Jana Fragemann
Spacewalker: Traversing Representation Spaces for Fast Interactive Exploration and Annotation of Unstructured Data
Sep 25, 2024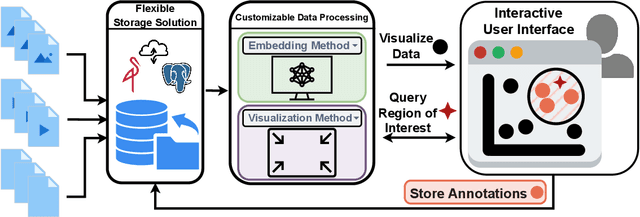
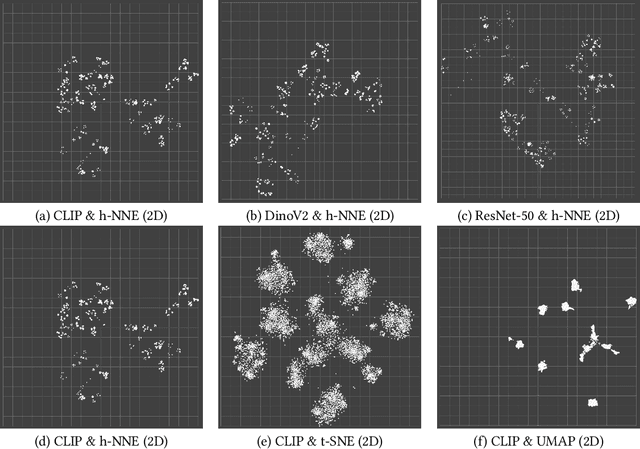
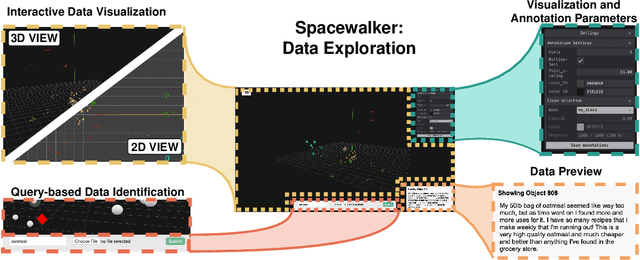

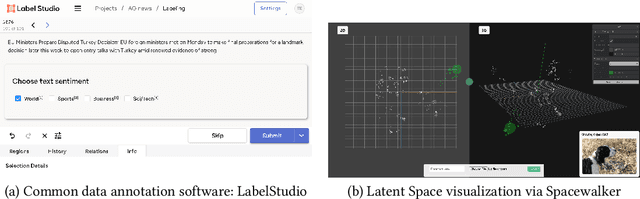
Abstract:Unstructured data in industries such as healthcare, finance, and manufacturing presents significant challenges for efficient analysis and decision making. Detecting patterns within this data and understanding their impact is critical but complex without the right tools. Traditionally, these tasks relied on the expertise of data analysts or labor-intensive manual reviews. In response, we introduce Spacewalker, an interactive tool designed to explore and annotate data across multiple modalities. Spacewalker allows users to extract data representations and visualize them in low-dimensional spaces, enabling the detection of semantic similarities. Through extensive user studies, we assess Spacewalker's effectiveness in data annotation and integrity verification. Results show that the tool's ability to traverse latent spaces and perform multi-modal queries significantly enhances the user's capacity to quickly identify relevant data. Moreover, Spacewalker allows for annotation speed-ups far superior to conventional methods, making it a promising tool for efficiently navigating unstructured data and improving decision making processes. The code of this work is open-source and can be found at: https://github.com/code-lukas/Spacewalker
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
Sep 12, 2023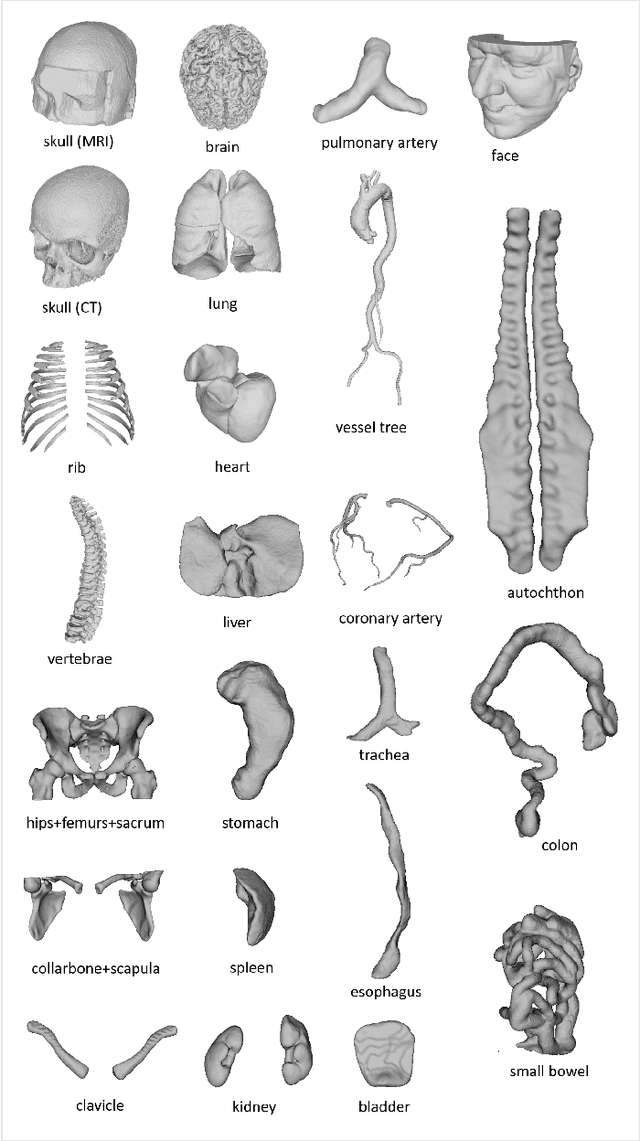

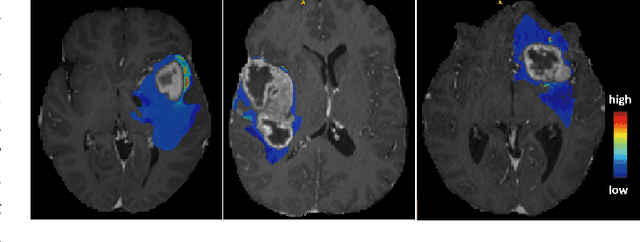
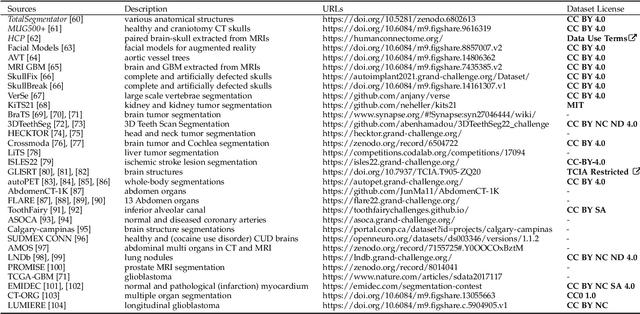
Abstract:We present MedShapeNet, a large collection of anatomical shapes (e.g., bones, organs, vessels) and 3D surgical instrument models. Prior to the deep learning era, the broad application of statistical shape models (SSMs) in medical image analysis is evidence that shapes have been commonly used to describe medical data. Nowadays, however, state-of-the-art (SOTA) deep learning algorithms in medical imaging are predominantly voxel-based. In computer vision, on the contrary, shapes (including, voxel occupancy grids, meshes, point clouds and implicit surface models) are preferred data representations in 3D, as seen from the numerous shape-related publications in premier vision conferences, such as the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), as well as the increasing popularity of ShapeNet (about 51,300 models) and Princeton ModelNet (127,915 models) in computer vision research. MedShapeNet is created as an alternative to these commonly used shape benchmarks to facilitate the translation of data-driven vision algorithms to medical applications, and it extends the opportunities to adapt SOTA vision algorithms to solve critical medical problems. Besides, the majority of the medical shapes in MedShapeNet are modeled directly on the imaging data of real patients, and therefore it complements well existing shape benchmarks comprising of computer-aided design (CAD) models. MedShapeNet currently includes more than 100,000 medical shapes, and provides annotations in the form of paired data. It is therefore also a freely available repository of 3D models for extended reality (virtual reality - VR, augmented reality - AR, mixed reality - MR) and medical 3D printing. This white paper describes in detail the motivations behind MedShapeNet, the shape acquisition procedures, the use cases, as well as the usage of the online shape search portal: https://medshapenet.ikim.nrw/
Training β-VAE by Aggregating a Learned Gaussian Posterior with a Decoupled Decoder
Sep 29, 2022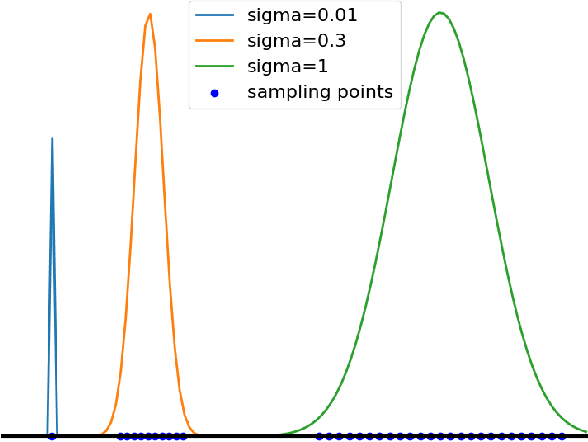
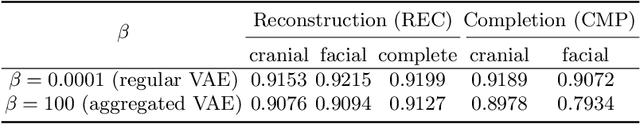
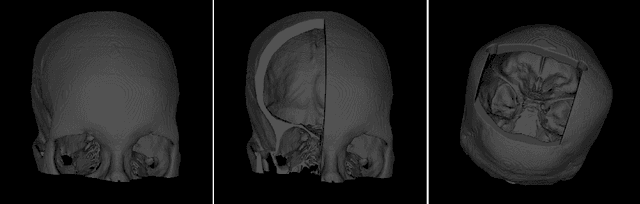
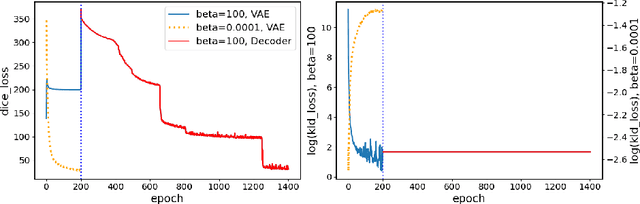
Abstract:The reconstruction loss and the Kullback-Leibler divergence (KLD) loss in a variational autoencoder (VAE) often play antagonistic roles, and tuning the weight of the KLD loss in $\beta$-VAE to achieve a balance between the two losses is a tricky and dataset-specific task. As a result, current practices in VAE training often result in a trade-off between the reconstruction fidelity and the continuity$/$disentanglement of the latent space, if the weight $\beta$ is not carefully tuned. In this paper, we present intuitions and a careful analysis of the antagonistic mechanism of the two losses, and propose, based on the insights, a simple yet effective two-stage method for training a VAE. Specifically, the method aggregates a learned Gaussian posterior $z \sim q_{\theta} (z|x)$ with a decoder decoupled from the KLD loss, which is trained to learn a new conditional distribution $p_{\phi} (x|z)$ of the input data $x$. Experimentally, we show that the aggregated VAE maximally satisfies the Gaussian assumption about the latent space, while still achieves a reconstruction error comparable to when the latent space is only loosely regularized by $\mathcal{N}(\mathbf{0},I)$. The proposed approach does not require hyperparameter (i.e., the KLD weight $\beta$) tuning given a specific dataset as required in common VAE training practices. We evaluate the method using a medical dataset intended for 3D skull reconstruction and shape completion, and the results indicate promising generative capabilities of the VAE trained using the proposed method. Besides, through guided manipulation of the latent variables, we establish a connection between existing autoencoder (AE)-based approaches and generative approaches, such as VAE, for the shape completion problem. Codes and pre-trained weights are available at https://github.com/Jianningli/skullVAE
AutoPET Challenge: Combining nn-Unet with Swin UNETR Augmented by Maximum Intensity Projection Classifier
Sep 02, 2022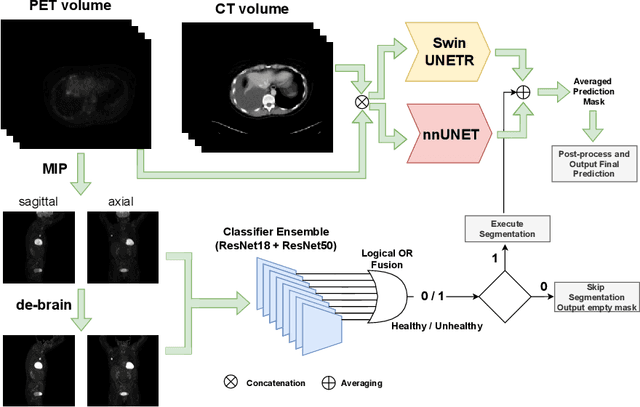
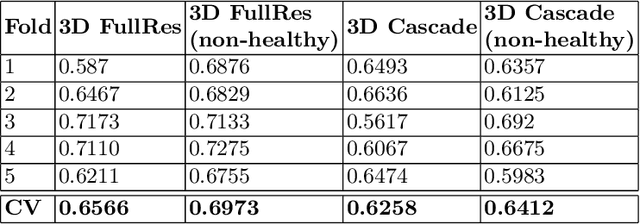
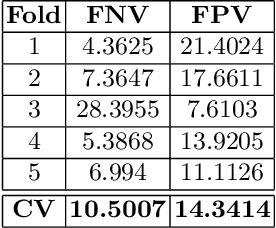
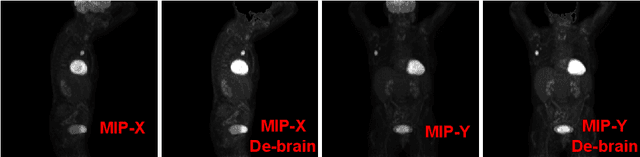
Abstract:Tumor volume and changes in tumor characteristics over time are important biomarkers for cancer therapy. In this context, FDG-PET/CT scans are routinely used for staging and re-staging of cancer, as the radiolabeled fluorodeoxyglucose is taken up in regions of high metabolism. Unfortunately, these regions with high metabolism are not specific to tumors and can also represent physiological uptake by normal functioning organs, inflammation, or infection, making detailed and reliable tumor segmentation in these scans a demanding task. This gap in research is addressed by the AutoPET challenge, which provides a public data set with FDG-PET/CT scans from 900 patients to encourage further improvement in this field. Our contribution to this challenge is an ensemble of two state-of-the-art segmentation models, the nn-Unet and the Swin UNETR, augmented by a maximum intensity projection classifier that acts like a gating mechanism. If it predicts the existence of lesions, both segmentations are combined by a late fusion approach. Our solution achieves a Dice score of 72.12\% on patients diagnosed with lung cancer, melanoma, and lymphoma in our cross-validation. Code: https://github.com/heiligerl/autopet_submission
Review of Disentanglement Approaches for Medical Applications -- Towards Solving the Gordian Knot of Generative Models in Healthcare
Mar 21, 2022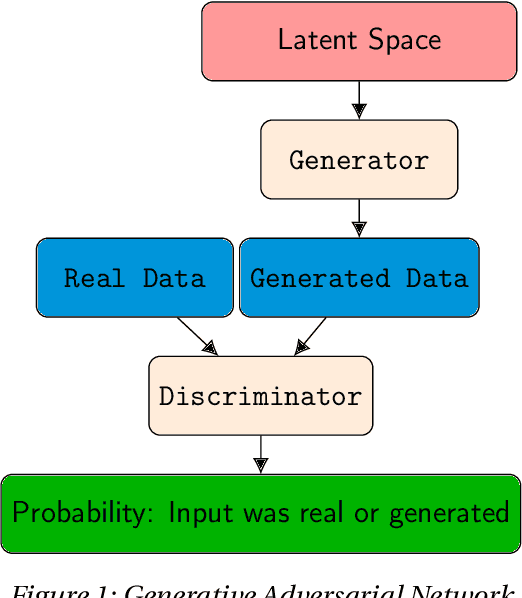

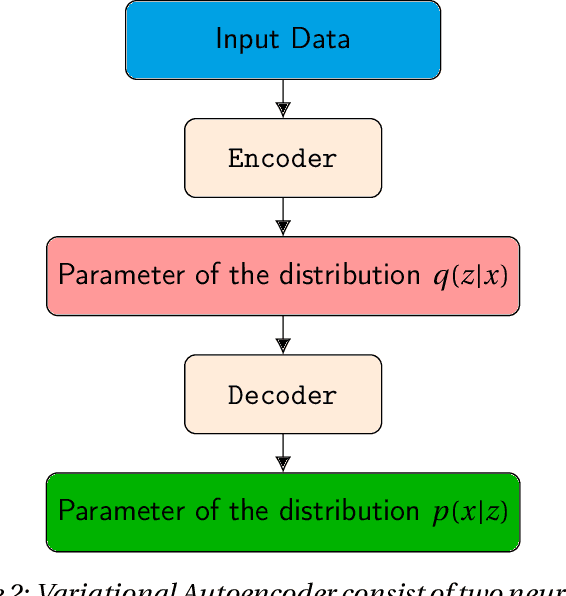
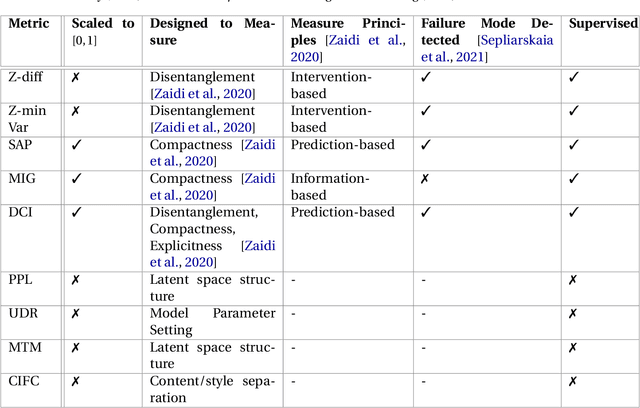
Abstract:Deep neural networks are commonly used for medical purposes such as image generation, segmentation, or classification. Besides this, they are often criticized as black boxes as their decision process is often not human interpretable. Encouraging the latent representation of a generative model to be disentangled offers new perspectives of control and interpretability. Understanding the data generation process could help to create artificial medical data sets without violating patient privacy, synthesizing different data modalities, or discovering data generating characteristics. These characteristics might unravel novel relationships that can be related to genetic traits or patient outcomes. In this paper, we give a comprehensive overview of popular generative models, like Generative Adversarial Networks (GANs), Variational Autoencoders (VAEs), and Flow-based Models. Furthermore, we summarize the different notions of disentanglement, review approaches to disentangle latent space representations and metrics to evaluate the degree of disentanglement. After introducing the theoretical frameworks, we give an overview of recent medical applications and discuss the impact and importance of disentanglement approaches for medical applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge