Giulia Baldini
SALT: Introducing a Framework for Hierarchical Segmentations in Medical Imaging using Softmax for Arbitrary Label Trees
Jul 11, 2024
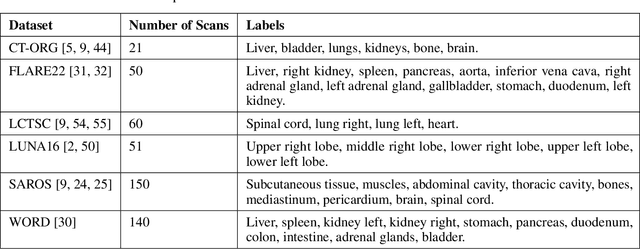
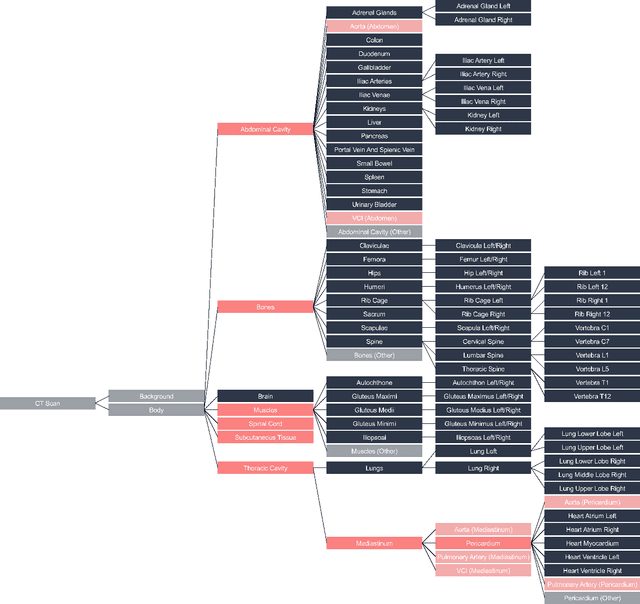
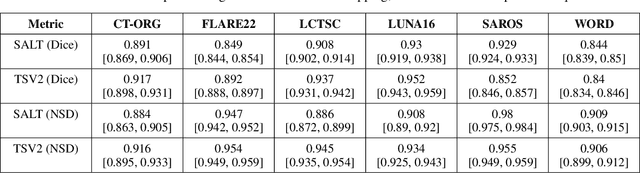
Abstract:Traditional segmentation networks approach anatomical structures as standalone elements, overlooking the intrinsic hierarchical connections among them. This study introduces Softmax for Arbitrary Label Trees (SALT), a novel approach designed to leverage the hierarchical relationships between labels, improving the efficiency and interpretability of the segmentations. This study introduces a novel segmentation technique for CT imaging, which leverages conditional probabilities to map the hierarchical structure of anatomical landmarks, such as the spine's division into lumbar, thoracic, and cervical regions and further into individual vertebrae. The model was developed using the SAROS dataset from The Cancer Imaging Archive (TCIA), comprising 900 body region segmentations from 883 patients. The dataset was further enhanced by generating additional segmentations with the TotalSegmentator, for a total of 113 labels. The model was trained on 600 scans, while validation and testing were conducted on 150 CT scans. Performance was assessed using the Dice score across various datasets, including SAROS, CT-ORG, FLARE22, LCTSC, LUNA16, and WORD. Among the evaluated datasets, SALT achieved its best results on the LUNA16 and SAROS datasets, with Dice scores of 0.93 and 0.929 respectively. The model demonstrated reliable accuracy across other datasets, scoring 0.891 on CT-ORG and 0.849 on FLARE22. The LCTSC dataset showed a score of 0.908 and the WORD dataset also showed good performance with a score of 0.844. SALT used the hierarchical structures inherent in the human body to achieve whole-body segmentations with an average of 35 seconds for 100 slices. This rapid processing underscores its potential for integration into clinical workflows, facilitating the automatic and efficient computation of full-body segmentations with each CT scan, thus enhancing diagnostic processes and patient care.
Comprehensive Study on German Language Models for Clinical and Biomedical Text Understanding
Apr 08, 2024



Abstract:Recent advances in natural language processing (NLP) can be largely attributed to the advent of pre-trained language models such as BERT and RoBERTa. While these models demonstrate remarkable performance on general datasets, they can struggle in specialized domains such as medicine, where unique domain-specific terminologies, domain-specific abbreviations, and varying document structures are common. This paper explores strategies for adapting these models to domain-specific requirements, primarily through continuous pre-training on domain-specific data. We pre-trained several German medical language models on 2.4B tokens derived from translated public English medical data and 3B tokens of German clinical data. The resulting models were evaluated on various German downstream tasks, including named entity recognition (NER), multi-label classification, and extractive question answering. Our results suggest that models augmented by clinical and translation-based pre-training typically outperform general domain models in medical contexts. We conclude that continuous pre-training has demonstrated the ability to match or even exceed the performance of clinical models trained from scratch. Furthermore, pre-training on clinical data or leveraging translated texts have proven to be reliable methods for domain adaptation in medical NLP tasks.
MRI Scan Synthesis Methods based on Clustering and Pix2Pix
Dec 08, 2023Abstract:We consider a missing data problem in the context of automatic segmentation methods for Magnetic Resonance Imaging (MRI) brain scans. Usually, automated MRI scan segmentation is based on multiple scans (e.g., T1-weighted, T2-weighted, T1CE, FLAIR). However, quite often a scan is blurry, missing or otherwise unusable. We investigate the question whether a missing scan can be synthesized. We exemplify that this is in principle possible by synthesizing a T2-weighted scan from a given T1-weighted scan. Our first aim is to compute a picture that resembles the missing scan closely, measured by average mean squared error (MSE). We develop/use several methods for this, including a random baseline approach, a clustering-based method and pixel-to-pixel translation method by (Pix2Pix) which is based on conditional GANs. The lowest MSE is achieved by our clustering-based method. Our second aim is to compare the methods with respect to the affect that using the synthesized scan has on the segmentation process. For this, we use a DeepMedic model trained with the four input scan modalities named above. We replace the T2-weighted scan by the synthesized picture and evaluate the segmentations with respect to the tumor identification, using Dice scores as numerical evaluation. The evaluation shows that the segmentation works well with synthesized scans (in particular, with Pix2Pix methods) in many cases.
CellViT: Vision Transformers for Precise Cell Segmentation and Classification
Jun 27, 2023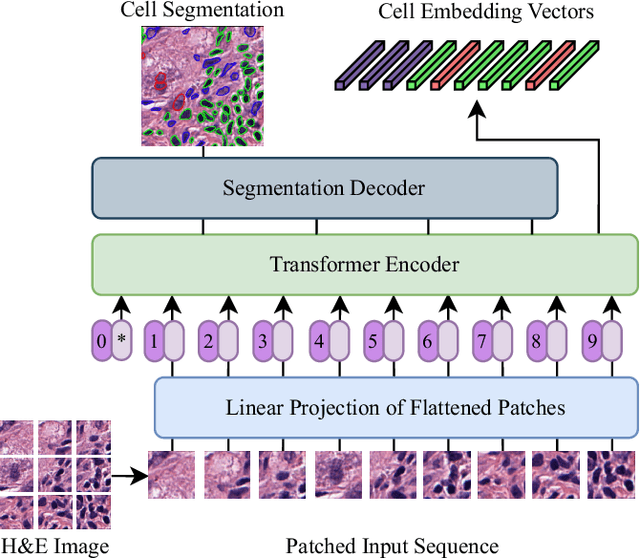
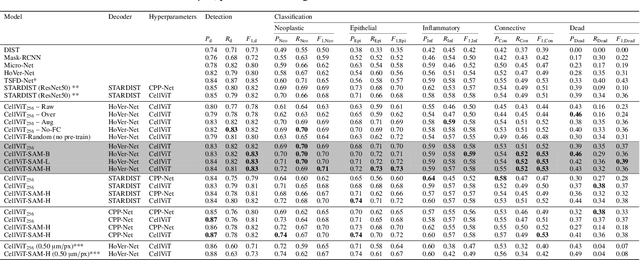
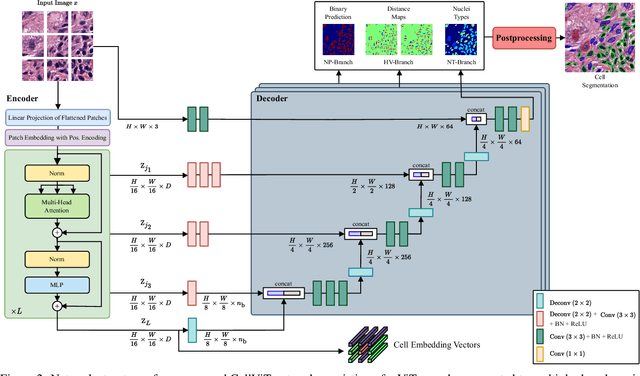

Abstract:Nuclei detection and segmentation in hematoxylin and eosin-stained (H&E) tissue images are important clinical tasks and crucial for a wide range of applications. However, it is a challenging task due to nuclei variances in staining and size, overlapping boundaries, and nuclei clustering. While convolutional neural networks have been extensively used for this task, we explore the potential of Transformer-based networks in this domain. Therefore, we introduce a new method for automated instance segmentation of cell nuclei in digitized tissue samples using a deep learning architecture based on Vision Transformer called CellViT. CellViT is trained and evaluated on the PanNuke dataset, which is one of the most challenging nuclei instance segmentation datasets, consisting of nearly 200,000 annotated Nuclei into 5 clinically important classes in 19 tissue types. We demonstrate the superiority of large-scale in-domain and out-of-domain pre-trained Vision Transformers by leveraging the recently published Segment Anything Model and a ViT-encoder pre-trained on 104 million histological image patches - achieving state-of-the-art nuclei detection and instance segmentation performance on the PanNuke dataset with a mean panoptic quality of 0.51 and an F1-detection score of 0.83. The code is publicly available at https://github.com/TIO-IKIM/CellViT
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge