Coen de Vente
on behalf of the MACUSTAR consortium
Uncertainty-aware retinal layer segmentation in OCT through probabilistic signed distance functions
Dec 06, 2024



Abstract:In this paper, we present a new approach for uncertainty-aware retinal layer segmentation in Optical Coherence Tomography (OCT) scans using probabilistic signed distance functions (SDF). Traditional pixel-wise and regression-based methods primarily encounter difficulties in precise segmentation and lack of geometrical grounding respectively. To address these shortcomings, our methodology refines the segmentation by predicting a signed distance function (SDF) that effectively parameterizes the retinal layer shape via level set. We further enhance the framework by integrating probabilistic modeling, applying Gaussian distributions to encapsulate the uncertainty in the shape parameterization. This ensures a robust representation of the retinal layer morphology even in the presence of ambiguous input, imaging noise, and unreliable segmentations. Both quantitative and qualitative evaluations demonstrate superior performance when compared to other methods. Additionally, we conducted experiments on artificially distorted datasets with various noise types-shadowing, blinking, speckle, and motion-common in OCT scans to showcase the effectiveness of our uncertainty estimation. Our findings demonstrate the possibility to obtain reliable segmentation of retinal layers, as well as an initial step towards the characterization of layer integrity, a key biomarker for disease progression. Our code is available at \url{https://github.com/niazoys/RLS_PSDF}.
Conditioning 3D Diffusion Models with 2D Images: Towards Standardized OCT Volumes through En Face-Informed Super-Resolution
Oct 13, 2024



Abstract:High anisotropy in volumetric medical images can lead to the inconsistent quantification of anatomical and pathological structures. Particularly in optical coherence tomography (OCT), slice spacing can substantially vary across and within datasets, studies, and clinical practices. We propose to standardize OCT volumes to less anisotropic volumes by conditioning 3D diffusion models with en face scanning laser ophthalmoscopy (SLO) imaging data, a 2D modality already commonly available in clinical practice. We trained and evaluated on data from the multicenter and multimodal MACUSTAR study. While upsampling the number of slices by a factor of 8, our method outperforms tricubic interpolation and diffusion models without en face conditioning in terms of perceptual similarity metrics. Qualitative results demonstrate improved coherence and structural similarity. Our approach allows for better informed generative decisions, potentially reducing hallucinations. We hope this work will provide the next step towards standardized high-quality volumetric imaging, enabling more consistent quantifications.
AIROGS: Artificial Intelligence for RObust Glaucoma Screening Challenge
Feb 10, 2023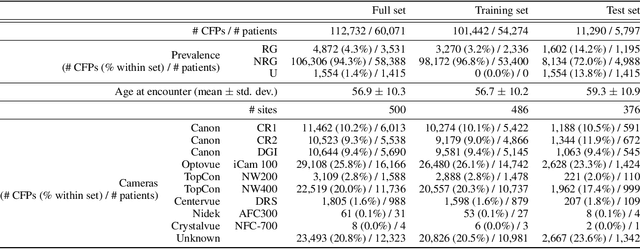
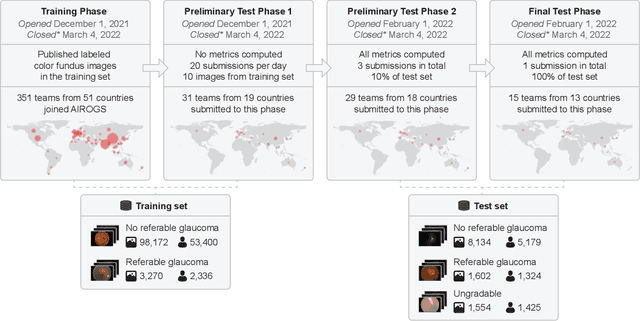
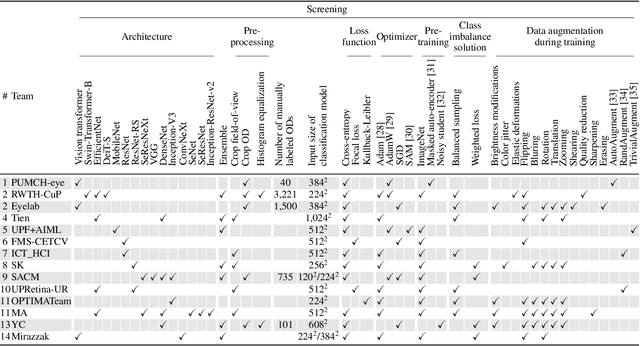
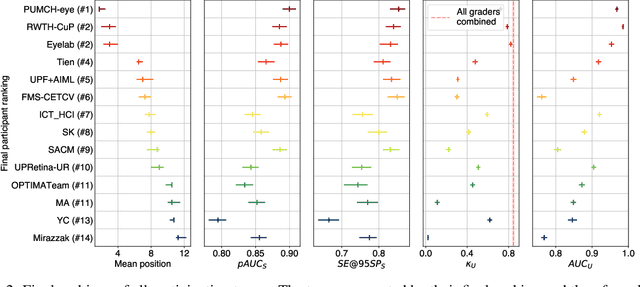
Abstract:The early detection of glaucoma is essential in preventing visual impairment. Artificial intelligence (AI) can be used to analyze color fundus photographs (CFPs) in a cost-effective manner, making glaucoma screening more accessible. While AI models for glaucoma screening from CFPs have shown promising results in laboratory settings, their performance decreases significantly in real-world scenarios due to the presence of out-of-distribution and low-quality images. To address this issue, we propose the Artificial Intelligence for Robust Glaucoma Screening (AIROGS) challenge. This challenge includes a large dataset of around 113,000 images from about 60,000 patients and 500 different screening centers, and encourages the development of algorithms that are robust to ungradable and unexpected input data. We evaluated solutions from 14 teams in this paper, and found that the best teams performed similarly to a set of 20 expert ophthalmologists and optometrists. The highest-scoring team achieved an area under the receiver operating characteristic curve of 0.99 (95% CI: 0.98-0.99) for detecting ungradable images on-the-fly. Additionally, many of the algorithms showed robust performance when tested on three other publicly available datasets. These results demonstrate the feasibility of robust AI-enabled glaucoma screening.
Uncertainty-Aware Multiple-Instance Learning for Reliable Classification: Application to Optical Coherence Tomography
Feb 06, 2023



Abstract:Deep learning classification models for medical image analysis often perform well on data from scanners that were used during training. However, when these models are applied to data from different vendors, their performance tends to drop substantially. Artifacts that only occur within scans from specific scanners are major causes of this poor generalizability. We aimed to improve the reliability of deep learning classification models by proposing Uncertainty-Based Instance eXclusion (UBIX). This technique, based on multiple-instance learning, reduces the effect of corrupted instances on the bag-classification by seamlessly integrating out-of-distribution (OOD) instance detection during inference. Although UBIX is generally applicable to different medical images and diverse classification tasks, we focused on staging of age-related macular degeneration in optical coherence tomography. After being trained using images from one vendor, UBIX showed a reliable behavior, with a slight decrease in performance (a decrease of the quadratic weighted kappa ($\kappa_w$) from 0.861 to 0.708), when applied to images from different vendors containing artifacts; while a state-of-the-art 3D neural network suffered from a significant detriment of performance ($\kappa_w$ from 0.852 to 0.084) on the same test set. We showed that instances with unseen artifacts can be identified with OOD detection and their contribution to the bag-level predictions can be reduced, improving reliability without the need for retraining on new data. This potentially increases the applicability of artificial intelligence models to data from other scanners than the ones for which they were developed.
A deep learning framework for the detection and quantification of drusen and reticular pseudodrusen on optical coherence tomography
Apr 05, 2022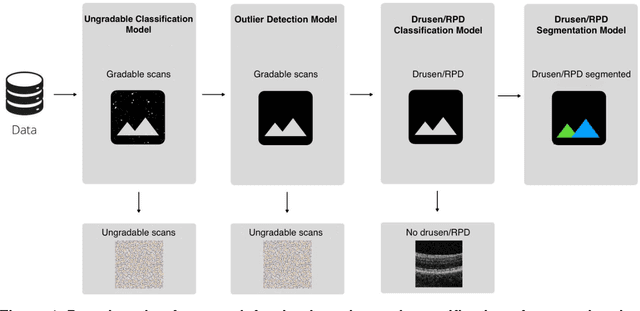
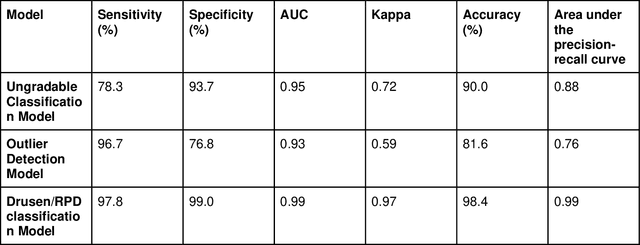
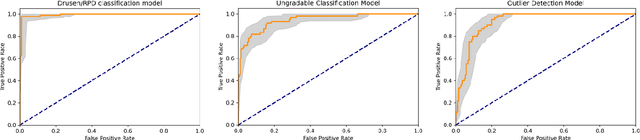
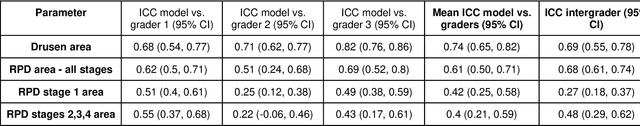
Abstract:Purpose - To develop and validate a deep learning (DL) framework for the detection and quantification of drusen and reticular pseudodrusen (RPD) on optical coherence tomography scans. Design - Development and validation of deep learning models for classification and feature segmentation. Methods - A DL framework was developed consisting of a classification model and an out-of-distribution (OOD) detection model for the identification of ungradable scans; a classification model to identify scans with drusen or RPD; and an image segmentation model to independently segment lesions as RPD or drusen. Data were obtained from 1284 participants in the UK Biobank (UKBB) with a self-reported diagnosis of age-related macular degeneration (AMD) and 250 UKBB controls. Drusen and RPD were manually delineated by five retina specialists. The main outcome measures were sensitivity, specificity, area under the ROC curve (AUC), kappa, accuracy and intraclass correlation coefficient (ICC). Results - The classification models performed strongly at their respective tasks (0.95, 0.93, and 0.99 AUC, respectively, for the ungradable scans classifier, the OOD model, and the drusen and RPD classification model). The mean ICC for drusen and RPD area vs. graders was 0.74 and 0.61, respectively, compared with 0.69 and 0.68 for intergrader agreement. FROC curves showed that the model's sensitivity was close to human performance. Conclusions - The models achieved high classification and segmentation performance, similar to human performance. Application of this robust framework will further our understanding of RPD as a separate entity from drusen in both research and clinical settings.
Improving Automated COVID-19 Grading with Convolutional Neural Networks in Computed Tomography Scans: An Ablation Study
Sep 21, 2020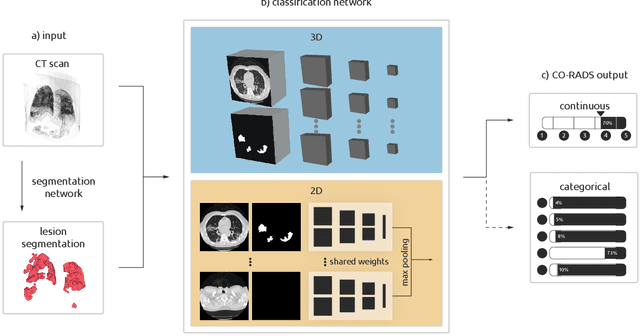
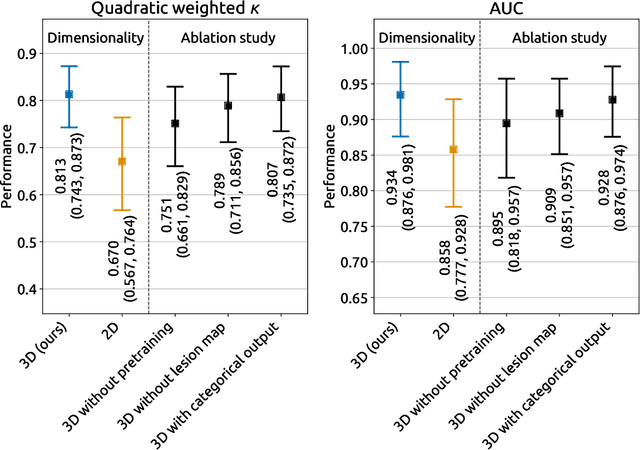
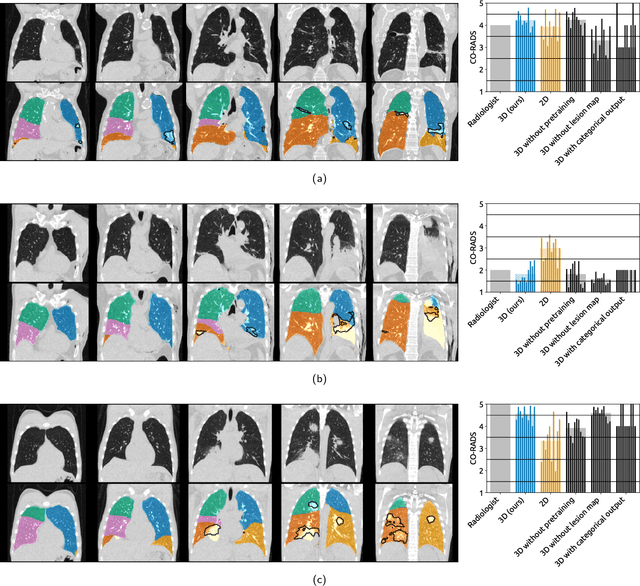
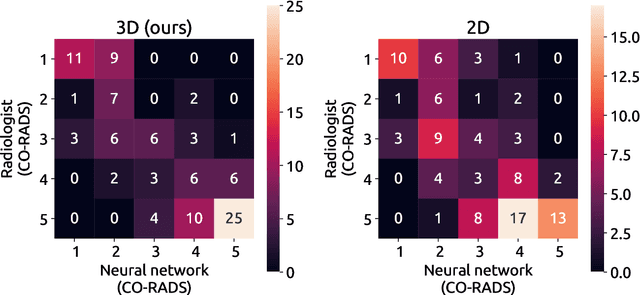
Abstract:Amidst the ongoing pandemic, several studies have shown that COVID-19 classification and grading using computed tomography (CT) images can be automated with convolutional neural networks (CNNs). Many of these studies focused on reporting initial results of algorithms that were assembled from commonly used components. The choice of these components was often pragmatic rather than systematic. For instance, several studies used 2D CNNs even though these might not be optimal for handling 3D CT volumes. This paper identifies a variety of components that increase the performance of CNN-based algorithms for COVID-19 grading from CT images. We investigated the effectiveness of using a 3D CNN instead of a 2D CNN, of using transfer learning to initialize the network, of providing automatically computed lesion maps as additional network input, and of predicting a continuous instead of a categorical output. A 3D CNN with these components achieved an area under the ROC curve (AUC) of 0.934 on our test set of 105 CT scans and an AUC of 0.923 on a publicly available set of 742 CT scans, a substantial improvement in comparison with a previously published 2D CNN. An ablation study demonstrated that in addition to using a 3D CNN instead of a 2D CNN transfer learning contributed the most and continuous output contributed the least to improving the model performance.
A Global Benchmark of Algorithms for Segmenting Late Gadolinium-Enhanced Cardiac Magnetic Resonance Imaging
May 07, 2020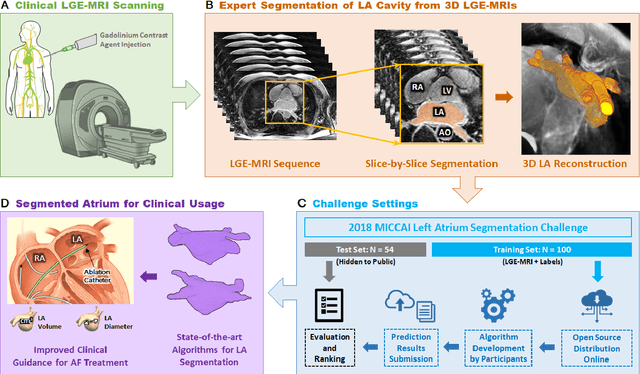
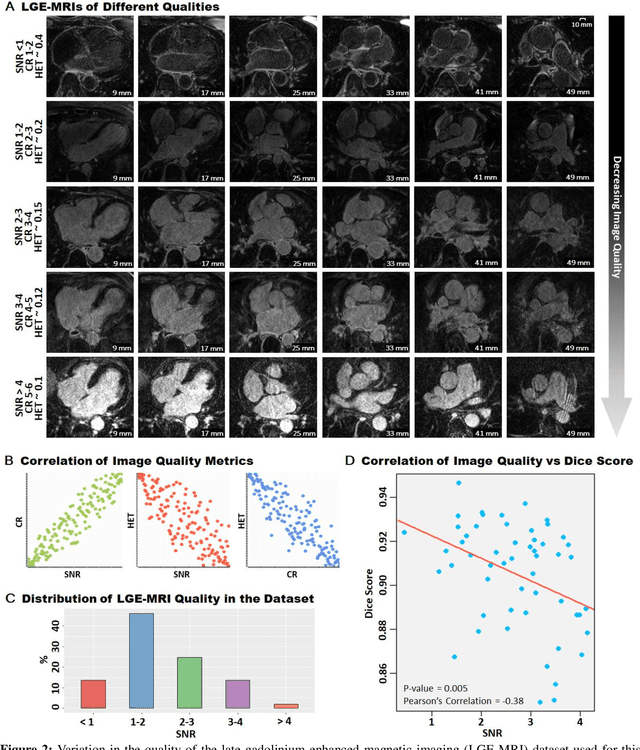
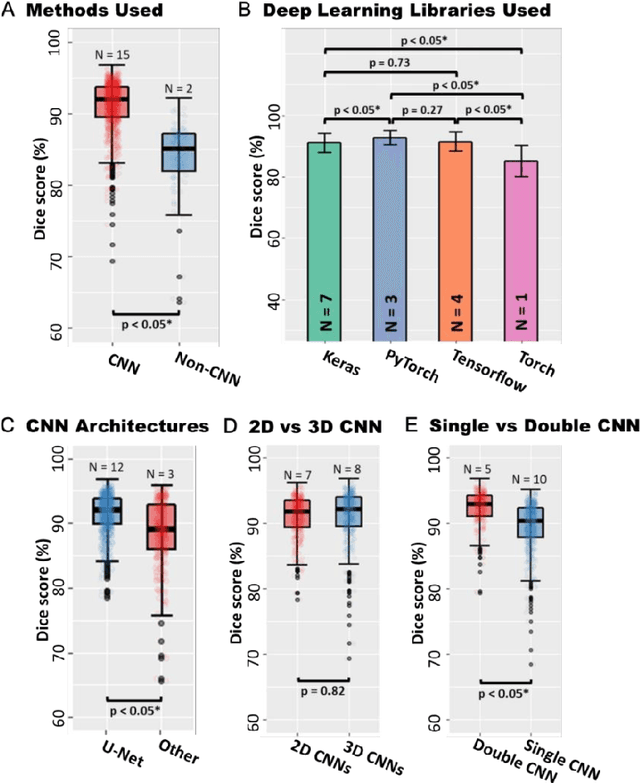
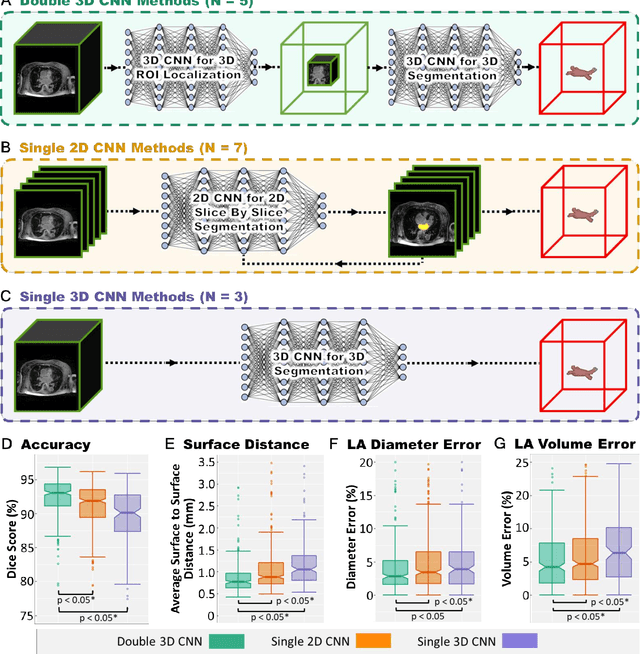
Abstract:Segmentation of cardiac images, particularly late gadolinium-enhanced magnetic resonance imaging (LGE-MRI) widely used for visualizing diseased cardiac structures, is a crucial first step for clinical diagnosis and treatment. However, direct segmentation of LGE-MRIs is challenging due to its attenuated contrast. Since most clinical studies have relied on manual and labor-intensive approaches, automatic methods are of high interest, particularly optimized machine learning approaches. To address this, we organized the "2018 Left Atrium Segmentation Challenge" using 154 3D LGE-MRIs, currently the world's largest cardiac LGE-MRI dataset, and associated labels of the left atrium segmented by three medical experts, ultimately attracting the participation of 27 international teams. In this paper, extensive analysis of the submitted algorithms using technical and biological metrics was performed by undergoing subgroup analysis and conducting hyper-parameter analysis, offering an overall picture of the major design choices of convolutional neural networks (CNNs) and practical considerations for achieving state-of-the-art left atrium segmentation. Results show the top method achieved a dice score of 93.2% and a mean surface to a surface distance of 0.7 mm, significantly outperforming prior state-of-the-art. Particularly, our analysis demonstrated that double, sequentially used CNNs, in which a first CNN is used for automatic region-of-interest localization and a subsequent CNN is used for refined regional segmentation, achieved far superior results than traditional methods and pipelines containing single CNNs. This large-scale benchmarking study makes a significant step towards much-improved segmentation methods for cardiac LGE-MRIs, and will serve as an important benchmark for evaluating and comparing the future works in the field.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge