Behrus Puladi
Enhancing Privacy: The Utility of Stand-Alone Synthetic CT and MRI for Tumor and Bone Segmentation
Jun 13, 2025Abstract:AI requires extensive datasets, while medical data is subject to high data protection. Anonymization is essential, but poses a challenge for some regions, such as the head, as identifying structures overlap with regions of clinical interest. Synthetic data offers a potential solution, but studies often lack rigorous evaluation of realism and utility. Therefore, we investigate to what extent synthetic data can replace real data in segmentation tasks. We employed head and neck cancer CT scans and brain glioma MRI scans from two large datasets. Synthetic data were generated using generative adversarial networks and diffusion models. We evaluated the quality of the synthetic data using MAE, MS-SSIM, Radiomics and a Visual Turing Test (VTT) performed by 5 radiologists and their usefulness in segmentation tasks using DSC. Radiomics indicates high fidelity of synthetic MRIs, but fall short in producing highly realistic CT tissue, with correlation coefficient of 0.8784 and 0.5461 for MRI and CT tumors, respectively. DSC results indicate limited utility of synthetic data: tumor segmentation achieved DSC=0.064 on CT and 0.834 on MRI, while bone segmentation a mean DSC=0.841. Relation between DSC and correlation is observed, but is limited by the complexity of the task. VTT results show synthetic CTs' utility, but with limited educational applications. Synthetic data can be used independently for the segmentation task, although limited by the complexity of the structures to segment. Advancing generative models to better tolerate heterogeneous inputs and learn subtle details is essential for enhancing their realism and expanding their application potential.
Efficient MedSAMs: Segment Anything in Medical Images on Laptop
Dec 20, 2024


Abstract:Promptable segmentation foundation models have emerged as a transformative approach to addressing the diverse needs in medical images, but most existing models require expensive computing, posing a big barrier to their adoption in clinical practice. In this work, we organized the first international competition dedicated to promptable medical image segmentation, featuring a large-scale dataset spanning nine common imaging modalities from over 20 different institutions. The top teams developed lightweight segmentation foundation models and implemented an efficient inference pipeline that substantially reduced computational requirements while maintaining state-of-the-art segmentation accuracy. Moreover, the post-challenge phase advanced the algorithms through the design of performance booster and reproducibility tasks, resulting in improved algorithms and validated reproducibility of the winning solution. Furthermore, the best-performing algorithms have been incorporated into the open-source software with a user-friendly interface to facilitate clinical adoption. The data and code are publicly available to foster the further development of medical image segmentation foundation models and pave the way for impactful real-world applications.
Comparative Analysis of nnUNet and MedNeXt for Head and Neck Tumor Segmentation in MRI-guided Radiotherapy
Nov 22, 2024Abstract:Radiation therapy (RT) is essential in treating head and neck cancer (HNC), with magnetic resonance imaging(MRI)-guided RT offering superior soft tissue contrast and functional imaging. However, manual tumor segmentation is time-consuming and complex, and therfore remains a challenge. In this study, we present our solution as team TUMOR to the HNTS-MRG24 MICCAI Challenge which is focused on automated segmentation of primary gross tumor volumes (GTVp) and metastatic lymph node gross tumor volume (GTVn) in pre-RT and mid-RT MRI images. We utilized the HNTS-MRG2024 dataset, which consists of 150 MRI scans from patients diagnosed with HNC, including original and registered pre-RT and mid-RT T2-weighted images with corresponding segmentation masks for GTVp and GTVn. We employed two state-of-the-art models in deep learning, nnUNet and MedNeXt. For Task 1, we pretrained models on pre-RT registered and mid-RT images, followed by fine-tuning on original pre-RT images. For Task 2, we combined registered pre-RT images, registered pre-RT segmentation masks, and mid-RT data as a multi-channel input for training. Our solution for Task 1 achieved 1st place in the final test phase with an aggregated Dice Similarity Coefficient of 0.8254, and our solution for Task 2 ranked 8th with a score of 0.7005. The proposed solution is publicly available at Github Repository.
Improved Multi-Task Brain Tumour Segmentation with Synthetic Data Augmentation
Nov 07, 2024



Abstract:This paper presents the winning solution of task 1 and the third-placed solution of task 3 of the BraTS challenge. The use of automated tools in clinical practice has increased due to the development of more and more sophisticated and reliable algorithms. However, achieving clinical standards and developing tools for real-life scenarios is a major challenge. To this end, BraTS has organised tasks to find the most advanced solutions for specific purposes. In this paper, we propose the use of synthetic data to train state-of-the-art frameworks in order to improve the segmentation of adult gliomas in a post-treatment scenario, and the segmentation of meningioma for radiotherapy planning. Our results suggest that the use of synthetic data leads to more robust algorithms, although the synthetic data generation pipeline is not directly suited to the meningioma task. The code for these tasks is available at https://github.com/ShadowTwin41/BraTS_2023_2024_solutions.
Brain Tumour Removing and Missing Modality Generation using 3D WDM
Nov 07, 2024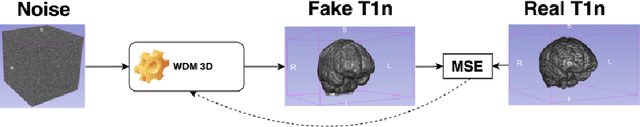
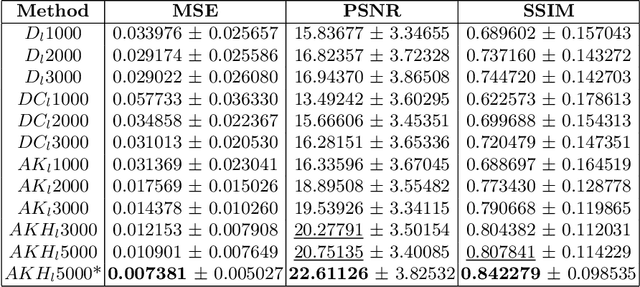
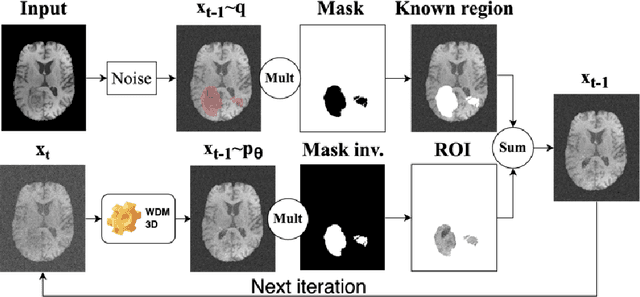
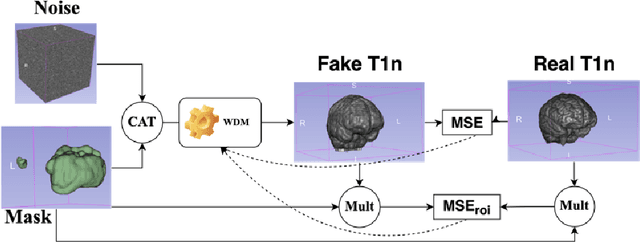
Abstract:This paper presents the second-placed solution for task 8 and the participation solution for task 7 of BraTS 2024. The adoption of automated brain analysis algorithms to support clinical practice is increasing. However, many of these algorithms struggle with the presence of brain lesions or the absence of certain MRI modalities. The alterations in the brain's morphology leads to high variability and thus poor performance of predictive models that were trained only on healthy brains. The lack of information that is usually provided by some of the missing MRI modalities also reduces the reliability of the prediction models trained with all modalities. In order to improve the performance of these models, we propose the use of conditional 3D wavelet diffusion models. The wavelet transform enabled full-resolution image training and prediction on a GPU with 48 GB VRAM, without patching or downsampling, preserving all information for prediction. For the inpainting task of BraTS 2024, the use of a large and variable number of healthy masks and the stability and efficiency of the 3D wavelet diffusion model resulted in 0.007, 22.61 and 0.842 in the validation set and 0.07 , 22.8 and 0.91 in the testing set (MSE, PSNR and SSIM respectively). The code for these tasks is available at https://github.com/ShadowTwin41/BraTS_2023_2024_solutions.
Deep Dive into MRI: Exploring Deep Learning Applications in 0.55T and 7T MRI
Jul 01, 2024



Abstract:The development of magnetic resonance imaging (MRI) for medical imaging has provided a leap forward in diagnosis, providing a safe, non-invasive alternative to techniques involving ionising radiation exposure for diagnostic purposes. It was described by Block and Purcel in 1946, and it was not until 1980 that the first clinical application of MRI became available. Since that time the MRI has gone through many advances and has altered the way diagnosing procedures are performed. Due to its ability to improve constantly, MRI has become a commonly used practice among several specialisations in medicine. Particularly starting 0.55T and 7T MRI technologies have pointed out enhanced preservation of image detail and advanced tissue characterisation. This review examines the integration of deep learning (DL) techniques into these MRI modalities, disseminating and exploring the study applications. It highlights how DL contributes to 0.55T and 7T MRI data, showcasing the potential of DL in improving and refining these technologies. The review ends with a brief overview of how MRI technology will evolve in the coming years.
An Ordinal Regression Framework for a Deep Learning Based Severity Assessment for Chest Radiographs
Feb 08, 2024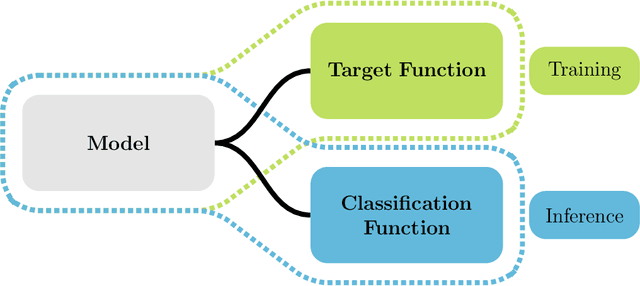
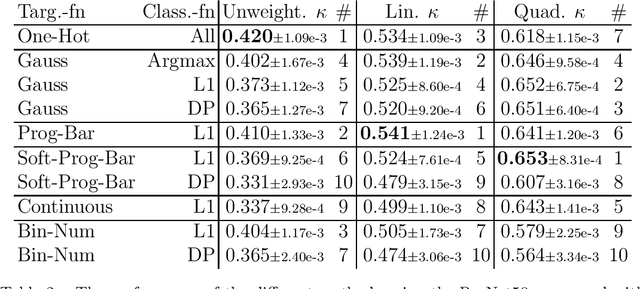
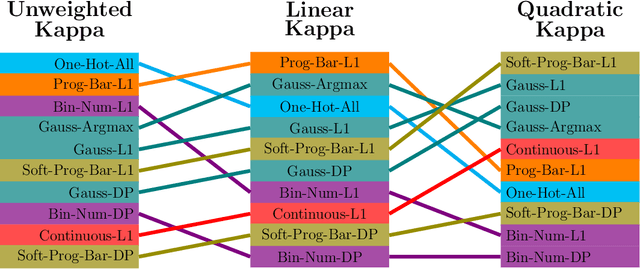
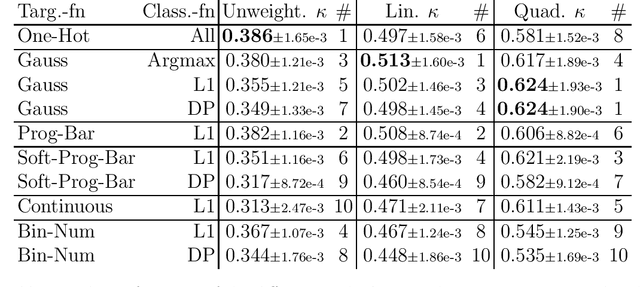
Abstract:This study investigates the application of ordinal regression methods for categorizing disease severity in chest radiographs. We propose a framework that divides the ordinal regression problem into three parts: a model, a target function, and a classification function. Different encoding methods, including one-hot, Gaussian, progress-bar, and our soft-progress-bar, are applied using ResNet50 and ViT-B-16 deep learning models. We show that the choice of encoding has a strong impact on performance and that the best encoding depends on the chosen weighting of Cohen's kappa and also on the model architecture used. We make our code publicly available on GitHub.
Deep PCCT: Photon Counting Computed Tomography Deep Learning Applications Review
Feb 06, 2024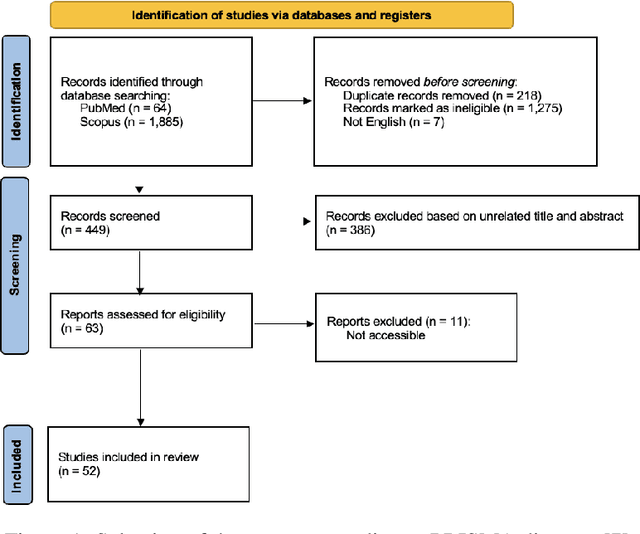
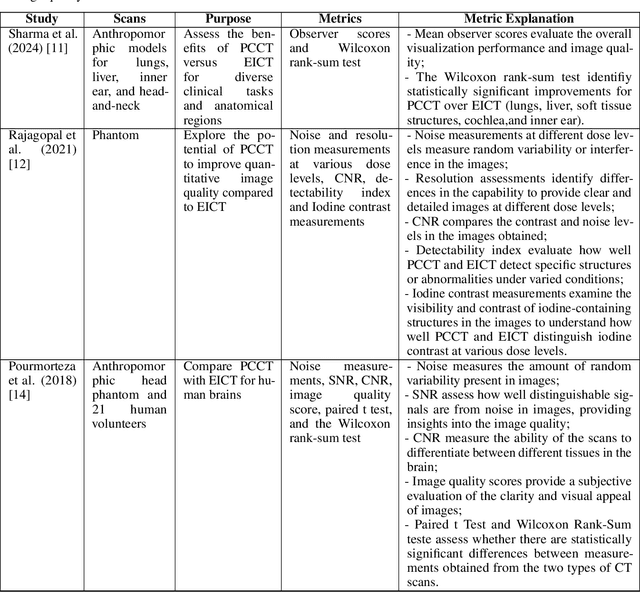
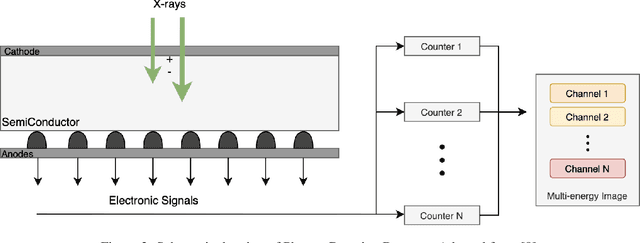
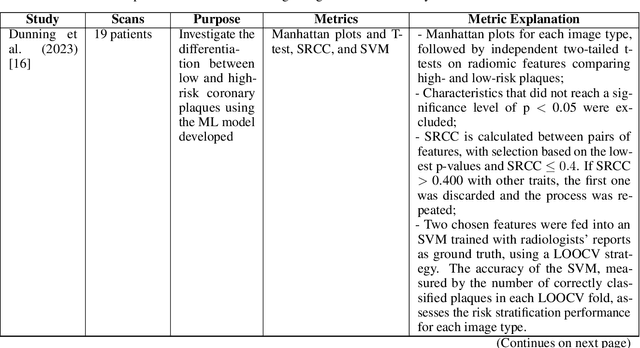
Abstract:Medical imaging faces challenges such as limited spatial resolution, interference from electronic noise and poor contrast-to-noise ratios. Photon Counting Computed Tomography (PCCT) has emerged as a solution, addressing these issues with its innovative technology. This review delves into the recent developments and applications of PCCT in pre-clinical research, emphasizing its potential to overcome traditional imaging limitations. For example PCCT has demonstrated remarkable efficacy in improving the detection of subtle abnormalities in breast, providing a level of detail previously unattainable. Examining the current literature on PCCT, it presents a comprehensive analysis of the technology, highlighting the main features of scanners and their varied applications. In addition, it explores the integration of deep learning into PCCT, along with the study of radiomic features, presenting successful applications in data processing. While acknowledging these advances, it also discusses the existing challenges in this field, paving the way for future research and improvements in medical imaging technologies. Despite the limited number of articles on this subject, due to the recent integration of PCCT at a clinical level, its potential benefits extend to various diagnostic applications.
Cyto R-CNN and CytoNuke Dataset: Towards reliable whole-cell segmentation in bright-field histological images
Feb 04, 2024Abstract:Background: Cell segmentation in bright-field histological slides is a crucial topic in medical image analysis. Having access to accurate segmentation allows researchers to examine the relationship between cellular morphology and clinical observations. Unfortunately, most segmentation methods known today are limited to nuclei and cannot segmentate the cytoplasm. Material & Methods: We present a new network architecture Cyto R-CNN that is able to accurately segment whole cells (with both the nucleus and the cytoplasm) in bright-field images. We also present a new dataset CytoNuke, consisting of multiple thousand manual annotations of head and neck squamous cell carcinoma cells. Utilizing this dataset, we compared the performance of Cyto R-CNN to other popular cell segmentation algorithms, including QuPath's built-in algorithm, StarDist and Cellpose. To evaluate segmentation performance, we calculated AP50, AP75 and measured 17 morphological and staining-related features for all detected cells. We compared these measurements to the gold standard of manual segmentation using the Kolmogorov-Smirnov test. Results: Cyto R-CNN achieved an AP50 of 58.65% and an AP75 of 11.56% in whole-cell segmentation, outperforming all other methods (QuPath $19.46/0.91\%$; StarDist $45.33/2.32\%$; Cellpose $31.85/5.61\%$). Cell features derived from Cyto R-CNN showed the best agreement to the gold standard ($\bar{D} = 0.15$) outperforming QuPath ($\bar{D} = 0.22$), StarDist ($\bar{D} = 0.25$) and Cellpose ($\bar{D} = 0.23$). Conclusion: Our newly proposed Cyto R-CNN architecture outperforms current algorithms in whole-cell segmentation while providing more reliable cell measurements than any other model. This could improve digital pathology workflows, potentially leading to improved diagnosis. Moreover, our published dataset can be used to develop further models in the future.
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
Sep 12, 2023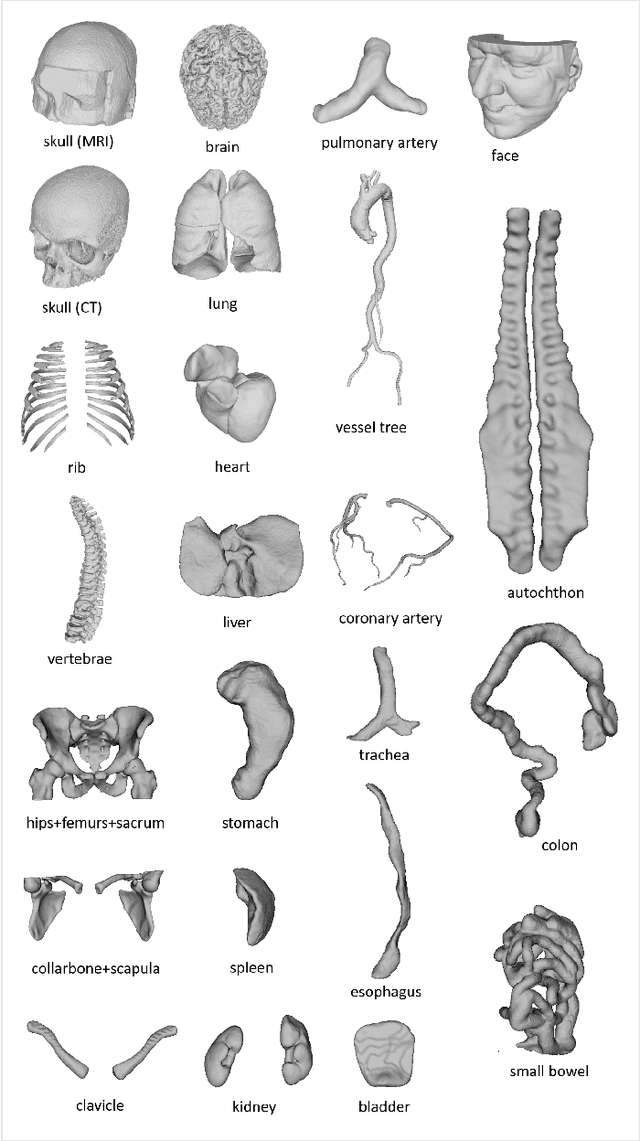

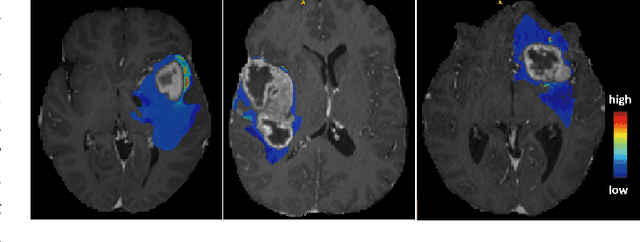
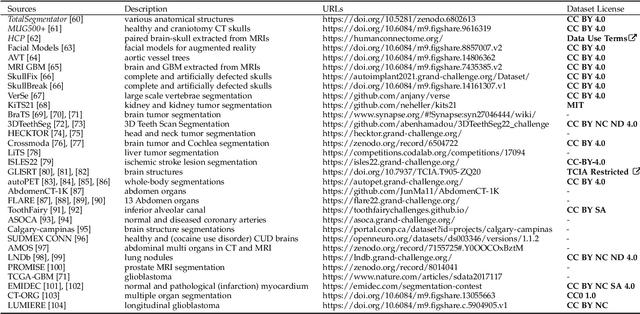
Abstract:We present MedShapeNet, a large collection of anatomical shapes (e.g., bones, organs, vessels) and 3D surgical instrument models. Prior to the deep learning era, the broad application of statistical shape models (SSMs) in medical image analysis is evidence that shapes have been commonly used to describe medical data. Nowadays, however, state-of-the-art (SOTA) deep learning algorithms in medical imaging are predominantly voxel-based. In computer vision, on the contrary, shapes (including, voxel occupancy grids, meshes, point clouds and implicit surface models) are preferred data representations in 3D, as seen from the numerous shape-related publications in premier vision conferences, such as the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), as well as the increasing popularity of ShapeNet (about 51,300 models) and Princeton ModelNet (127,915 models) in computer vision research. MedShapeNet is created as an alternative to these commonly used shape benchmarks to facilitate the translation of data-driven vision algorithms to medical applications, and it extends the opportunities to adapt SOTA vision algorithms to solve critical medical problems. Besides, the majority of the medical shapes in MedShapeNet are modeled directly on the imaging data of real patients, and therefore it complements well existing shape benchmarks comprising of computer-aided design (CAD) models. MedShapeNet currently includes more than 100,000 medical shapes, and provides annotations in the form of paired data. It is therefore also a freely available repository of 3D models for extended reality (virtual reality - VR, augmented reality - AR, mixed reality - MR) and medical 3D printing. This white paper describes in detail the motivations behind MedShapeNet, the shape acquisition procedures, the use cases, as well as the usage of the online shape search portal: https://medshapenet.ikim.nrw/
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge