 22, 2023



Abstract:Medical systematic reviews are crucial for informing clinical decision making and healthcare policy. But producing such reviews is onerous and time-consuming. Thus, high-quality evidence synopses are not available for many questions and may be outdated even when they are available. Large language models (LLMs) are now capable of generating long-form texts, suggesting the tantalizing possibility of automatically generating literature reviews on demand. However, LLMs sometimes generate inaccurate (and potentially misleading) texts by hallucinating or omitting important information. In the healthcare context, this may render LLMs unusable at best and dangerous at worst. Most discussion surrounding the benefits and risks of LLMs have been divorced from specific applications. In this work, we seek to qualitatively characterize the potential utility and risks of LLMs for assisting in production of medical evidence reviews. We conducted 16 semi-structured interviews with international experts in systematic reviews, grounding discussion in the context of generating evidence reviews. Domain experts indicated that LLMs could aid writing reviews, as a tool for drafting or creating plain language summaries, generating templates or suggestions, distilling information, crosschecking, and synthesizing or interpreting text inputs. But they also identified issues with model outputs and expressed concerns about potential downstream harms of confidently composed but inaccurate LLM outputs which might mislead. Other anticipated potential downstream harms included lessened accountability and proliferation of automatically generated reviews that might be of low quality. Informed by this qualitative analysis, we identify criteria for rigorous evaluation of biomedical LLMs aligned with domain expert views.
Summarizing, Simplifying, and Synthesizing Medical Evidence Using GPT-3 (with Varying Success)
May 11, 2023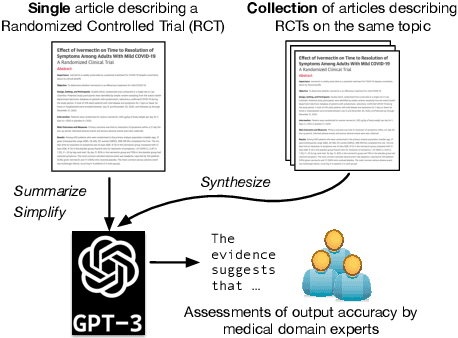

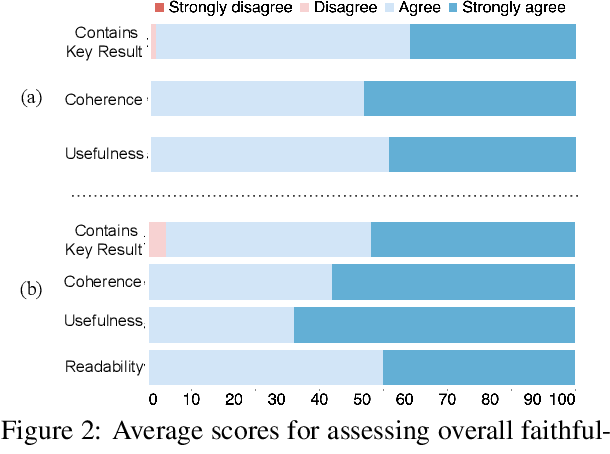

Abstract:Large language models, particularly GPT-3, are able to produce high quality summaries of general domain news articles in few- and zero-shot settings. However, it is unclear if such models are similarly capable in more specialized, high-stakes domains such as biomedicine. In this paper, we enlist domain experts (individuals with medical training) to evaluate summaries of biomedical articles generated by GPT-3, given zero supervision. We consider both single- and multi-document settings. In the former, GPT-3 is tasked with generating regular and plain-language summaries of articles describing randomized controlled trials; in the latter, we assess the degree to which GPT-3 is able to \emph{synthesize} evidence reported across a collection of articles. We design an annotation scheme for evaluating model outputs, with an emphasis on assessing the factual accuracy of generated summaries. We find that while GPT-3 is able to summarize and simplify single biomedical articles faithfully, it struggles to provide accurate aggregations of findings over multiple documents. We release all data and annotations used in this work.
Do Multi-Document Summarization Models Synthesize?
Jan 31, 2023Abstract:Multi-document summarization entails producing concise synopses of collections of inputs. For some applications, the synopsis should accurately \emph{synthesize} inputs with respect to a key property or aspect. For example, a synopsis of film reviews all written about a particular movie should reflect the average critic consensus. As a more consequential example, consider narrative summaries that accompany biomedical \emph{systematic reviews} of clinical trial results. These narratives should fairly summarize the potentially conflicting results from individual trials. In this paper we ask: To what extent do modern multi-document summarization models implicitly perform this type of synthesis? To assess this we perform a suite of experiments that probe the degree to which conditional generation models trained for summarization using standard methods yield outputs that appropriately synthesize inputs. We find that existing models do partially perform synthesis, but do so imperfectly. In particular, they are over-sensitive to changes in input ordering and under-sensitive to changes in input compositions (e.g., the ratio of positive to negative movie reviews). We propose a simple, general method for improving model synthesis capabilities by generating an explicitly diverse set of candidate outputs, and then selecting from these the string best aligned with the expected aggregate measure for the inputs, or \emph{abstaining} when the model produces no good candidate. This approach improves model synthesis performance. We hope highlighting the need for synthesis (in some summarization settings), motivates further research into multi-document summarization methods and learning objectives that explicitly account for the need to synthesize.
What Would it Take to get Biomedical QA Systems into Practice?
Sep 21, 2021



Abstract:Medical question answering (QA) systems have the potential to answer clinicians uncertainties about treatment and diagnosis on demand, informed by the latest evidence. However, despite the significant progress in general QA made by the NLP community, medical QA systems are still not widely used in clinical environments. One likely reason for this is that clinicians may not readily trust QA system outputs, in part because transparency, trustworthiness, and provenance have not been key considerations in the design of such models. In this paper we discuss a set of criteria that, if met, we argue would likely increase the utility of biomedical QA systems, which may in turn lead to adoption of such systems in practice. We assess existing models, tasks, and datasets with respect to these criteria, highlighting shortcomings of previously proposed approaches and pointing toward what might be more usable QA systems.
Paragraph-level Simplification of Medical Texts
Apr 12, 2021



Abstract:We consider the problem of learning to simplify medical texts. This is important because most reliable, up-to-date information in biomedicine is dense with jargon and thus practically inaccessible to the lay audience. Furthermore, manual simplification does not scale to the rapidly growing body of biomedical literature, motivating the need for automated approaches. Unfortunately, there are no large-scale resources available for this task. In this work we introduce a new corpus of parallel texts in English comprising technical and lay summaries of all published evidence pertaining to different clinical topics. We then propose a new metric based on likelihood scores from a masked language model pretrained on scientific texts. We show that this automated measure better differentiates between technical and lay summaries than existing heuristics. We introduce and evaluate baseline encoder-decoder Transformer models for simplification and propose a novel augmentation to these in which we explicitly penalize the decoder for producing "jargon" terms; we find that this yields improvements over baselines in terms of readability.
Understanding Clinical Trial Reports: Extracting Medical Entities and Their Relations
Oct 08, 2020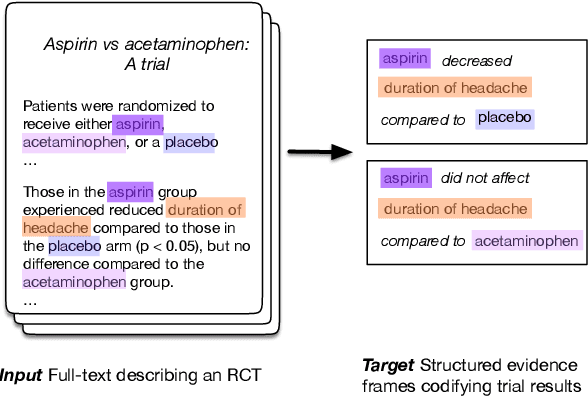
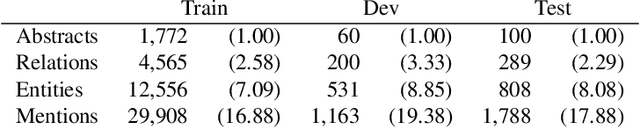
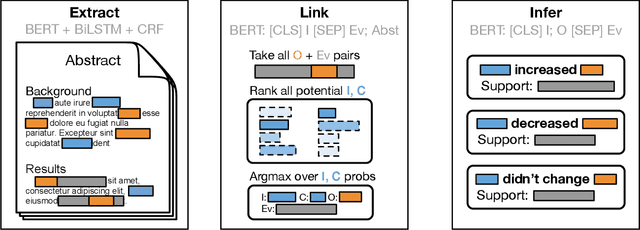

Abstract:The best evidence concerning comparative treatment effectiveness comes from clinical trials, the results of which are reported in unstructured articles. Medical experts must manually extract information from articles to inform decision-making, which is time-consuming and expensive. Here we consider the end-to-end task of both (a) extracting treatments and outcomes from full-text articles describing clinical trials (entity identification) and, (b) inferring the reported results for the former with respect to the latter (relation extraction). We introduce new data for this task, and evaluate models that have recently achieved state-of-the-art results on similar tasks in Natural Language Processing. We then propose a new method motivated by how trial results are typically presented that outperforms these purely data-driven baselines. Finally, we run a fielded evaluation of the model with a non-profit seeking to identify existing drugs that might be re-purposed for cancer, showing the potential utility of end-to-end evidence extraction systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge