Fabian Hörst
Efficient Complex-Valued Vision Transformers for MRI Classification Directly from k-Space
Jan 26, 2026Abstract:Deep learning applications in Magnetic Resonance Imaging (MRI) predominantly operate on reconstructed magnitude images, a process that discards phase information and requires computationally expensive transforms. Standard neural network architectures rely on local operations (convolutions or grid-patches) that are ill-suited for the global, non-local nature of raw frequency-domain (k-Space) data. In this work, we propose a novel complex-valued Vision Transformer (kViT) designed to perform classification directly on k-Space data. To bridge the geometric disconnect between current architectures and MRI physics, we introduce a radial k-Space patching strategy that respects the spectral energy distribution of the frequency-domain. Extensive experiments on the fastMRI and in-house datasets demonstrate that our approach achieves classification performance competitive with state-of-the-art image-domain baselines (ResNet, EfficientNet, ViT). Crucially, kViT exhibits superior robustness to high acceleration factors and offers a paradigm shift in computational efficiency, reducing VRAM consumption during training by up to 68$\times$ compared to standard methods. This establishes a pathway for resource-efficient, direct-from-scanner AI analysis.
CT-GRAPH: Hierarchical Graph Attention Network for Anatomy-Guided CT Report Generation
Aug 07, 2025Abstract:As medical imaging is central to diagnostic processes, automating the generation of radiology reports has become increasingly relevant to assist radiologists with their heavy workloads. Most current methods rely solely on global image features, failing to capture fine-grained organ relationships crucial for accurate reporting. To this end, we propose CT-GRAPH, a hierarchical graph attention network that explicitly models radiological knowledge by structuring anatomical regions into a graph, linking fine-grained organ features to coarser anatomical systems and a global patient context. Our method leverages pretrained 3D medical feature encoders to obtain global and organ-level features by utilizing anatomical masks. These features are further refined within the graph and then integrated into a large language model to generate detailed medical reports. We evaluate our approach for the task of report generation on the large-scale chest CT dataset CT-RATE. We provide an in-depth analysis of pretrained feature encoders for CT report generation and show that our method achieves a substantial improvement of absolute 7.9\% in F1 score over current state-of-the-art methods. The code is publicly available at https://github.com/hakal104/CT-GRAPH.
PhaseGen: A Diffusion-Based Approach for Complex-Valued MRI Data Generation
Apr 10, 2025Abstract:Magnetic resonance imaging (MRI) raw data, or k-Space data, is complex-valued, containing both magnitude and phase information. However, clinical and existing Artificial Intelligence (AI)-based methods focus only on magnitude images, discarding the phase data despite its potential for downstream tasks, such as tumor segmentation and classification. In this work, we introduce $\textit{PhaseGen}$, a novel complex-valued diffusion model for generating synthetic MRI raw data conditioned on magnitude images, commonly used in clinical practice. This enables the creation of artificial complex-valued raw data, allowing pretraining for models that require k-Space information. We evaluate PhaseGen on two tasks: skull-stripping directly in k-Space and MRI reconstruction using the publicly available FastMRI dataset. Our results show that training with synthetic phase data significantly improves generalization for skull-stripping on real-world data, with an increased segmentation accuracy from $41.1\%$ to $80.1\%$, and enhances MRI reconstruction when combined with limited real-world data. This work presents a step forward in utilizing generative AI to bridge the gap between magnitude-based datasets and the complex-valued nature of MRI raw data. This approach allows researchers to leverage the vast amount of avaliable image domain data in combination with the information-rich k-Space data for more accurate and efficient diagnostic tasks. We make our code publicly $\href{https://github.com/TIO-IKIM/PhaseGen}{\text{available here}}$.
Cracking the PUMA Challenge in 24 Hours with CellViT++ and nnU-Net
Mar 15, 2025
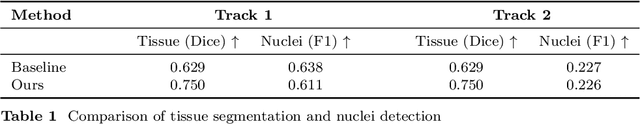
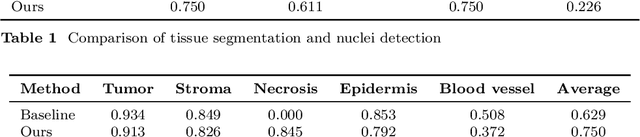
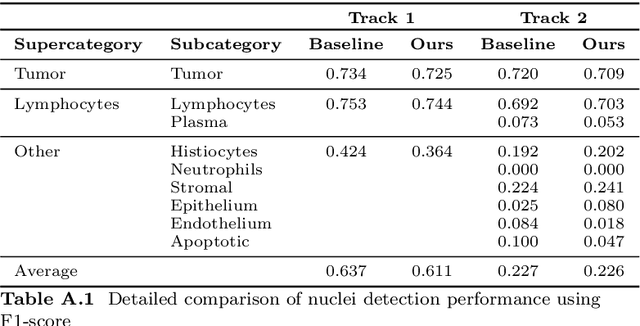
Abstract:Automatic tissue segmentation and nuclei detection is an important task in pathology, aiding in biomarker extraction and discovery. The panoptic segmentation of nuclei and tissue in advanced melanoma (PUMA) challenge aims to improve tissue segmentation and nuclei detection in melanoma histopathology. Unlike many challenge submissions focusing on extensive model tuning, our approach emphasizes delivering a deployable solution within a 24-hour development timeframe, using out-of-the-box frameworks. The pipeline combines two models, namely CellViT++ for nuclei detection and nnU-Net for tissue segmentation. Our results demonstrate a significant improvement in tissue segmentation, achieving a Dice score of 0.750, surpassing the baseline score of 0.629. For nuclei detection, we obtained results comparable to the baseline in both challenge tracks. The code is publicly available at https://github.com/TIO-IKIM/PUMA.
CellViT++: Energy-Efficient and Adaptive Cell Segmentation and Classification Using Foundation Models
Jan 09, 2025



Abstract:Digital Pathology is a cornerstone in the diagnosis and treatment of diseases. A key task in this field is the identification and segmentation of cells in hematoxylin and eosin-stained images. Existing methods for cell segmentation often require extensive annotated datasets for training and are limited to a predefined cell classification scheme. To overcome these limitations, we propose $\text{CellViT}^{{\scriptscriptstyle ++}}$, a framework for generalized cell segmentation in digital pathology. $\text{CellViT}^{{\scriptscriptstyle ++}}$ utilizes Vision Transformers with foundation models as encoders to compute deep cell features and segmentation masks simultaneously. To adapt to unseen cell types, we rely on a computationally efficient approach. It requires minimal data for training and leads to a drastically reduced carbon footprint. We demonstrate excellent performance on seven different datasets, covering a broad spectrum of cell types, organs, and clinical settings. The framework achieves remarkable zero-shot segmentation and data-efficient cell-type classification. Furthermore, we show that $\text{CellViT}^{{\scriptscriptstyle ++}}$ can leverage immunofluorescence stainings to generate training datasets without the need for pathologist annotations. The automated dataset generation approach surpasses the performance of networks trained on manually labeled data, demonstrating its effectiveness in creating high-quality training datasets without expert annotations. To advance digital pathology, $\text{CellViT}^{{\scriptscriptstyle ++}}$ is available as an open-source framework featuring a user-friendly, web-based interface for visualization and annotation. The code is available under https://github.com/TIO-IKIM/CellViT-plus-plus.
De-Identification of Medical Imaging Data: A Comprehensive Tool for Ensuring Patient Privacy
Oct 16, 2024



Abstract:Medical data employed in research frequently comprises sensitive patient health information (PHI), which is subject to rigorous legal frameworks such as the General Data Protection Regulation (GDPR) or the Health Insurance Portability and Accountability Act (HIPAA). Consequently, these types of data must be pseudonymized prior to utilisation, which presents a significant challenge for many researchers. Given the vast array of medical data, it is necessary to employ a variety of de-identification techniques. To facilitate the anonymization process for medical imaging data, we have developed an open-source tool that can be used to de-identify DICOM magnetic resonance images, computer tomography images, whole slide images and magnetic resonance twix raw data. Furthermore, the implementation of a neural network enables the removal of text within the images. The proposed tool automates an elaborate anonymization pipeline for multiple types of inputs, reducing the need for additional tools used for de-identification of imaging data. We make our code publicly available at https://github.com/code-lukas/medical_image_deidentification.
Spacewalker: Traversing Representation Spaces for Fast Interactive Exploration and Annotation of Unstructured Data
Sep 25, 2024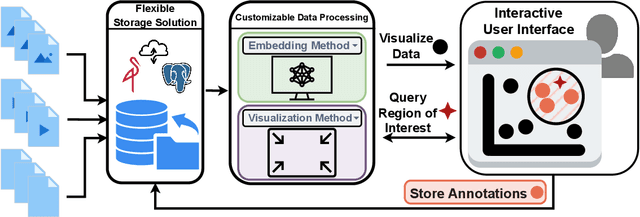
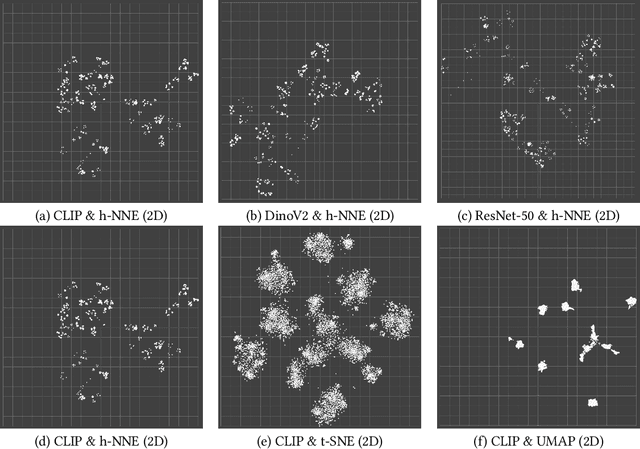
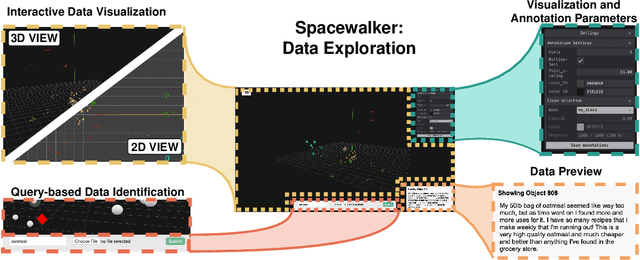

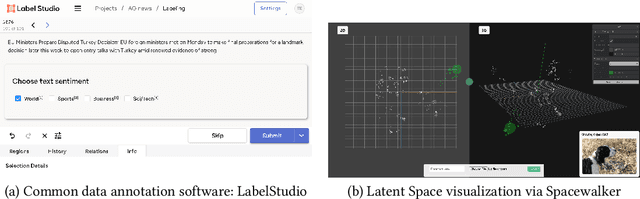
Abstract:Unstructured data in industries such as healthcare, finance, and manufacturing presents significant challenges for efficient analysis and decision making. Detecting patterns within this data and understanding their impact is critical but complex without the right tools. Traditionally, these tasks relied on the expertise of data analysts or labor-intensive manual reviews. In response, we introduce Spacewalker, an interactive tool designed to explore and annotate data across multiple modalities. Spacewalker allows users to extract data representations and visualize them in low-dimensional spaces, enabling the detection of semantic similarities. Through extensive user studies, we assess Spacewalker's effectiveness in data annotation and integrity verification. Results show that the tool's ability to traverse latent spaces and perform multi-modal queries significantly enhances the user's capacity to quickly identify relevant data. Moreover, Spacewalker allows for annotation speed-ups far superior to conventional methods, making it a promising tool for efficiently navigating unstructured data and improving decision making processes. The code of this work is open-source and can be found at: https://github.com/code-lukas/Spacewalker
Autopet III challenge: Incorporating anatomical knowledge into nnUNet for lesion segmentation in PET/CT
Sep 18, 2024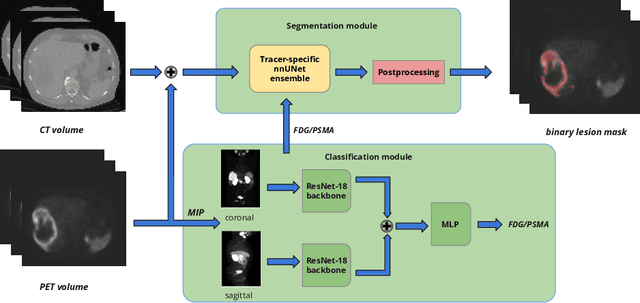
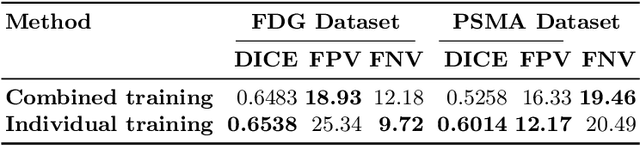
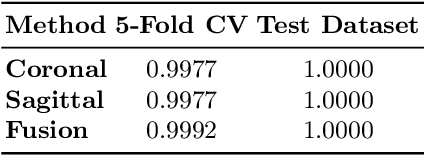

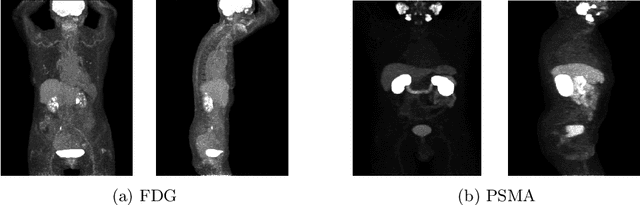

Abstract:Lesion segmentation in PET/CT imaging is essential for precise tumor characterization, which supports personalized treatment planning and enhances diagnostic precision in oncology. However, accurate manual segmentation of lesions is time-consuming and prone to inter-observer variability. Given the rising demand and clinical use of PET/CT, automated segmentation methods, particularly deep-learning-based approaches, have become increasingly more relevant. The autoPET III Challenge focuses on advancing automated segmentation of tumor lesions in PET/CT images in a multitracer multicenter setting, addressing the clinical need for quantitative, robust, and generalizable solutions. Building on previous challenges, the third iteration of the autoPET challenge introduces a more diverse dataset featuring two different tracers (FDG and PSMA) from two clinical centers. To this extent, we developed a classifier that identifies the tracer of the given PET/CT based on the Maximum Intensity Projection of the PET scan. We trained two individual nnUNet-ensembles for each tracer where anatomical labels are included as a multi-label task to enhance the model's performance. Our final submission achieves cross-validation Dice scores of 76.90% and 61.33% for the publicly available FDG and PSMA datasets, respectively. The code is available at https://github.com/hakal104/autoPETIII/ .
Cyto R-CNN and CytoNuke Dataset: Towards reliable whole-cell segmentation in bright-field histological images
Feb 04, 2024Abstract:Background: Cell segmentation in bright-field histological slides is a crucial topic in medical image analysis. Having access to accurate segmentation allows researchers to examine the relationship between cellular morphology and clinical observations. Unfortunately, most segmentation methods known today are limited to nuclei and cannot segmentate the cytoplasm. Material & Methods: We present a new network architecture Cyto R-CNN that is able to accurately segment whole cells (with both the nucleus and the cytoplasm) in bright-field images. We also present a new dataset CytoNuke, consisting of multiple thousand manual annotations of head and neck squamous cell carcinoma cells. Utilizing this dataset, we compared the performance of Cyto R-CNN to other popular cell segmentation algorithms, including QuPath's built-in algorithm, StarDist and Cellpose. To evaluate segmentation performance, we calculated AP50, AP75 and measured 17 morphological and staining-related features for all detected cells. We compared these measurements to the gold standard of manual segmentation using the Kolmogorov-Smirnov test. Results: Cyto R-CNN achieved an AP50 of 58.65% and an AP75 of 11.56% in whole-cell segmentation, outperforming all other methods (QuPath $19.46/0.91\%$; StarDist $45.33/2.32\%$; Cellpose $31.85/5.61\%$). Cell features derived from Cyto R-CNN showed the best agreement to the gold standard ($\bar{D} = 0.15$) outperforming QuPath ($\bar{D} = 0.22$), StarDist ($\bar{D} = 0.25$) and Cellpose ($\bar{D} = 0.23$). Conclusion: Our newly proposed Cyto R-CNN architecture outperforms current algorithms in whole-cell segmentation while providing more reliable cell measurements than any other model. This could improve digital pathology workflows, potentially leading to improved diagnosis. Moreover, our published dataset can be used to develop further models in the future.
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
Sep 12, 2023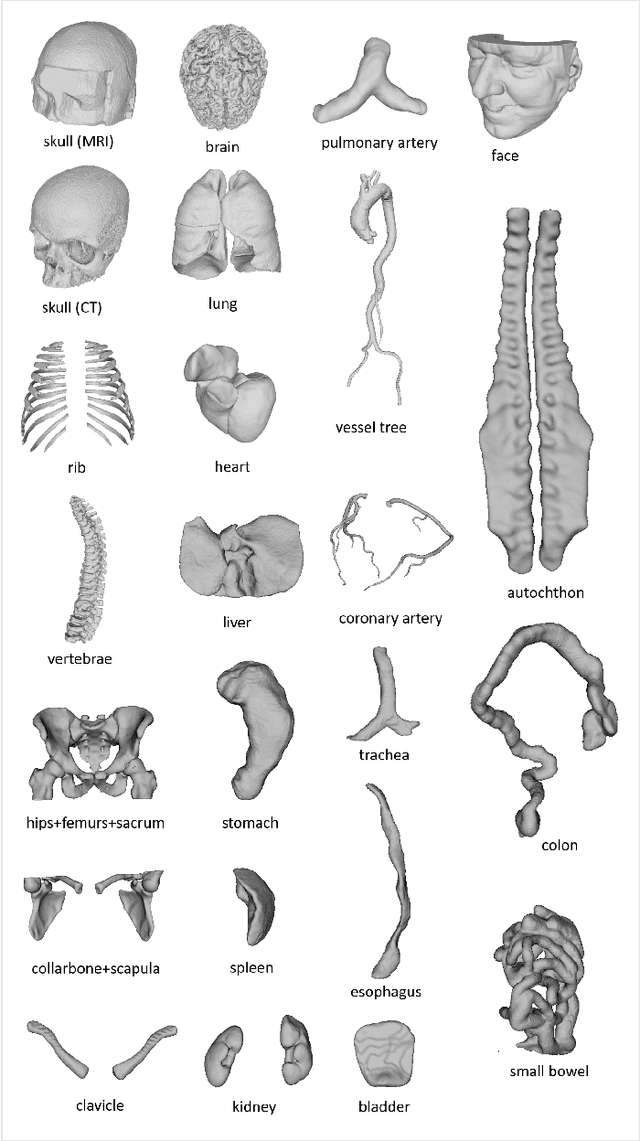

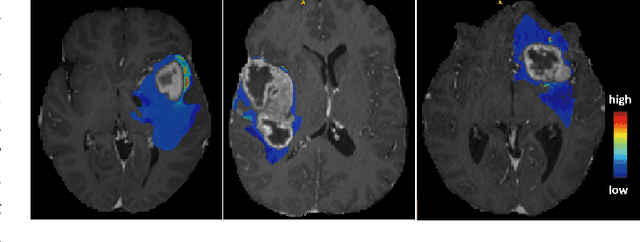
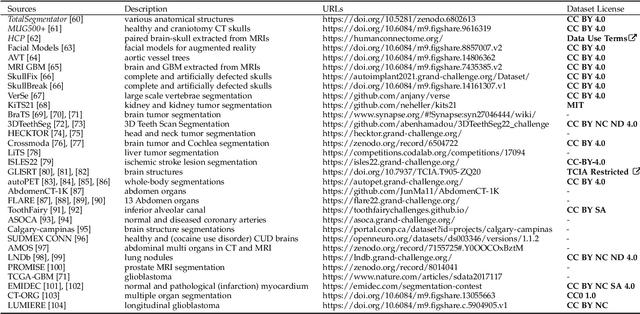
Abstract:We present MedShapeNet, a large collection of anatomical shapes (e.g., bones, organs, vessels) and 3D surgical instrument models. Prior to the deep learning era, the broad application of statistical shape models (SSMs) in medical image analysis is evidence that shapes have been commonly used to describe medical data. Nowadays, however, state-of-the-art (SOTA) deep learning algorithms in medical imaging are predominantly voxel-based. In computer vision, on the contrary, shapes (including, voxel occupancy grids, meshes, point clouds and implicit surface models) are preferred data representations in 3D, as seen from the numerous shape-related publications in premier vision conferences, such as the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), as well as the increasing popularity of ShapeNet (about 51,300 models) and Princeton ModelNet (127,915 models) in computer vision research. MedShapeNet is created as an alternative to these commonly used shape benchmarks to facilitate the translation of data-driven vision algorithms to medical applications, and it extends the opportunities to adapt SOTA vision algorithms to solve critical medical problems. Besides, the majority of the medical shapes in MedShapeNet are modeled directly on the imaging data of real patients, and therefore it complements well existing shape benchmarks comprising of computer-aided design (CAD) models. MedShapeNet currently includes more than 100,000 medical shapes, and provides annotations in the form of paired data. It is therefore also a freely available repository of 3D models for extended reality (virtual reality - VR, augmented reality - AR, mixed reality - MR) and medical 3D printing. This white paper describes in detail the motivations behind MedShapeNet, the shape acquisition procedures, the use cases, as well as the usage of the online shape search portal: https://medshapenet.ikim.nrw/
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge