Jakob Nikolas Kather
Agentic Systems in Radiology: Design, Applications, Evaluation, and Challenges
Oct 10, 2025Abstract:Building agents, systems that perceive and act upon their environment with a degree of autonomy, has long been a focus of AI research. This pursuit has recently become vastly more practical with the emergence of large language models (LLMs) capable of using natural language to integrate information, follow instructions, and perform forms of "reasoning" and planning across a wide range of tasks. With its multimodal data streams and orchestrated workflows spanning multiple systems, radiology is uniquely suited to benefit from agents that can adapt to context and automate repetitive yet complex tasks. In radiology, LLMs and their multimodal variants have already demonstrated promising performance for individual tasks such as information extraction and report summarization. However, using LLMs in isolation underutilizes their potential to support complex, multi-step workflows where decisions depend on evolving context from multiple information sources. Equipping LLMs with external tools and feedback mechanisms enables them to drive systems that exhibit a spectrum of autonomy, ranging from semi-automated workflows to more adaptive agents capable of managing complex processes. This review examines the design of such LLM-driven agentic systems, highlights key applications, discusses evaluation methods for planning and tool use, and outlines challenges such as error cascades, tool-use efficiency, and health IT integration.
Resolution scaling governs DINOv3 transfer performance in chest radiograph classification
Oct 08, 2025Abstract:Self-supervised learning (SSL) has advanced visual representation learning, but its value in chest radiography, a high-volume imaging modality with fine-grained findings, remains unclear. Meta's DINOv3 extends earlier SSL models through Gram-anchored self-distillation. Whether these design choices improve transfer learning for chest radiography has not been systematically tested. We benchmarked DINOv3 against DINOv2 and ImageNet initialization across seven datasets (n>814,000). Two representative backbones were evaluated: ViT-B/16 and ConvNeXt-B. Images were analyzed at 224x224, 512x512, and 1024x1024 pixels. We additionally assessed frozen features from a 7B model. The primary outcome was mean AUROC across labels. At 224x224, DINOv3 and DINOv2 achieved comparable performance on adult datasets. Increasing resolution to 512x512 yielded consistent improvements for DINOv3 over both DINOv2 and ImageNet. In contrast, results in pediatric cohort showed no differences across initializations. Across all settings, ConvNeXt-B outperformed ViT-B/16. Models using frozen DINOv3-7B features underperformed relative to fully finetuned 86-89M-parameter backbones, highlighting the importance of domain adaptation. Scaling to 1024x1024 did not further improve accuracy. Resolution-related gains were most evident for boundary-dependent and small focal abnormalities. In chest radiography, higher input resolution is critical for leveraging the benefits of modern self-supervised models. 512x512 pixels represent a practical upper limit where DINOv3-initialized ConvNeXt-B networks provide the strongest performance, while larger inputs offer minimal return on cost. Clinically, these findings support use of finetuned, mid-sized backbones at 512x512 for chest radiograph interpretation, with the greatest gains expected in detecting subtle or boundary-centered lesions relevant to emergency and critical care settings.
MedicalPatchNet: A Patch-Based Self-Explainable AI Architecture for Chest X-ray Classification
Sep 09, 2025Abstract:Deep neural networks excel in radiological image classification but frequently suffer from poor interpretability, limiting clinical acceptance. We present MedicalPatchNet, an inherently self-explainable architecture for chest X-ray classification that transparently attributes decisions to distinct image regions. MedicalPatchNet splits images into non-overlapping patches, independently classifies each patch, and aggregates predictions, enabling intuitive visualization of each patch's diagnostic contribution without post-hoc techniques. Trained on the CheXpert dataset (223,414 images), MedicalPatchNet matches the classification performance (AUROC 0.907 vs. 0.908) of EfficientNet-B0, while substantially improving interpretability: MedicalPatchNet demonstrates substantially improved interpretability with higher pathology localization accuracy (mean hit-rate 0.485 vs. 0.376 with Grad-CAM) on the CheXlocalize dataset. By providing explicit, reliable explanations accessible even to non-AI experts, MedicalPatchNet mitigates risks associated with shortcut learning, thus improving clinical trust. Our model is publicly available with reproducible training and inference scripts and contributes to safer, explainable AI-assisted diagnostics across medical imaging domains. We make the code publicly available: https://github.com/TruhnLab/MedicalPatchNet
A deep learning framework for efficient pathology image analysis
Feb 18, 2025



Abstract:Artificial intelligence (AI) has transformed digital pathology by enabling biomarker prediction from high-resolution whole slide images (WSIs). However, current methods are computationally inefficient, processing thousands of redundant tiles per WSI and requiring complex aggregator models. We introduce EAGLE (Efficient Approach for Guided Local Examination), a deep learning framework that emulates pathologists by selectively analyzing informative regions. EAGLE incorporates two foundation models: CHIEF for efficient tile selection and Virchow2 for extracting high-quality features. Benchmarking was conducted against leading slide- and tile-level foundation models across 31 tasks from four cancer types, spanning morphology, biomarker prediction and prognosis. EAGLE outperformed state-of-the-art foundation models by up to 23% and achieved the highest AUROC overall. It processed a slide in 2.27 seconds, reducing computational time by more than 99% compared to existing models. This efficiency enables real-time workflows, allows pathologists to validate all tiles which are used by the model during analysis, and eliminates dependence on high-performance computing, making AI-powered pathology more accessible. By reliably identifying meaningful regions and minimizing artifacts, EAGLE provides robust and interpretable outputs, supporting rapid slide searches, integration into multi-omics pipelines and emerging clinical foundation models.
LLM Agents Making Agent Tools
Feb 17, 2025
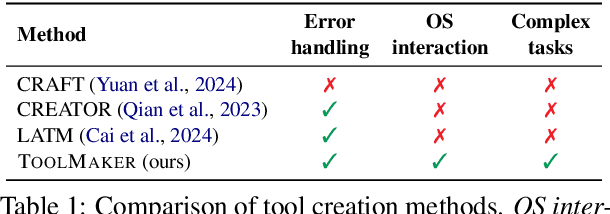
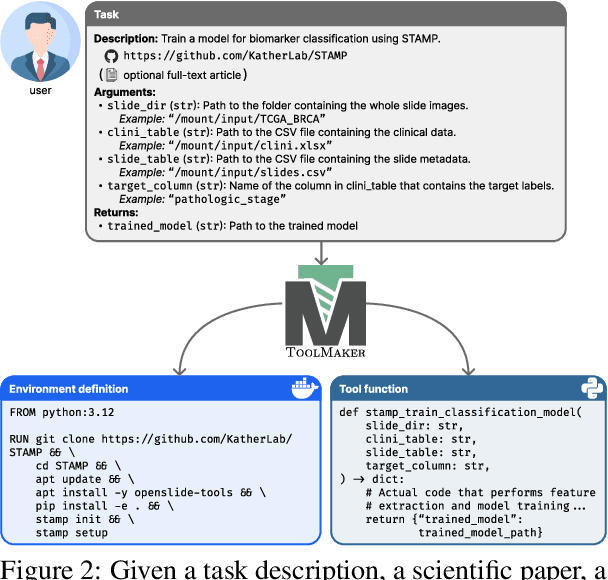


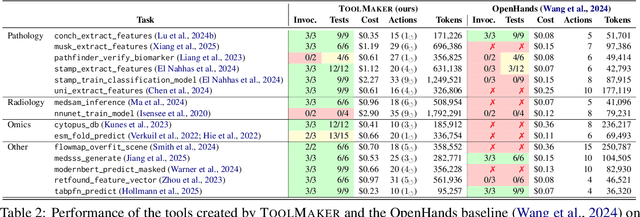
Abstract:Tool use has turned large language models (LLMs) into powerful agents that can perform complex multi-step tasks by dynamically utilising external software components. However, these tools must be implemented in advance by human developers, hindering the applicability of LLM agents in domains which demand large numbers of highly specialised tools, like in life sciences and medicine. Motivated by the growing trend of scientific studies accompanied by public code repositories, we propose ToolMaker, a novel agentic framework that autonomously transforms papers with code into LLM-compatible tools. Given a short task description and a repository URL, ToolMaker autonomously installs required dependencies and generates code to perform the task, using a closed-loop self-correction mechanism to iteratively diagnose and rectify errors. To evaluate our approach, we introduce a benchmark comprising 15 diverse and complex computational tasks spanning both medical and non-medical domains with over 100 unit tests to objectively assess tool correctness and robustness. ToolMaker correctly implements 80% of the tasks, substantially outperforming current state-of-the-art software engineering agents. ToolMaker therefore is a step towards fully autonomous agent-based scientific workflows.
Abnormality-Driven Representation Learning for Radiology Imaging
Nov 25, 2024



Abstract:To date, the most common approach for radiology deep learning pipelines is the use of end-to-end 3D networks based on models pre-trained on other tasks, followed by fine-tuning on the task at hand. In contrast, adjacent medical fields such as pathology, which focus on 2D images, have effectively adopted task-agnostic foundational models based on self-supervised learning (SSL), combined with weakly-supervised deep learning (DL). However, the field of radiology still lacks task-agnostic representation models due to the computational and data demands of 3D imaging and the anatomical complexity inherent to radiology scans. To address this gap, we propose CLEAR, a framework for radiology images that uses extracted embeddings from 2D slices along with attention-based aggregation for efficiently predicting clinical endpoints. As part of this framework, we introduce lesion-enhanced contrastive learning (LeCL), a novel approach to obtain visual representations driven by abnormalities in 2D axial slices across different locations of the CT scans. Specifically, we trained single-domain contrastive learning approaches using three different architectures: Vision Transformers, Vision State Space Models and Gated Convolutional Neural Networks. We evaluate our approach across three clinical tasks: tumor lesion location, lung disease detection, and patient staging, benchmarking against four state-of-the-art foundation models, including BiomedCLIP. Our findings demonstrate that CLEAR using representations learned through LeCL, outperforms existing foundation models, while being substantially more compute- and data-efficient.
Medical Slice Transformer: Improved Diagnosis and Explainability on 3D Medical Images with DINOv2
Nov 24, 2024



Abstract:MRI and CT are essential clinical cross-sectional imaging techniques for diagnosing complex conditions. However, large 3D datasets with annotations for deep learning are scarce. While methods like DINOv2 are encouraging for 2D image analysis, these methods have not been applied to 3D medical images. Furthermore, deep learning models often lack explainability due to their "black-box" nature. This study aims to extend 2D self-supervised models, specifically DINOv2, to 3D medical imaging while evaluating their potential for explainable outcomes. We introduce the Medical Slice Transformer (MST) framework to adapt 2D self-supervised models for 3D medical image analysis. MST combines a Transformer architecture with a 2D feature extractor, i.e., DINOv2. We evaluate its diagnostic performance against a 3D convolutional neural network (3D ResNet) across three clinical datasets: breast MRI (651 patients), chest CT (722 patients), and knee MRI (1199 patients). Both methods were tested for diagnosing breast cancer, predicting lung nodule dignity, and detecting meniscus tears. Diagnostic performance was assessed by calculating the Area Under the Receiver Operating Characteristic Curve (AUC). Explainability was evaluated through a radiologist's qualitative comparison of saliency maps based on slice and lesion correctness. P-values were calculated using Delong's test. MST achieved higher AUC values compared to ResNet across all three datasets: breast (0.94$\pm$0.01 vs. 0.91$\pm$0.02, P=0.02), chest (0.95$\pm$0.01 vs. 0.92$\pm$0.02, P=0.13), and knee (0.85$\pm$0.04 vs. 0.69$\pm$0.05, P=0.001). Saliency maps were consistently more precise and anatomically correct for MST than for ResNet. Self-supervised 2D models like DINOv2 can be effectively adapted for 3D medical imaging using MST, offering enhanced diagnostic accuracy and explainability compared to convolutional neural networks.
Unsupervised Foundation Model-Agnostic Slide-Level Representation Learning
Nov 20, 2024



Abstract:Representation learning of pathology whole-slide images (WSIs) has primarily relied on weak supervision with Multiple Instance Learning (MIL). This approach leads to slide representations highly tailored to a specific clinical task. Self-supervised learning (SSL) has been successfully applied to train histopathology foundation models (FMs) for patch embedding generation. However, generating patient or slide level embeddings remains challenging. Existing approaches for slide representation learning extend the principles of SSL from patch level learning to entire slides by aligning different augmentations of the slide or by utilizing multimodal data. By integrating tile embeddings from multiple FMs, we propose a new single modality SSL method in feature space that generates useful slide representations. Our contrastive pretraining strategy, called COBRA, employs multiple FMs and an architecture based on Mamba-2. COBRA exceeds performance of state-of-the-art slide encoders on four different public CPTAC cohorts on average by at least +3.8% AUC, despite only being pretrained on 3048 WSIs from TCGA. Additionally, COBRA is readily compatible at inference time with previously unseen feature extractors.
Pathologist-like explainable AI for interpretable Gleason grading in prostate cancer
Oct 19, 2024



Abstract:The aggressiveness of prostate cancer, the most common cancer in men worldwide, is primarily assessed based on histopathological data using the Gleason scoring system. While artificial intelligence (AI) has shown promise in accurately predicting Gleason scores, these predictions often lack inherent explainability, potentially leading to distrust in human-machine interactions. To address this issue, we introduce a novel dataset of 1,015 tissue microarray core images, annotated by an international group of 54 pathologists. The annotations provide detailed localized pattern descriptions for Gleason grading in line with international guidelines. Utilizing this dataset, we develop an inherently explainable AI system based on a U-Net architecture that provides predictions leveraging pathologists' terminology. This approach circumvents post-hoc explainability methods while maintaining or exceeding the performance of methods trained directly for Gleason pattern segmentation (Dice score: 0.713 $\pm$ 0.003 trained on explanations vs. 0.691 $\pm$ 0.010 trained on Gleason patterns). By employing soft labels during training, we capture the intrinsic uncertainty in the data, yielding strong results in Gleason pattern segmentation even in the context of high interobserver variability. With the release of this dataset, we aim to encourage further research into segmentation in medical tasks with high levels of subjectivity and to advance the understanding of pathologists' reasoning processes.
Large Language Models-Enabled Digital Twins for Precision Medicine in Rare Gynecological Tumors
Aug 31, 2024Abstract:Rare gynecological tumors (RGTs) present major clinical challenges due to their low incidence and heterogeneity. The lack of clear guidelines leads to suboptimal management and poor prognosis. Molecular tumor boards accelerate access to effective therapies by tailoring treatment based on biomarkers, beyond cancer type. Unstructured data that requires manual curation hinders efficient use of biomarker profiling for therapy matching. This study explores the use of large language models (LLMs) to construct digital twins for precision medicine in RGTs. Our proof-of-concept digital twin system integrates clinical and biomarker data from institutional and published cases (n=21) and literature-derived data (n=655 publications with n=404,265 patients) to create tailored treatment plans for metastatic uterine carcinosarcoma, identifying options potentially missed by traditional, single-source analysis. LLM-enabled digital twins efficiently model individual patient trajectories. Shifting to a biology-based rather than organ-based tumor definition enables personalized care that could advance RGT management and thus enhance patient outcomes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge