Shujian Yu
Modeling Higher-Order Brain Interactions via a Multi-View Information Bottleneck Framework for fMRI-based Psychiatric Diagnosis
Apr 20, 2026Abstract:Resting-state functional magnetic resonance imaging (fMRI) has emerged as a cornerstone for psychiatric diagnosis, yet most approaches rely on pairwise brain cortical or sub-cortical connectivities that overlooks higher-order interactions (HOIs) central to complex brain dynamics. While hypergraph methods encode HOIs through predefined hyperedges, their construction typically relies on heuristic similarity metrics and does not explicitly characterize whether interactions are synergy- or redundancy-dominated. In this paper, we introduce $O$-information, a signed measure that characterizes the informational nature of HOIs, and integrate third- and fourth-order $O$-information into a unified multi-view information bottleneck framework for fMRI-based psychiatric diagnosis. To enable scalable $O$-information estimation, we further develop two independent acceleration strategies: a Gaussian analytical approximation and a randomized matrix-based Rényi entropy estimator, achieving over a 30-fold computational speedup compared with conventional estimators. Our tri-view architecture systematically fuses pairwise, triadic, and tetradic brain interactions, capturing comprehensive brain connectivity while explicitly penalizing redundancy. Extensive evaluation across four benchmark datasets (REST-meta-MDD, ABIDE, UCLA, ADNI) demonstrates consistent improvements, outperforming 11 baseline methods including state-of-the-art graph neural network (GNN) and hypergraph based approaches. Moreover, our method reveals interpretable region-level synergy-redundancy patterns which are not explicitly characterized by conventional hypergraph formulations.
Continual Learning for fMRI-Based Brain Disorder Diagnosis via Functional Connectivity Matrices Generative Replay
Apr 15, 2026Abstract:Functional magnetic resonance imaging (fMRI) is widely used for studying and diagnosing brain disorders, with functional connectivity (FC) matrices providing powerful representations of large-scale neural interactions. However, existing diagnostic models are trained either on a single site or under full multi-site access, making them unsuitable for real-world scenarios where clinical data arrive sequentially from different institutions. This results in limited generalization and severe catastrophic forgetting. This paper presents the first continual learning framework specifically designed for fMRI-based diagnosis across heterogeneous clinical sites. Our framework introduces a structure-aware variational autoencoder that synthesizes realistic FC matrices for both patient and control groups. Built on this generative backbone, we develop a multi-level knowledge distillation strategy that aligns predictions and graph representations between new-site data and replayed samples. To further enhance efficiency, we incorporate a hierarchical contextual bandit scheme for adaptive replay sampling. Experiments on multi-site datasets for major depressive disorder (MDD), schizophrenia (SZ), and autism spectrum disorder (ASD) show that the proposed generative model enhances data augmentation quality, and the overall continual learning framework substantially outperforms existing methods in mitigating catastrophic forgetting. Our code is available at https://github.com/4me808/FORGE.
PIME: Prototype-based Interpretable MCTS-Enhanced Brain Network Analysis for Disorder Diagnosis
Feb 24, 2026Abstract:Recent deep learning methods for fMRI-based diagnosis have achieved promising accuracy by modeling functional connectivity networks. However, standard approaches often struggle with noisy interactions, and conventional post-hoc attribution methods may lack reliability, potentially highlighting dataset-specific artifacts. To address these challenges, we introduce PIME, an interpretable framework that bridges intrinsic interpretability with minimal-sufficient subgraph optimization by integrating prototype-based classification and consistency training with structural perturbations during learning. This encourages a structured latent space and enables Monte Carlo Tree Search (MCTS) under a prototype-consistent objective to extract compact minimal-sufficient explanatory subgraphs post-training. Experiments on three benchmark fMRI datasets demonstrate that PIME achieves state-of-the-art performance. Furthermore, by constraining the search space via learned prototypes, PIME identifies critical brain regions that are consistent with established neuroimaging findings. Stability analysis shows 90% reproducibility and consistent explanations across atlases.
Towards Uniformity and Alignment for Multimodal Representation Learning
Feb 10, 2026Abstract:Multimodal representation learning aims to construct a shared embedding space in which heterogeneous modalities are semantically aligned. Despite strong empirical results, InfoNCE-based objectives introduce inherent conflicts that yield distribution gaps across modalities. In this work, we identify two conflicts in the multimodal regime, both exacerbated as the number of modalities increases: (i) an alignment-uniformity conflict, whereby the repulsion of uniformity undermines pairwise alignment, and (ii) an intra-alignment conflict, where aligning multiple modalities induces competing alignment directions. To address these issues, we propose a principled decoupling of alignment and uniformity for multimodal representations, providing a conflict-free recipe for multimodal learning that simultaneously supports discriminative and generative use cases without task-specific modules. We then provide a theoretical guarantee that our method acts as an efficient proxy for a global Hölder divergence over multiple modality distributions, and thus reduces the distribution gap among modalities. Extensive experiments on retrieval and UnCLIP-style generation demonstrate consistent gains.
HFMCA: Orthonormal Feature Learning for EEG-based Brain Decoding
Feb 04, 2026Abstract:Electroencephalography (EEG) analysis is critical for brain-computer interfaces and neuroscience, but the intrinsic noise and high dimensionality of EEG signals hinder effective feature learning. We propose a self-supervised framework based on the Hierarchical Functional Maximal Correlation Algorithm (HFMCA), which learns orthonormal EEG representations by enforcing feature decorrelation and reducing redundancy. This design enables robust capture of essential brain dynamics for various EEG recognition tasks. We validate HFMCA on two benchmark datasets, SEED and BCIC-2A, where pretraining with HFMCA consistently outperforms competitive self-supervised baselines, achieving notable gains in classification accuracy. Across diverse EEG tasks, our method demonstrates superior cross-subject generalization under leave-one-subject-out validation, advancing state-of-the-art by 2.71\% on SEED emotion recognition and 2.57\% on BCIC-2A motor imagery classification.
Deep Deterministic Nonlinear ICA via Total Correlation Minimization with Matrix-Based Entropy Functional
Dec 31, 2025Abstract:Blind source separation, particularly through independent component analysis (ICA), is widely utilized across various signal processing domains for disentangling underlying components from observed mixed signals, owing to its fully data-driven nature that minimizes reliance on prior assumptions. However, conventional ICA methods rely on an assumption of linear mixing, limiting their ability to capture complex nonlinear relationships and to maintain robustness in noisy environments. In this work, we present deep deterministic nonlinear independent component analysis (DDICA), a novel deep neural network-based framework designed to address these limitations. DDICA leverages a matrix-based entropy function to directly optimize the independence criterion via stochastic gradient descent, bypassing the need for variational approximations or adversarial schemes. This results in a streamlined training process and improved resilience to noise. We validated the effectiveness and generalizability of DDICA across a range of applications, including simulated signal mixtures, hyperspectral image unmixing, modeling of primary visual receptive fields, and resting-state functional magnetic resonance imaging (fMRI) data analysis. Experimental results demonstrate that DDICA effectively separates independent components with high accuracy across a range of applications. These findings suggest that DDICA offers a robust and versatile solution for blind source separation in diverse signal processing tasks.
Multimodal Functional Maximum Correlation for Emotion Recognition
Dec 28, 2025Abstract:Emotional states manifest as coordinated yet heterogeneous physiological responses across central and autonomic systems, posing a fundamental challenge for multimodal representation learning in affective computing. Learning such joint dynamics is further complicated by the scarcity and subjectivity of affective annotations, which motivates the use of self-supervised learning (SSL). However, most existing SSL approaches rely on pairwise alignment objectives, which are insufficient to characterize dependencies among more than two modalities and fail to capture higher-order interactions arising from coordinated brain and autonomic responses. To address this limitation, we propose Multimodal Functional Maximum Correlation (MFMC), a principled SSL framework that maximizes higher-order multimodal dependence through a Dual Total Correlation (DTC) objective. By deriving a tight sandwich bound and optimizing it using a functional maximum correlation analysis (FMCA) based trace surrogate, MFMC captures joint multimodal interactions directly, without relying on pairwise contrastive losses. Experiments on three public affective computing benchmarks demonstrate that MFMC consistently achieves state-of-the-art or competitive performance under both subject-dependent and subject-independent evaluation protocols, highlighting its robustness to inter-subject variability. In particular, MFMC improves subject-dependent accuracy on CEAP-360VR from 78.9% to 86.8%, and subject-independent accuracy from 27.5% to 33.1% using the EDA signal alone. Moreover, MFMC remains within 0.8 percentage points of the best-performing method on the most challenging EEG subject-independent split of MAHNOB-HCI. Our code is available at https://github.com/DY9910/MFMC.
Explainable Multimodal Regression via Information Decomposition
Dec 26, 2025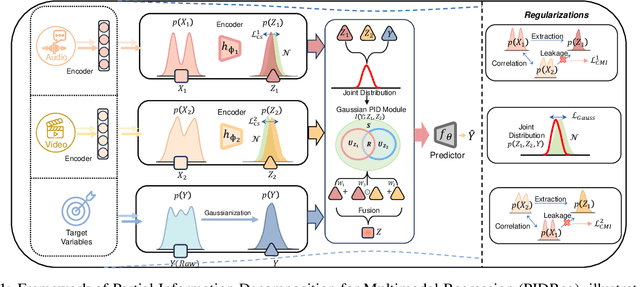

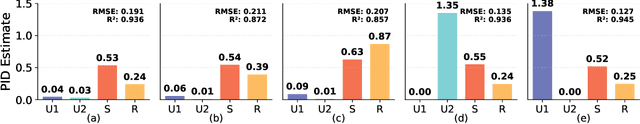
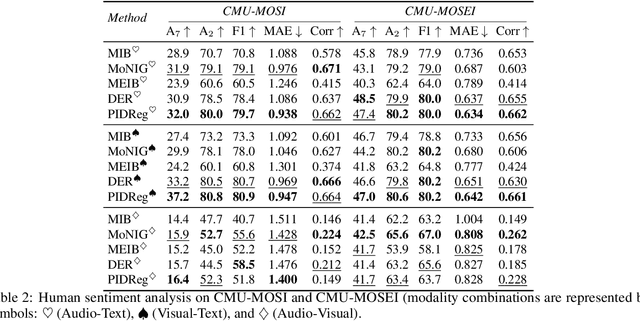
Abstract:Multimodal regression aims to predict a continuous target from heterogeneous input sources and typically relies on fusion strategies such as early or late fusion. However, existing methods lack principled tools to disentangle and quantify the individual contributions of each modality and their interactions, limiting the interpretability of multimodal fusion. We propose a novel multimodal regression framework grounded in Partial Information Decomposition (PID), which decomposes modality-specific representations into unique, redundant, and synergistic components. The basic PID framework is inherently underdetermined. To resolve this, we introduce inductive bias by enforcing Gaussianity in the joint distribution of latent representations and the transformed response variable (after inverse normal transformation), thereby enabling analytical computation of the PID terms. Additionally, we derive a closed-form conditional independence regularizer to promote the isolation of unique information within each modality. Experiments on six real-world datasets, including a case study on large-scale brain age prediction from multimodal neuroimaging data, demonstrate that our framework outperforms state-of-the-art methods in both predictive accuracy and interpretability, while also enabling informed modality selection for efficient inference. Implementation is available at https://github.com/zhaozhaoma/PIDReg.
When Brain Foundation Model Meets Cauchy-Schwarz Divergence: A New Framework for Cross-Subject Motor Imagery Decoding
Jul 28, 2025Abstract:Decoding motor imagery (MI) electroencephalogram (EEG) signals, a key non-invasive brain-computer interface (BCI) paradigm for controlling external systems, has been significantly advanced by deep learning. However, MI-EEG decoding remains challenging due to substantial inter-subject variability and limited labeled target data, which necessitate costly calibration for new users. Many existing multi-source domain adaptation (MSDA) methods indiscriminately incorporate all available source domains, disregarding the large inter-subject differences in EEG signals, which leads to negative transfer and excessive computational costs. Moreover, while many approaches focus on feature distribution alignment, they often neglect the explicit dependence between features and decision-level outputs, limiting their ability to preserve discriminative structures. To address these gaps, we propose a novel MSDA framework that leverages a pretrained large Brain Foundation Model (BFM) for dynamic and informed source subject selection, ensuring only relevant sources contribute to adaptation. Furthermore, we employ Cauchy-Schwarz (CS) and Conditional CS (CCS) divergences to jointly perform feature-level and decision-level alignment, enhancing domain invariance while maintaining class discriminability. Extensive evaluations on two benchmark MI-EEG datasets demonstrate that our framework outperforms a broad range of state-of-the-art baselines. Additional experiments with a large source pool validate the scalability and efficiency of BFM-guided selection, which significantly reduces training time without sacrificing performance.
MvHo-IB: Multi-View Higher-Order Information Bottleneck for Brain Disorder Diagnosis
Jul 03, 2025Abstract:Recent evidence suggests that modeling higher-order interactions (HOIs) in functional magnetic resonance imaging (fMRI) data can enhance the diagnostic accuracy of machine learning systems. However, effectively extracting and utilizing HOIs remains a significant challenge. In this work, we propose MvHo-IB, a novel multi-view learning framework that integrates both pairwise interactions and HOIs for diagnostic decision-making, while automatically compressing task-irrelevant redundant information. MvHo-IB introduces several key innovations: (1) a principled method that combines O-information from information theory with a matrix-based Renyi alpha-order entropy estimator to quantify and extract HOIs, (2) a purpose-built Brain3DCNN encoder to effectively utilize these interactions, and (3) a new multi-view learning information bottleneck objective to enhance representation learning. Experiments on three benchmark fMRI datasets demonstrate that MvHo-IB achieves state-of-the-art performance, significantly outperforming previous methods, including recent hypergraph-based techniques. The implementation of MvHo-IB is available at https://github.com/zky04/MvHo-IB.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge