Dyke Ferber
LLM Agents Making Agent Tools
Feb 17, 2025
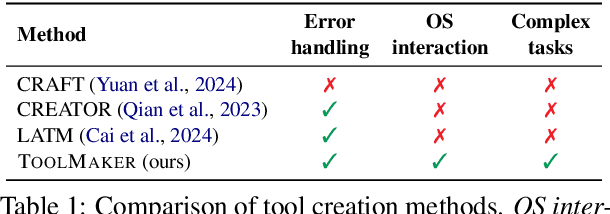
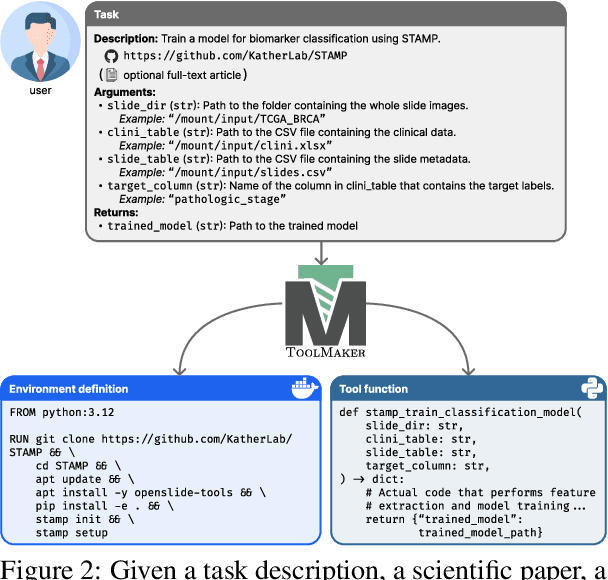


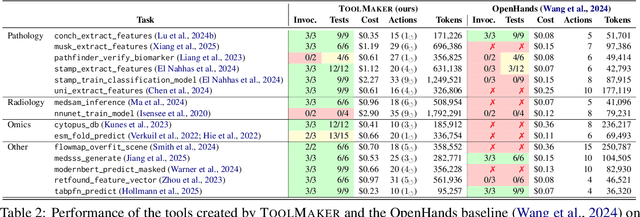
Abstract:Tool use has turned large language models (LLMs) into powerful agents that can perform complex multi-step tasks by dynamically utilising external software components. However, these tools must be implemented in advance by human developers, hindering the applicability of LLM agents in domains which demand large numbers of highly specialised tools, like in life sciences and medicine. Motivated by the growing trend of scientific studies accompanied by public code repositories, we propose ToolMaker, a novel agentic framework that autonomously transforms papers with code into LLM-compatible tools. Given a short task description and a repository URL, ToolMaker autonomously installs required dependencies and generates code to perform the task, using a closed-loop self-correction mechanism to iteratively diagnose and rectify errors. To evaluate our approach, we introduce a benchmark comprising 15 diverse and complex computational tasks spanning both medical and non-medical domains with over 100 unit tests to objectively assess tool correctness and robustness. ToolMaker correctly implements 80% of the tasks, substantially outperforming current state-of-the-art software engineering agents. ToolMaker therefore is a step towards fully autonomous agent-based scientific workflows.
Large Language Models-Enabled Digital Twins for Precision Medicine in Rare Gynecological Tumors
Aug 31, 2024Abstract:Rare gynecological tumors (RGTs) present major clinical challenges due to their low incidence and heterogeneity. The lack of clear guidelines leads to suboptimal management and poor prognosis. Molecular tumor boards accelerate access to effective therapies by tailoring treatment based on biomarkers, beyond cancer type. Unstructured data that requires manual curation hinders efficient use of biomarker profiling for therapy matching. This study explores the use of large language models (LLMs) to construct digital twins for precision medicine in RGTs. Our proof-of-concept digital twin system integrates clinical and biomarker data from institutional and published cases (n=21) and literature-derived data (n=655 publications with n=404,265 patients) to create tailored treatment plans for metastatic uterine carcinosarcoma, identifying options potentially missed by traditional, single-source analysis. LLM-enabled digital twins efficiently model individual patient trajectories. Shifting to a biology-based rather than organ-based tumor definition enables personalized care that could advance RGT management and thus enhance patient outcomes.
Prompt Injection Attacks on Large Language Models in Oncology
Jul 23, 2024



Abstract:Vision-language artificial intelligence models (VLMs) possess medical knowledge and can be employed in healthcare in numerous ways, including as image interpreters, virtual scribes, and general decision support systems. However, here, we demonstrate that current VLMs applied to medical tasks exhibit a fundamental security flaw: they can be attacked by prompt injection attacks, which can be used to output harmful information just by interacting with the VLM, without any access to its parameters. We performed a quantitative study to evaluate the vulnerabilities to these attacks in four state of the art VLMs which have been proposed to be of utility in healthcare: Claude 3 Opus, Claude 3.5 Sonnet, Reka Core, and GPT-4o. Using a set of N=297 attacks, we show that all of these models are susceptible. Specifically, we show that embedding sub-visual prompts in medical imaging data can cause the model to provide harmful output, and that these prompts are non-obvious to human observers. Thus, our study demonstrates a key vulnerability in medical VLMs which should be mitigated before widespread clinical adoption.
RadioRAG: Factual Large Language Models for Enhanced Diagnostics in Radiology Using Dynamic Retrieval Augmented Generation
Jul 22, 2024



Abstract:Large language models (LLMs) have advanced the field of artificial intelligence (AI) in medicine. However LLMs often generate outdated or inaccurate information based on static training datasets. Retrieval augmented generation (RAG) mitigates this by integrating outside data sources. While previous RAG systems used pre-assembled, fixed databases with limited flexibility, we have developed Radiology RAG (RadioRAG) as an end-to-end framework that retrieves data from authoritative radiologic online sources in real-time. RadioRAG is evaluated using a dedicated radiologic question-and-answer dataset (RadioQA). We evaluate the diagnostic accuracy of various LLMs when answering radiology-specific questions with and without access to additional online information via RAG. Using 80 questions from RSNA Case Collection across radiologic subspecialties and 24 additional expert-curated questions, for which the correct gold-standard answers were available, LLMs (GPT-3.5-turbo, GPT-4, Mistral-7B, Mixtral-8x7B, and Llama3 [8B and 70B]) were prompted with and without RadioRAG. RadioRAG retrieved context-specific information from www.radiopaedia.org in real-time and incorporated them into its reply. RadioRAG consistently improved diagnostic accuracy across all LLMs, with relative improvements ranging from 2% to 54%. It matched or exceeded question answering without RAG across radiologic subspecialties, particularly in breast imaging and emergency radiology. However, degree of improvement varied among models; GPT-3.5-turbo and Mixtral-8x7B-instruct-v0.1 saw notable gains, while Mistral-7B-instruct-v0.2 showed no improvement, highlighting variability in its effectiveness. LLMs benefit when provided access to domain-specific data beyond their training data. For radiology, RadioRAG establishes a robust framework that substantially improves diagnostic accuracy and factuality in radiological question answering.
End-To-End Clinical Trial Matching with Large Language Models
Jul 18, 2024


Abstract:Matching cancer patients to clinical trials is essential for advancing treatment and patient care. However, the inconsistent format of medical free text documents and complex trial eligibility criteria make this process extremely challenging and time-consuming for physicians. We investigated whether the entire trial matching process - from identifying relevant trials among 105,600 oncology-related clinical trials on clinicaltrials.gov to generating criterion-level eligibility matches - could be automated using Large Language Models (LLMs). Using GPT-4o and a set of 51 synthetic Electronic Health Records (EHRs), we demonstrate that our approach identifies relevant candidate trials in 93.3% of cases and achieves a preliminary accuracy of 88.0% when matching patient-level information at the criterion level against a baseline defined by human experts. Utilizing LLM feedback reveals that 39.3% criteria that were initially considered incorrect are either ambiguous or inaccurately annotated, leading to a total model accuracy of 92.7% after refining our human baseline. In summary, we present an end-to-end pipeline for clinical trial matching using LLMs, demonstrating high precision in screening and matching trials to individual patients, even outperforming the performance of qualified medical doctors. Our fully end-to-end pipeline can operate autonomously or with human supervision and is not restricted to oncology, offering a scalable solution for enhancing patient-trial matching in real-world settings.
Autonomous Artificial Intelligence Agents for Clinical Decision Making in Oncology
Apr 06, 2024

Abstract:Multimodal artificial intelligence (AI) systems have the potential to enhance clinical decision-making by interpreting various types of medical data. However, the effectiveness of these models across all medical fields is uncertain. Each discipline presents unique challenges that need to be addressed for optimal performance. This complexity is further increased when attempting to integrate different fields into a single model. Here, we introduce an alternative approach to multimodal medical AI that utilizes the generalist capabilities of a large language model (LLM) as a central reasoning engine. This engine autonomously coordinates and deploys a set of specialized medical AI tools. These tools include text, radiology and histopathology image interpretation, genomic data processing, web searches, and document retrieval from medical guidelines. We validate our system across a series of clinical oncology scenarios that closely resemble typical patient care workflows. We show that the system has a high capability in employing appropriate tools (97%), drawing correct conclusions (93.6%), and providing complete (94%), and helpful (89.2%) recommendations for individual patient cases while consistently referencing relevant literature (82.5%) upon instruction. This work provides evidence that LLMs can effectively plan and execute domain-specific models to retrieve or synthesize new information when used as autonomous agents. This enables them to function as specialist, patient-tailored clinical assistants. It also simplifies regulatory compliance by allowing each component tool to be individually validated and approved. We believe, that our work can serve as a proof-of-concept for more advanced LLM-agents in the medical domain.
In-context learning enables multimodal large language models to classify cancer pathology images
Mar 12, 2024Abstract:Medical image classification requires labeled, task-specific datasets which are used to train deep learning networks de novo, or to fine-tune foundation models. However, this process is computationally and technically demanding. In language processing, in-context learning provides an alternative, where models learn from within prompts, bypassing the need for parameter updates. Yet, in-context learning remains underexplored in medical image analysis. Here, we systematically evaluate the model Generative Pretrained Transformer 4 with Vision capabilities (GPT-4V) on cancer image processing with in-context learning on three cancer histopathology tasks of high importance: Classification of tissue subtypes in colorectal cancer, colon polyp subtyping and breast tumor detection in lymph node sections. Our results show that in-context learning is sufficient to match or even outperform specialized neural networks trained for particular tasks, while only requiring a minimal number of samples. In summary, this study demonstrates that large vision language models trained on non-domain specific data can be applied out-of-the box to solve medical image-processing tasks in histopathology. This democratizes access of generalist AI models to medical experts without technical background especially for areas where annotated data is scarce.
A Good Feature Extractor Is All You Need for Weakly Supervised Learning in Histopathology
Nov 29, 2023Abstract:Deep learning is revolutionising pathology, offering novel opportunities in disease prognosis and personalised treatment. Historically, stain normalisation has been a crucial preprocessing step in computational pathology pipelines, and persists into the deep learning era. Yet, with the emergence of feature extractors trained using self-supervised learning (SSL) on diverse pathology datasets, we call this practice into question. In an empirical evaluation of publicly available feature extractors, we find that omitting stain normalisation and image augmentations does not compromise downstream performance, while incurring substantial savings in memory and compute. Further, we show that the top-performing feature extractors are remarkably robust to variations in stain and augmentations like rotation in their latent space. Contrary to previous patch-level benchmarking studies, our approach emphasises clinical relevance by focusing on slide-level prediction tasks in a weakly supervised setting with external validation cohorts. This work represents the most comprehensive robustness evaluation of public pathology SSL feature extractors to date, involving more than 6,000 training runs across nine tasks, five datasets, three downstream architectures, and various preprocessing setups. Our findings stand to streamline digital pathology workflows by minimising preprocessing needs and informing the selection of feature extractors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge