Firas Khader
Medical Slice Transformer: Improved Diagnosis and Explainability on 3D Medical Images with DINOv2
Nov 24, 2024



Abstract:MRI and CT are essential clinical cross-sectional imaging techniques for diagnosing complex conditions. However, large 3D datasets with annotations for deep learning are scarce. While methods like DINOv2 are encouraging for 2D image analysis, these methods have not been applied to 3D medical images. Furthermore, deep learning models often lack explainability due to their "black-box" nature. This study aims to extend 2D self-supervised models, specifically DINOv2, to 3D medical imaging while evaluating their potential for explainable outcomes. We introduce the Medical Slice Transformer (MST) framework to adapt 2D self-supervised models for 3D medical image analysis. MST combines a Transformer architecture with a 2D feature extractor, i.e., DINOv2. We evaluate its diagnostic performance against a 3D convolutional neural network (3D ResNet) across three clinical datasets: breast MRI (651 patients), chest CT (722 patients), and knee MRI (1199 patients). Both methods were tested for diagnosing breast cancer, predicting lung nodule dignity, and detecting meniscus tears. Diagnostic performance was assessed by calculating the Area Under the Receiver Operating Characteristic Curve (AUC). Explainability was evaluated through a radiologist's qualitative comparison of saliency maps based on slice and lesion correctness. P-values were calculated using Delong's test. MST achieved higher AUC values compared to ResNet across all three datasets: breast (0.94$\pm$0.01 vs. 0.91$\pm$0.02, P=0.02), chest (0.95$\pm$0.01 vs. 0.92$\pm$0.02, P=0.13), and knee (0.85$\pm$0.04 vs. 0.69$\pm$0.05, P=0.001). Saliency maps were consistently more precise and anatomically correct for MST than for ResNet. Self-supervised 2D models like DINOv2 can be effectively adapted for 3D medical imaging using MST, offering enhanced diagnostic accuracy and explainability compared to convolutional neural networks.
On Instabilities of Unsupervised Denoising Diffusion Models in Magnetic Resonance Imaging Reconstruction
Jun 23, 2024Abstract:Denoising diffusion models offer a promising approach to accelerating magnetic resonance imaging (MRI) and producing diagnostic-level images in an unsupervised manner. However, our study demonstrates that even tiny worst-case potential perturbations transferred from a surrogate model can cause these models to generate fake tissue structures that may mislead clinicians. The transferability of such worst-case perturbations indicates that the robustness of image reconstruction may be compromised due to MR system imperfections or other sources of noise. Moreover, at larger perturbation strengths, diffusion models exhibit Gaussian noise-like artifacts that are distinct from those observed in supervised models and are more challenging to detect. Our results highlight the vulnerability of current state-of-the-art diffusion-based reconstruction models to possible worst-case perturbations and underscore the need for further research to improve their robustness and reliability in clinical settings.
Compute-Efficient Medical Image Classification with Softmax-Free Transformers and Sequence Normalization
Jun 03, 2024Abstract:The Transformer model has been pivotal in advancing fields such as natural language processing, speech recognition, and computer vision. However, a critical limitation of this model is its quadratic computational and memory complexity relative to the sequence length, which constrains its application to longer sequences. This is especially crucial in medical imaging where high-resolution images can reach gigapixel scale. Efforts to address this issue have predominantely focused on complex techniques, such as decomposing the softmax operation integral to the Transformer's architecture. This paper addresses this quadratic computational complexity of Transformer models and introduces a remarkably simple and effective method that circumvents this issue by eliminating the softmax function from the attention mechanism and adopting a sequence normalization technique for the key, query, and value tokens. Coupled with a reordering of matrix multiplications this approach reduces the memory- and compute complexity to a linear scale. We evaluate this approach across various medical imaging datasets comprising fundoscopic, dermascopic, radiologic and histologic imaging data. Our findings highlight that these models exhibit a comparable performance to traditional transformer models, while efficiently handling longer sequences.
Joint multi-task learning improves weakly-supervised biomarker prediction in computational pathology
Mar 06, 2024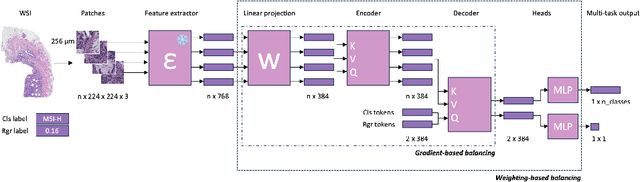


Abstract:Deep Learning (DL) can predict biomarkers directly from digitized cancer histology in a weakly-supervised setting. Recently, the prediction of continuous biomarkers through regression-based DL has seen an increasing interest. Nonetheless, clinical decision making often requires a categorical outcome. Consequently, we developed a weakly-supervised joint multi-task Transformer architecture which has been trained and evaluated on four public patient cohorts for the prediction of two key predictive biomarkers, microsatellite instability (MSI) and homologous recombination deficiency (HRD), trained with auxiliary regression tasks related to the tumor microenvironment. Moreover, we perform a comprehensive benchmark of 16 approaches of task balancing for weakly-supervised joint multi-task learning in computational pathology. Using our novel approach, we improve over the state-of-the-art area under the receiver operating characteristic by +7.7% and +4.1%, as well as yielding better clustering of latent embeddings by +8% and +5% for the prediction of MSI and HRD in external cohorts, respectively.
An Ordinal Regression Framework for a Deep Learning Based Severity Assessment for Chest Radiographs
Feb 08, 2024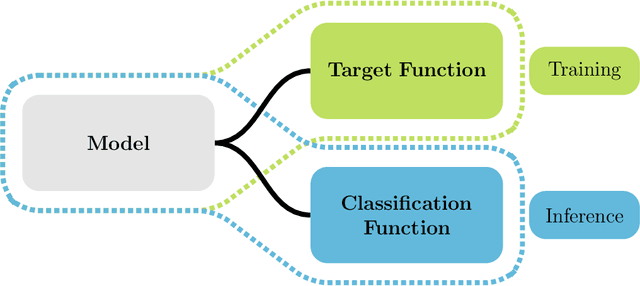
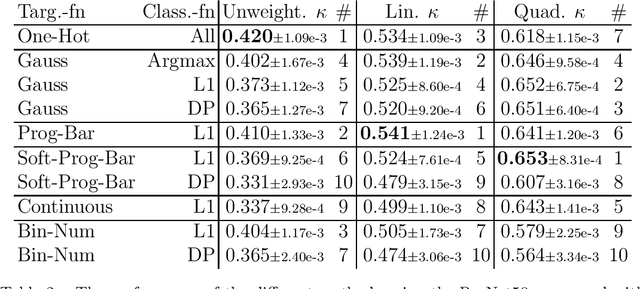
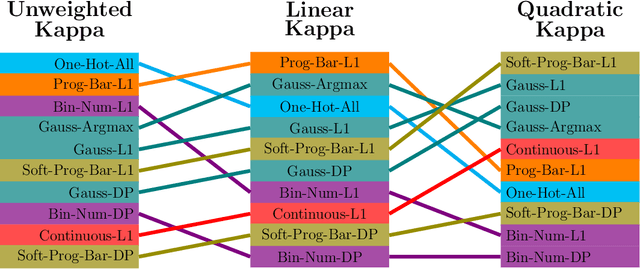
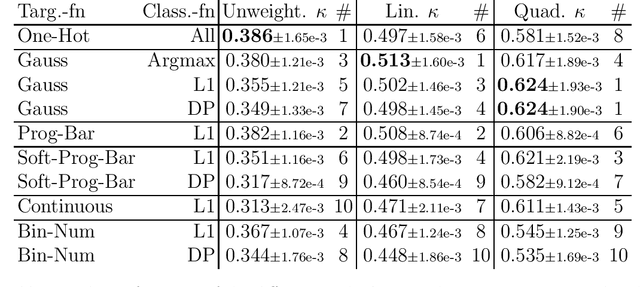
Abstract:This study investigates the application of ordinal regression methods for categorizing disease severity in chest radiographs. We propose a framework that divides the ordinal regression problem into three parts: a model, a target function, and a classification function. Different encoding methods, including one-hot, Gaussian, progress-bar, and our soft-progress-bar, are applied using ResNet50 and ViT-B-16 deep learning models. We show that the choice of encoding has a strong impact on performance and that the best encoding depends on the chosen weighting of Cohen's kappa and also on the model architecture used. We make our code publicly available on GitHub.
From Whole-slide Image to Biomarker Prediction: A Protocol for End-to-End Deep Learning in Computational Pathology
Dec 18, 2023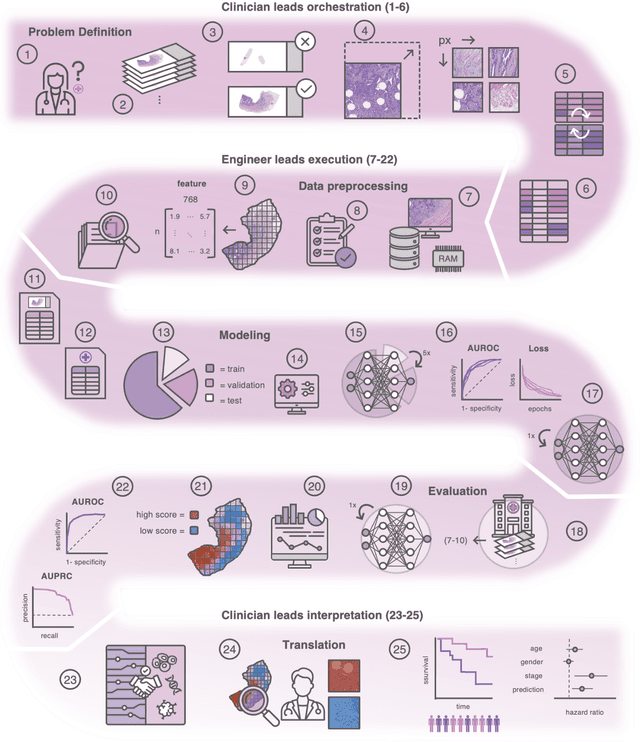
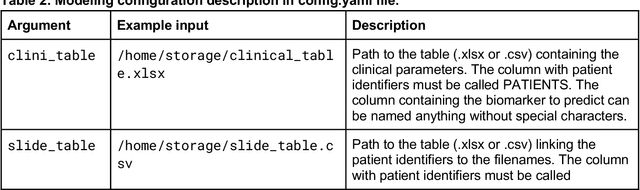
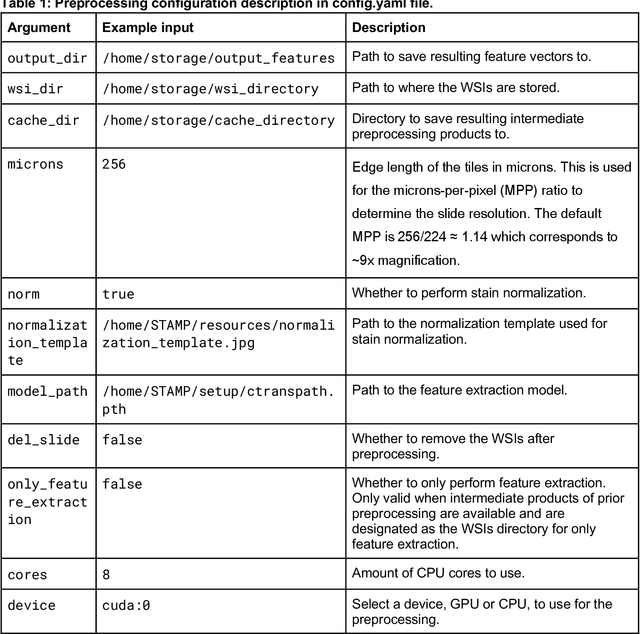
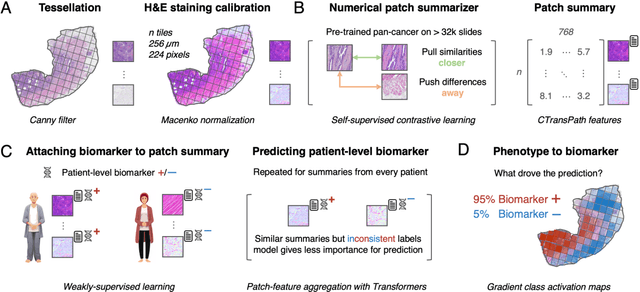

Abstract:Hematoxylin- and eosin (H&E) stained whole-slide images (WSIs) are the foundation of diagnosis of cancer. In recent years, development of deep learning-based methods in computational pathology enabled the prediction of biomarkers directly from WSIs. However, accurately linking tissue phenotype to biomarkers at scale remains a crucial challenge for democratizing complex biomarkers in precision oncology. This protocol describes a practical workflow for solid tumor associative modeling in pathology (STAMP), enabling prediction of biomarkers directly from WSIs using deep learning. The STAMP workflow is biomarker agnostic and allows for genetic- and clinicopathologic tabular data to be included as an additional input, together with histopathology images. The protocol consists of five main stages which have been successfully applied to various research problems: formal problem definition, data preprocessing, modeling, evaluation and clinical translation. The STAMP workflow differentiates itself through its focus on serving as a collaborative framework that can be used by clinicians and engineers alike for setting up research projects in the field of computational pathology. As an example task, we applied STAMP to the prediction of microsatellite instability (MSI) status in colorectal cancer, showing accurate performance for the identification of MSI-high tumors. Moreover, we provide an open-source codebase which has been deployed at several hospitals across the globe to set up computational pathology workflows. The STAMP workflow requires one workday of hands-on computational execution and basic command line knowledge.
Medical Foundation Models are Susceptible to Targeted Misinformation Attacks
Sep 29, 2023



Abstract:Large language models (LLMs) have broad medical knowledge and can reason about medical information across many domains, holding promising potential for diverse medical applications in the near future. In this study, we demonstrate a concerning vulnerability of LLMs in medicine. Through targeted manipulation of just 1.1% of the model's weights, we can deliberately inject an incorrect biomedical fact. The erroneous information is then propagated in the model's output, whilst its performance on other biomedical tasks remains intact. We validate our findings in a set of 1,038 incorrect biomedical facts. This peculiar susceptibility raises serious security and trustworthiness concerns for the application of LLMs in healthcare settings. It accentuates the need for robust protective measures, thorough verification mechanisms, and stringent management of access to these models, ensuring their reliable and safe use in medical practice.
Reconstruction of Patient-Specific Confounders in AI-based Radiologic Image Interpretation using Generative Pretraining
Sep 29, 2023
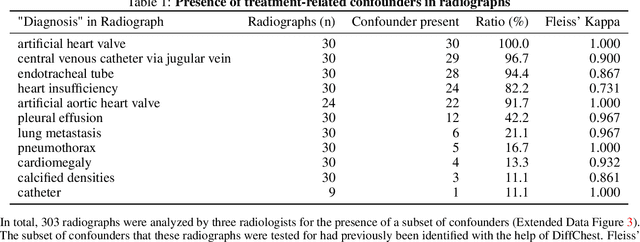
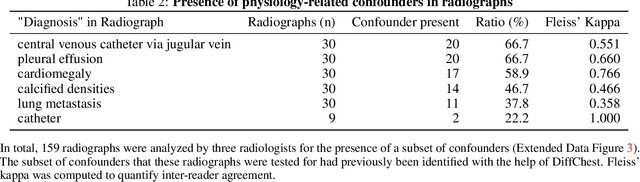

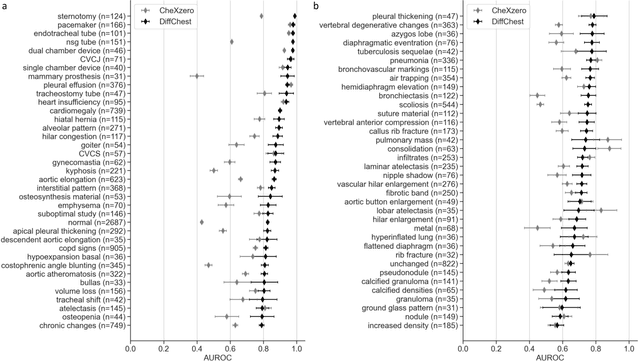

Abstract:Detecting misleading patterns in automated diagnostic assistance systems, such as those powered by Artificial Intelligence, is critical to ensuring their reliability, particularly in healthcare. Current techniques for evaluating deep learning models cannot visualize confounding factors at a diagnostic level. Here, we propose a self-conditioned diffusion model termed DiffChest and train it on a dataset of 515,704 chest radiographs from 194,956 patients from multiple healthcare centers in the United States and Europe. DiffChest explains classifications on a patient-specific level and visualizes the confounding factors that may mislead the model. We found high inter-reader agreement when evaluating DiffChest's capability to identify treatment-related confounders, with Fleiss' Kappa values of 0.8 or higher across most imaging findings. Confounders were accurately captured with 11.1% to 100% prevalence rates. Furthermore, our pretraining process optimized the model to capture the most relevant information from the input radiographs. DiffChest achieved excellent diagnostic accuracy when diagnosing 11 chest conditions, such as pleural effusion and cardiac insufficiency, and at least sufficient diagnostic accuracy for the remaining conditions. Our findings highlight the potential of pretraining based on diffusion models in medical image classification, specifically in providing insights into confounding factors and model robustness.
Cascaded Cross-Attention Networks for Data-Efficient Whole-Slide Image Classification Using Transformers
May 11, 2023Abstract:Whole-Slide Imaging allows for the capturing and digitization of high-resolution images of histological specimen. An automated analysis of such images using deep learning models is therefore of high demand. The transformer architecture has been proposed as a possible candidate for effectively leveraging the high-resolution information. Here, the whole-slide image is partitioned into smaller image patches and feature tokens are extracted from these image patches. However, while the conventional transformer allows for a simultaneous processing of a large set of input tokens, the computational demand scales quadratically with the number of input tokens and thus quadratically with the number of image patches. To address this problem we propose a novel cascaded cross-attention network (CCAN) based on the cross-attention mechanism that scales linearly with the number of extracted patches. Our experiments demonstrate that this architecture is at least on-par with and even outperforms other attention-based state-of-the-art methods on two public datasets: On the use-case of lung cancer (TCGA NSCLC) our model reaches a mean area under the receiver operating characteristic (AUC) of 0.970 $\pm$ 0.008 and on renal cancer (TCGA RCC) reaches a mean AUC of 0.985 $\pm$ 0.004. Furthermore, we show that our proposed model is efficient in low-data regimes, making it a promising approach for analyzing whole-slide images in resource-limited settings. To foster research in this direction, we make our code publicly available on GitHub: XXX.
Transformers for CT Reconstruction From Monoplanar and Biplanar Radiographs
May 11, 2023



Abstract:Computed Tomography (CT) scans provide detailed and accurate information of internal structures in the body. They are constructed by sending x-rays through the body from different directions and combining this information into a three-dimensional volume. Such volumes can then be used to diagnose a wide range of conditions and allow for volumetric measurements of organs. In this work, we tackle the problem of reconstructing CT images from biplanar x-rays only. X-rays are widely available and even if the CT reconstructed from these radiographs is not a replacement of a complete CT in the diagnostic setting, it might serve to spare the patients from radiation where a CT is only acquired for rough measurements such as determining organ size. We propose a novel method based on the transformer architecture, by framing the underlying task as a language translation problem. Radiographs and CT images are first embedded into latent quantized codebook vectors using two different autoencoder networks. We then train a GPT model, to reconstruct the codebook vectors of the CT image, conditioned on the codebook vectors of the x-rays and show that this approach leads to realistic looking images. To encourage further research in this direction, we make our code publicly available on GitHub: XXX.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge