Shijie Huang
Benchmarking Retrieval-Augmented Multimomal Generation for Document Question Answering
May 22, 2025
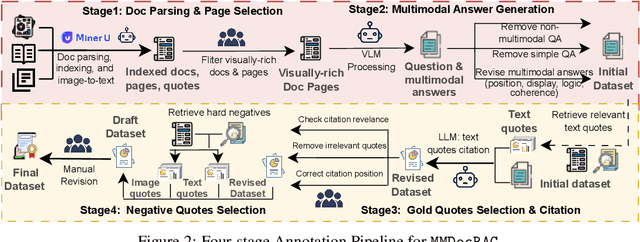
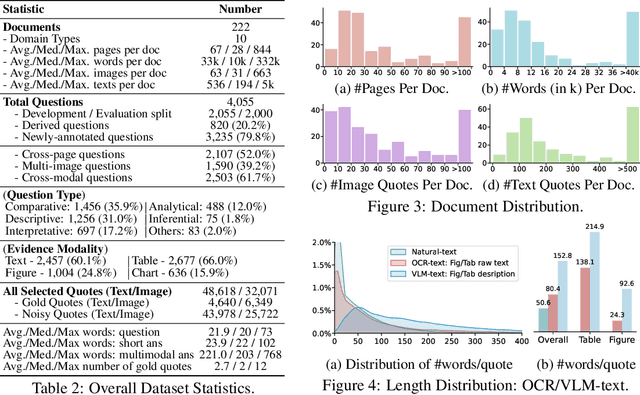
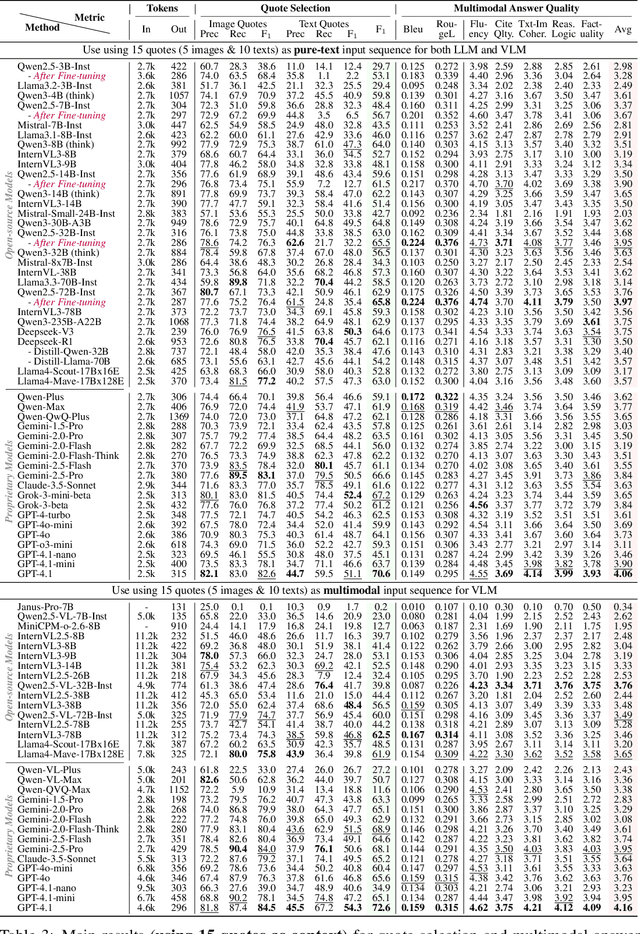
Abstract:Document Visual Question Answering (DocVQA) faces dual challenges in processing lengthy multimodal documents (text, images, tables) and performing cross-modal reasoning. Current document retrieval-augmented generation (DocRAG) methods remain limited by their text-centric approaches, frequently missing critical visual information. The field also lacks robust benchmarks for assessing multimodal evidence selection and integration. We introduce MMDocRAG, a comprehensive benchmark featuring 4,055 expert-annotated QA pairs with multi-page, cross-modal evidence chains. Our framework introduces innovative metrics for evaluating multimodal quote selection and enables answers that interleave text with relevant visual elements. Through large-scale experiments with 60 VLM/LLM models and 14 retrieval systems, we identify persistent challenges in multimodal evidence retrieval, selection, and integration.Key findings reveal advanced proprietary LVMs show superior performance than open-sourced alternatives. Also, they show moderate advantages using multimodal inputs over text-only inputs, while open-source alternatives show significant performance degradation. Notably, fine-tuned LLMs achieve substantial improvements when using detailed image descriptions. MMDocRAG establishes a rigorous testing ground and provides actionable insights for developing more robust multimodal DocVQA systems. Our benchmark and code are available at https://mmdocrag.github.io/MMDocRAG/.
Open-Sora 2.0: Training a Commercial-Level Video Generation Model in $200k
Mar 12, 2025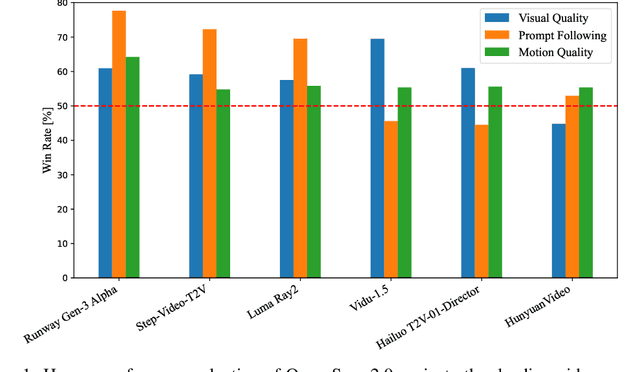
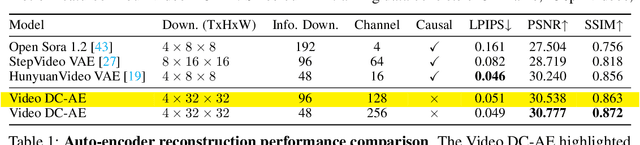

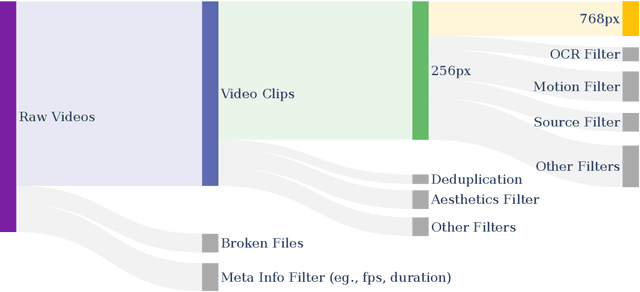
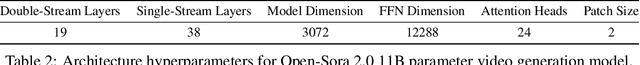
Abstract:Video generation models have achieved remarkable progress in the past year. The quality of AI video continues to improve, but at the cost of larger model size, increased data quantity, and greater demand for training compute. In this report, we present Open-Sora 2.0, a commercial-level video generation model trained for only $200k. With this model, we demonstrate that the cost of training a top-performing video generation model is highly controllable. We detail all techniques that contribute to this efficiency breakthrough, including data curation, model architecture, training strategy, and system optimization. According to human evaluation results and VBench scores, Open-Sora 2.0 is comparable to global leading video generation models including the open-source HunyuanVideo and the closed-source Runway Gen-3 Alpha. By making Open-Sora 2.0 fully open-source, we aim to democratize access to advanced video generation technology, fostering broader innovation and creativity in content creation. All resources are publicly available at: https://github.com/hpcaitech/Open-Sora.
PhotoDoodle: Learning Artistic Image Editing from Few-Shot Pairwise Data
Feb 23, 2025Abstract:We introduce PhotoDoodle, a novel image editing framework designed to facilitate photo doodling by enabling artists to overlay decorative elements onto photographs. Photo doodling is challenging because the inserted elements must appear seamlessly integrated with the background, requiring realistic blending, perspective alignment, and contextual coherence. Additionally, the background must be preserved without distortion, and the artist's unique style must be captured efficiently from limited training data. These requirements are not addressed by previous methods that primarily focus on global style transfer or regional inpainting. The proposed method, PhotoDoodle, employs a two-stage training strategy. Initially, we train a general-purpose image editing model, OmniEditor, using large-scale data. Subsequently, we fine-tune this model with EditLoRA using a small, artist-curated dataset of before-and-after image pairs to capture distinct editing styles and techniques. To enhance consistency in the generated results, we introduce a positional encoding reuse mechanism. Additionally, we release a PhotoDoodle dataset featuring six high-quality styles. Extensive experiments demonstrate the advanced performance and robustness of our method in customized image editing, opening new possibilities for artistic creation.
DVasMesh: Deep Structured Mesh Reconstruction from Vascular Images for Dynamics Modeling of Vessels
Dec 01, 2024



Abstract:Vessel dynamics simulation is vital in studying the relationship between geometry and vascular disease progression. Reliable dynamics simulation relies on high-quality vascular meshes. Most of the existing mesh generation methods highly depend on manual annotation, which is time-consuming and laborious, usually facing challenges such as branch merging and vessel disconnection. This will hinder vessel dynamics simulation, especially for the population study. To address this issue, we propose a deep learning-based method, dubbed as DVasMesh to directly generate structured hexahedral vascular meshes from vascular images. Our contributions are threefold. First, we propose to formally formulate each vertex of the vascular graph by a four-element vector, including coordinates of the centerline point and the radius. Second, a vectorized graph template is employed to guide DVasMesh to estimate the vascular graph. Specifically, we introduce a sampling operator, which samples the extracted features of the vascular image (by a segmentation network) according to the vertices in the template graph. Third, we employ a graph convolution network (GCN) and take the sampled features as nodes to estimate the deformation between vertices of the template graph and target graph, and the deformed graph template is used to build the mesh. Taking advantage of end-to-end learning and discarding direct dependency on annotated labels, our DVasMesh demonstrates outstanding performance in generating structured vascular meshes on cardiac and cerebral vascular images. It shows great potential for clinical applications by reducing mesh generation time from 2 hours (manual) to 30 seconds (automatic).
ProcessPainter: Learn Painting Process from Sequence Data
Jun 10, 2024



Abstract:The painting process of artists is inherently stepwise and varies significantly among different painters and styles. Generating detailed, step-by-step painting processes is essential for art education and research, yet remains largely underexplored. Traditional stroke-based rendering methods break down images into sequences of brushstrokes, yet they fall short of replicating the authentic processes of artists, with limitations confined to basic brushstroke modifications. Text-to-image models utilizing diffusion processes generate images through iterative denoising, also diverge substantially from artists' painting process. To address these challenges, we introduce ProcessPainter, a text-to-video model that is initially pre-trained on synthetic data and subsequently fine-tuned with a select set of artists' painting sequences using the LoRA model. This approach successfully generates painting processes from text prompts for the first time. Furthermore, we introduce an Artwork Replication Network capable of accepting arbitrary-frame input, which facilitates the controlled generation of painting processes, decomposing images into painting sequences, and completing semi-finished artworks. This paper offers new perspectives and tools for advancing art education and image generation technology.
Tissue Segmentation of Thick-Slice Fetal Brain MR Scans with Guidance from High-Quality Isotropic Volumes
Aug 13, 2023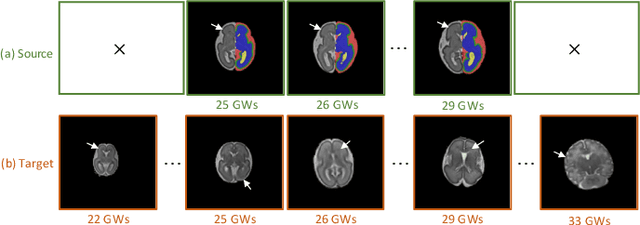
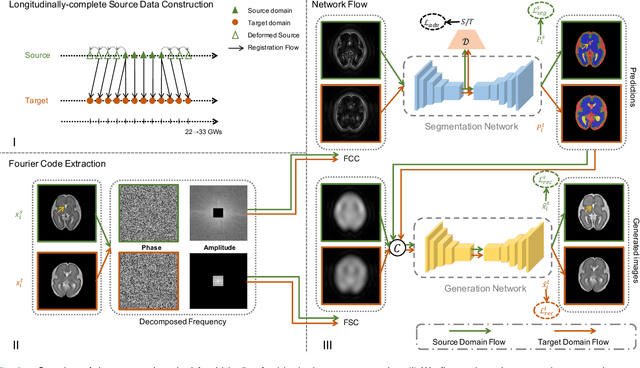
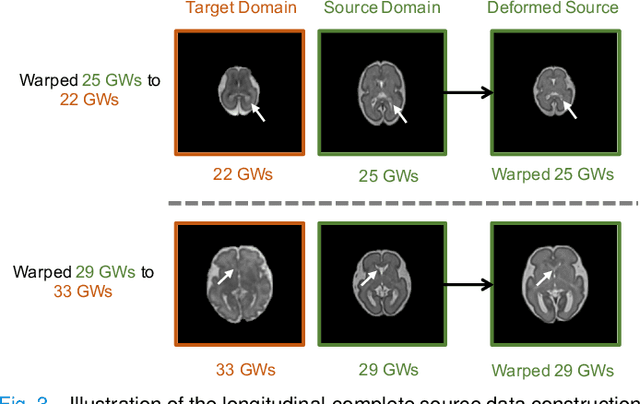
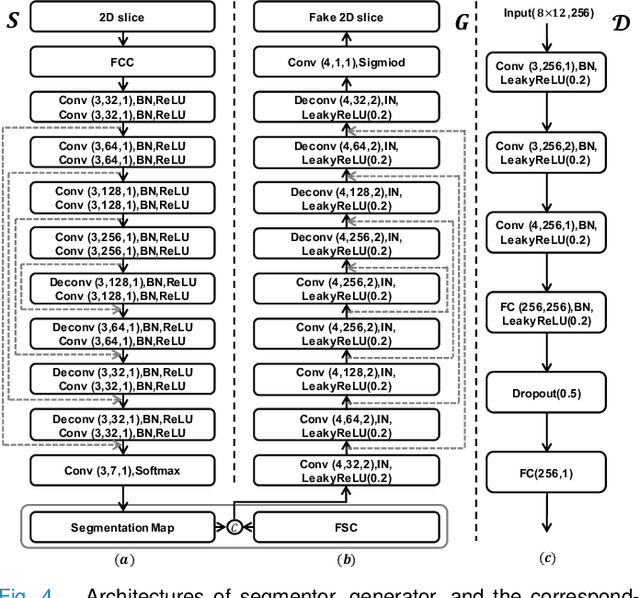
Abstract:Accurate tissue segmentation of thick-slice fetal brain magnetic resonance (MR) scans is crucial for both reconstruction of isotropic brain MR volumes and the quantification of fetal brain development. However, this task is challenging due to the use of thick-slice scans in clinically-acquired fetal brain data. To address this issue, we propose to leverage high-quality isotropic fetal brain MR volumes (and also their corresponding annotations) as guidance for segmentation of thick-slice scans. Due to existence of significant domain gap between high-quality isotropic volume (i.e., source data) and thick-slice scans (i.e., target data), we employ a domain adaptation technique to achieve the associated knowledge transfer (from high-quality <source> volumes to thick-slice <target> scans). Specifically, we first register the available high-quality isotropic fetal brain MR volumes across different gestational weeks to construct longitudinally-complete source data. To capture domain-invariant information, we then perform Fourier decomposition to extract image content and style codes. Finally, we propose a novel Cycle-Consistent Domain Adaptation Network (C2DA-Net) to efficiently transfer the knowledge learned from high-quality isotropic volumes for accurate tissue segmentation of thick-slice scans. Our C2DA-Net can fully utilize a small set of annotated isotropic volumes to guide tissue segmentation on unannotated thick-slice scans. Extensive experiments on a large-scale dataset of 372 clinically acquired thick-slice MR scans demonstrate that our C2DA-Net achieves much better performance than cutting-edge methods quantitatively and qualitatively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge