Varun Buch
Development and Validation of a Deep Learning Model for Prediction of Severe Outcomes in Suspected COVID-19 Infection
Mar 29, 2021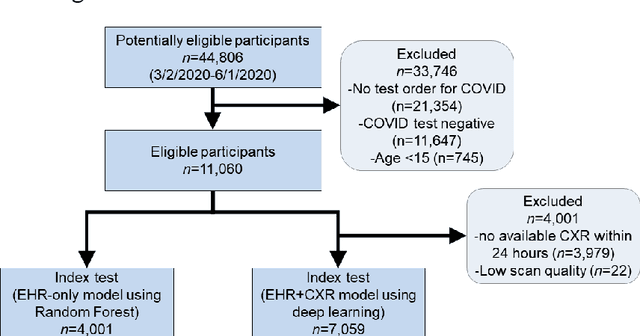

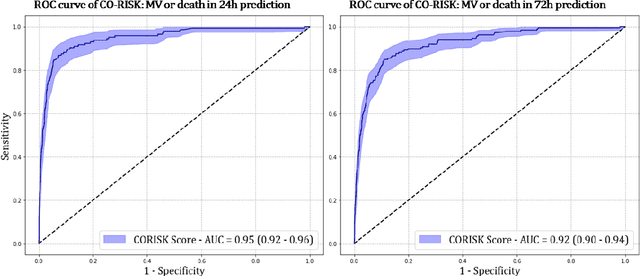
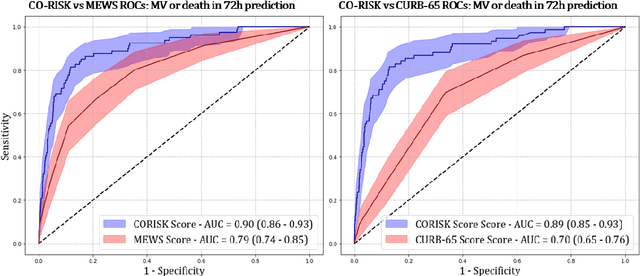
Abstract:COVID-19 patient triaging with predictive outcome of the patients upon first present to emergency department (ED) is crucial for improving patient prognosis, as well as better hospital resources management and cross-infection control. We trained a deep feature fusion model to predict patient outcomes, where the model inputs were EHR data including demographic information, co-morbidities, vital signs and laboratory measurements, plus patient's CXR images. The model output was patient outcomes defined as the most insensitive oxygen therapy required. For patients without CXR images, we employed Random Forest method for the prediction. Predictive risk scores for COVID-19 severe outcomes ("CO-RISK" score) were derived from model output and evaluated on the testing dataset, as well as compared to human performance. The study's dataset (the "MGB COVID Cohort") was constructed from all patients presenting to the Mass General Brigham (MGB) healthcare system from March 1st to June 1st, 2020. ED visits with incomplete or erroneous data were excluded. Patients with no test order for COVID or confirmed negative test results were excluded. Patients under the age of 15 were also excluded. Finally, electronic health record (EHR) data from a total of 11060 COVID-19 confirmed or suspected patients were used in this study. Chest X-ray (CXR) images were also collected from each patient if available. Results show that CO-RISK score achieved area under the Curve (AUC) of predicting MV/death (i.e. severe outcomes) in 24 hours of 0.95, and 0.92 in 72 hours on the testing dataset. The model shows superior performance to the commonly used risk scores in ED (CURB-65 and MEWS). Comparing with physician's decisions, CO-RISK score has demonstrated superior performance to human in making ICU/floor decisions.
Deep Metric Learning-based Image Retrieval System for Chest Radiograph and its Clinical Applications in COVID-19
Nov 26, 2020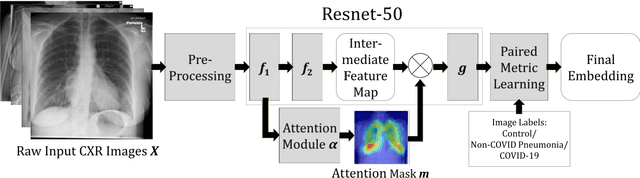
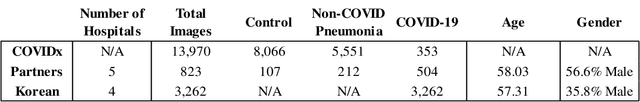
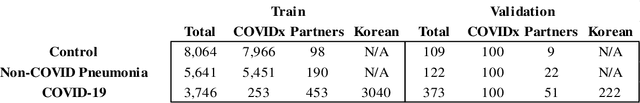
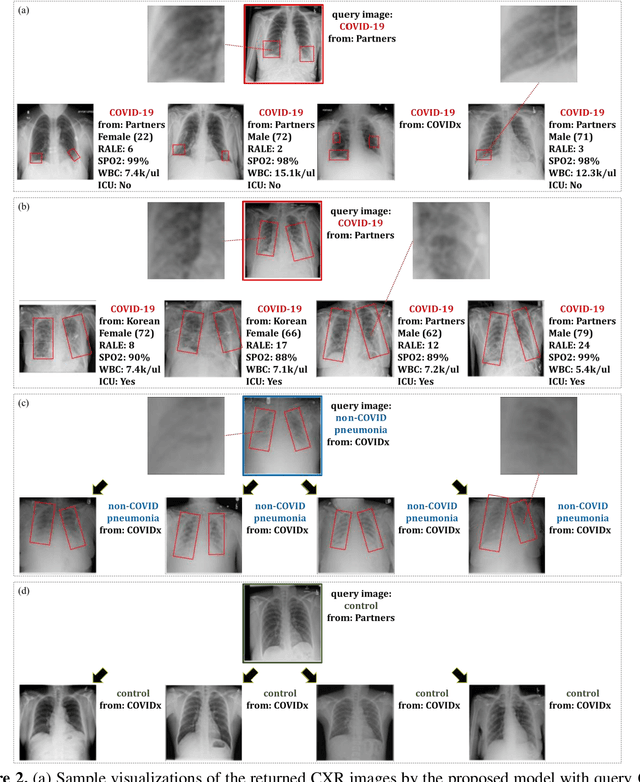
Abstract:In recent years, deep learning-based image analysis methods have been widely applied in computer-aided detection, diagnosis and prognosis, and has shown its value during the public health crisis of the novel coronavirus disease 2019 (COVID-19) pandemic. Chest radiograph (CXR) has been playing a crucial role in COVID-19 patient triaging, diagnosing and monitoring, particularly in the United States. Considering the mixed and unspecific signals in CXR, an image retrieval model of CXR that provides both similar images and associated clinical information can be more clinically meaningful than a direct image diagnostic model. In this work we develop a novel CXR image retrieval model based on deep metric learning. Unlike traditional diagnostic models which aims at learning the direct mapping from images to labels, the proposed model aims at learning the optimized embedding space of images, where images with the same labels and similar contents are pulled together. It utilizes multi-similarity loss with hard-mining sampling strategy and attention mechanism to learn the optimized embedding space, and provides similar images to the query image. The model is trained and validated on an international multi-site COVID-19 dataset collected from 3 different sources. Experimental results of COVID-19 image retrieval and diagnosis tasks show that the proposed model can serve as a robust solution for CXR analysis and patient management for COVID-19. The model is also tested on its transferability on a different clinical decision support task, where the pre-trained model is applied to extract image features from a new dataset without any further training. These results demonstrate our deep metric learning based image retrieval model is highly efficient in the CXR retrieval, diagnosis and prognosis, and thus has great clinical value for the treatment and management of COVID-19 patients.
Federated Learning for Breast Density Classification: A Real-World Implementation
Sep 17, 2020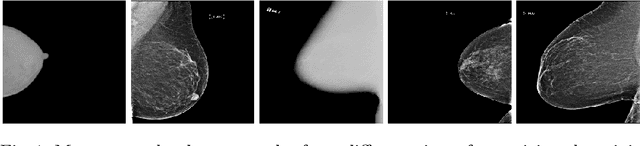
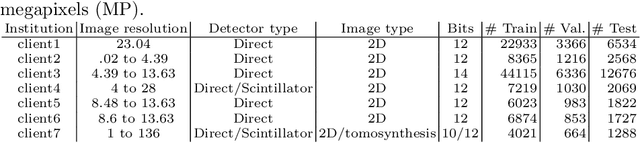

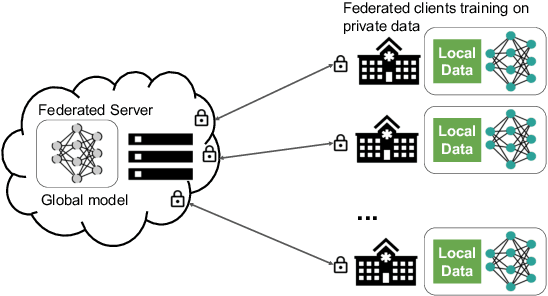
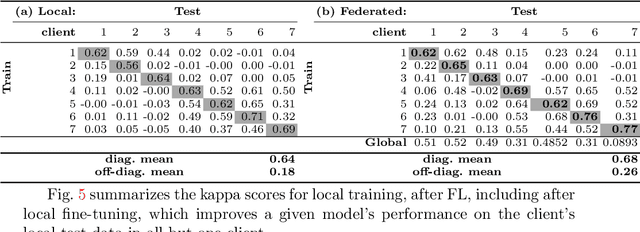
Abstract:Building robust deep learning-based models requires large quantities of diverse training data. In this study, we investigate the use of federated learning (FL) to build medical imaging classification models in a real-world collaborative setting. Seven clinical institutions from across the world joined this FL effort to train a model for breast density classification based on Breast Imaging, Reporting & Data System (BI-RADS). We show that despite substantial differences among the datasets from all sites (mammography system, class distribution, and data set size) and without centralizing data, we can successfully train AI models in federation. The results show that models trained using FL perform 6.3% on average better than their counterparts trained on an institute's local data alone. Furthermore, we show a 45.8% relative improvement in the models' generalizability when evaluated on the other participating sites' testing data.
An Overview and Case Study of the Clinical AI Model Development Life Cycle for Healthcare Systems
Mar 26, 2020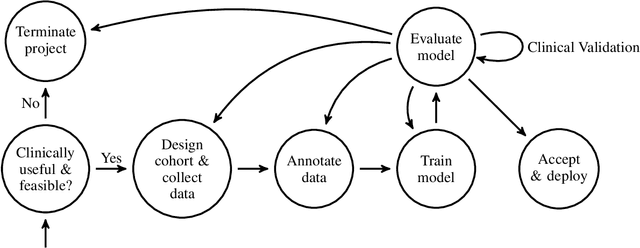
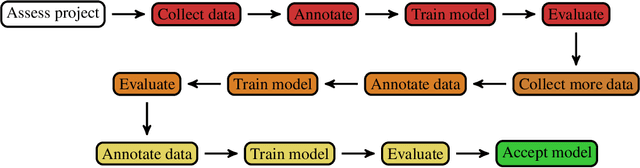
Abstract:Healthcare is one of the most promising areas for machine learning models to make a positive impact. However, successful adoption of AI-based systems in healthcare depends on engaging and educating stakeholders from diverse backgrounds about the development process of AI models. We present a broadly accessible overview of the development life cycle of clinical AI models that is general enough to be adapted to most machine learning projects, and then give an in-depth case study of the development process of a deep learning based system to detect aortic aneurysms in Computed Tomography (CT) exams. We hope other healthcare institutions and clinical practitioners find the insights we share about the development process useful in informing their own model development efforts and to increase the likelihood of successful deployment and integration of AI in healthcare.
Semi-Supervised Natural Language Approach for Fine-Grained Classification of Medical Reports
Nov 14, 2019
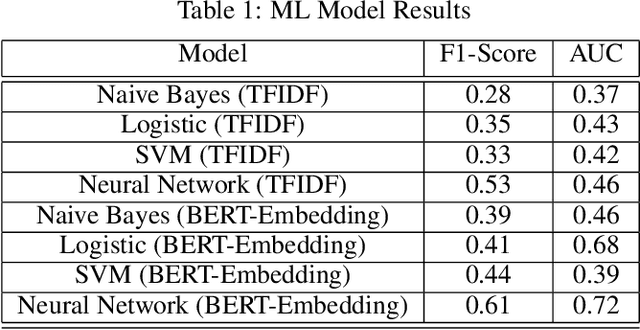
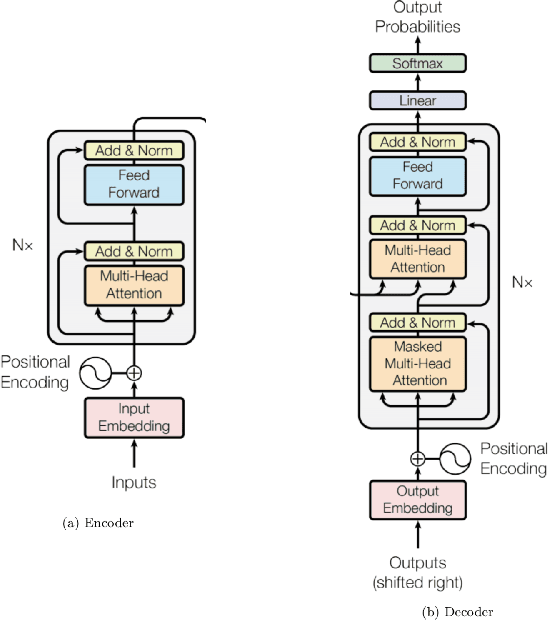
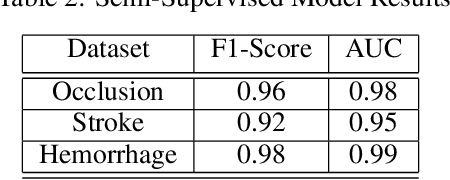
Abstract:Although machine learning has become a powerful tool to augment doctors in clinical analysis, the immense amount of labeled data that is necessary to train supervised learning approaches burdens each development task as time and resource intensive. The vast majority of dense clinical information is stored in written reports, detailing pertinent patient information. The challenge with utilizing natural language data for standard model development is due to the complex nature of the modality. In this research, a model pipeline was developed to utilize an unsupervised approach to train an encoder-language model, a recurrent network, to generate document encodings; which then can be used as features passed into a decoder-classifier model that requires magnitudes less labeled data than previous approaches to differentiate between fine-grained disease classes accurately. The language model was trained on unlabeled radiology reports from the Massachusetts General Hospital Radiology Department (n=218,159) and terminated with a loss of 1.62. The classification models were trained on three labeled datasets of head CT studies of reported patients, presenting large vessel occlusion (n=1403), acute ischemic strokes (n=331), and intracranial hemorrhage (n=4350), to identify a variety of different findings directly from the radiology report data; resulting in AUCs of 0.98, 0.95, and 0.99, respectively, for the large vessel occlusion, acute ischemic stroke, and intracranial hemorrhage datasets. The output encodings are able to be used in conjunction with imaging data, to create models that can process a multitude of different modalities. The ability to automatically extract relevant features from textual data allows for faster model development and integration of textual modality, overall, allowing clinical reports to become a more viable input for more encompassing and accurate deep learning models.
DeepAAA: clinically applicable and generalizable detection of abdominal aortic aneurysm using deep learning
Jul 04, 2019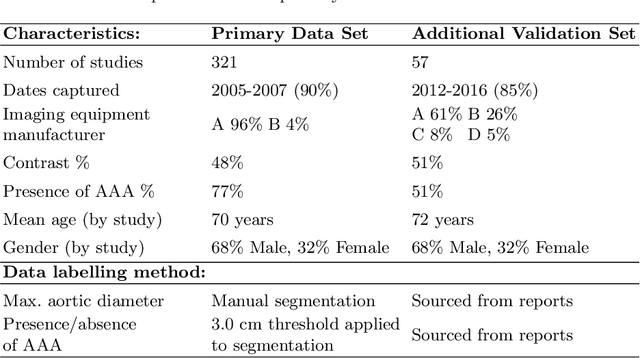
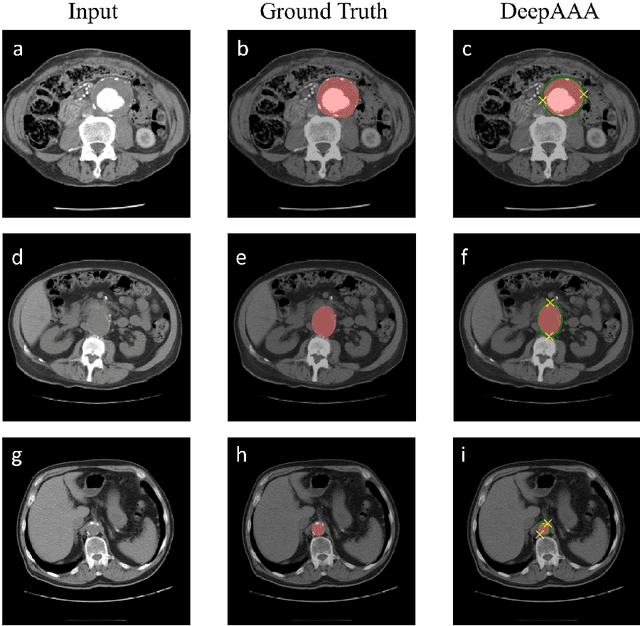
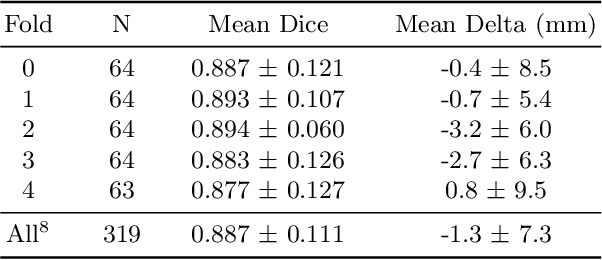
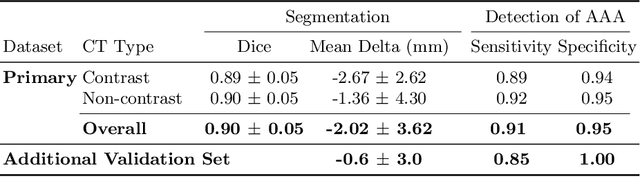
Abstract:We propose a deep learning-based technique for detection and quantification of abdominal aortic aneurysms (AAAs). The condition, which leads to more than 10,000 deaths per year in the United States, is asymptomatic, often detected incidentally, and often missed by radiologists. Our model architecture is a modified 3D U-Net combined with ellipse fitting that performs aorta segmentation and AAA detection. The study uses 321 abdominal-pelvic CT examinations performed by Massachusetts General Hospital Department of Radiology for training and validation. The model is then further tested for generalizability on a separate set of 57 examinations with differing patient demographics and acquisition characteristics than the original dataset. DeepAAA achieves high performance on both sets of data (sensitivity/specificity 0.91/0.95 and 0.85 / 1.0 respectively), on contrast and non-contrast CT scans and works with image volumes with varying numbers of images. We find that DeepAAA exceeds literature-reported performance of radiologists on incidental AAA detection. It is expected that the model can serve as an effective background detector in routine CT examinations to prevent incidental AAAs from being missed.
4D CNN for semantic segmentation of cardiac volumetric sequences
Jun 17, 2019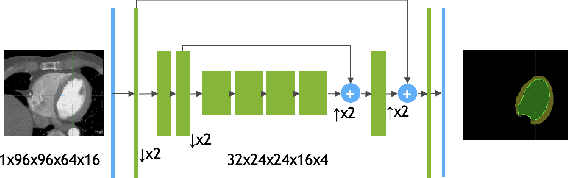
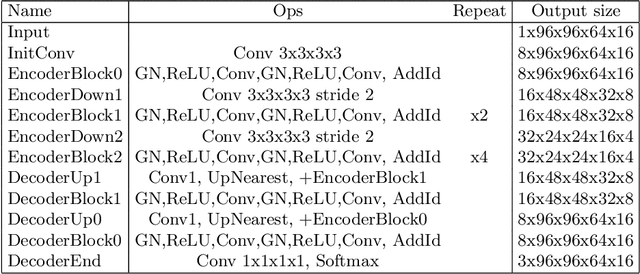
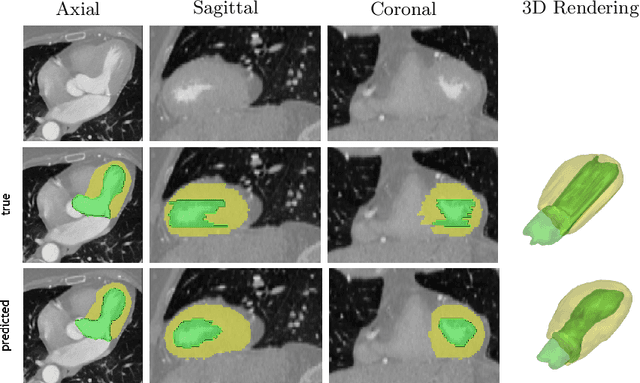
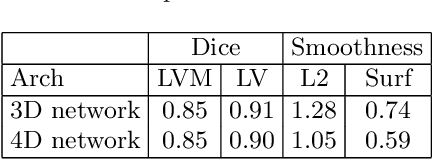
Abstract:We propose a 4D convolutional neural network (CNN) for the segmentation of retrospective ECG-gated cardiac CT, a series of single-channel volumetric data over time. While only a small subset of volumes in the temporal sequence are annotated, we define a sparse loss function on available labels to allow the network to leverage unlabeled images during training and generate a fully segmented sequence. We investigate the accuracy of the proposed 4D network to predict temporally consistent segmentations and compare with traditional 3D segmentation approaches. We demonstrate the feasibility of the 4D CNN and establish its performance on cardiac 4D CCTA.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge