Henning Schäfer
A Multimodal Pipeline for Clinical Data Extraction: Applying Vision-Language Models to Scans of Transfusion Reaction Reports
Apr 28, 2025Abstract:Despite the growing adoption of electronic health records, many processes still rely on paper documents, reflecting the heterogeneous real-world conditions in which healthcare is delivered. The manual transcription process is time-consuming and prone to errors when transferring paper-based data to digital formats. To streamline this workflow, this study presents an open-source pipeline that extracts and categorizes checkbox data from scanned documents. Demonstrated on transfusion reaction reports, the design supports adaptation to other checkbox-rich document types. The proposed method integrates checkbox detection, multilingual optical character recognition (OCR) and multilingual vision-language models (VLMs). The pipeline achieves high precision and recall compared against annually compiled gold-standards from 2017 to 2024. The result is a reduction in administrative workload and accurate regulatory reporting. The open-source availability of this pipeline encourages self-hosted parsing of checkbox forms.
WisPerMed at BioLaySumm: Adapting Autoregressive Large Language Models for Lay Summarization of Scientific Articles
May 20, 2024Abstract:This paper details the efforts of the WisPerMed team in the BioLaySumm2024 Shared Task on automatic lay summarization in the biomedical domain, aimed at making scientific publications accessible to non-specialists. Large language models (LLMs), specifically the BioMistral and Llama3 models, were fine-tuned and employed to create lay summaries from complex scientific texts. The summarization performance was enhanced through various approaches, including instruction tuning, few-shot learning, and prompt variations tailored to incorporate specific context information. The experiments demonstrated that fine-tuning generally led to the best performance across most evaluated metrics. Few-shot learning notably improved the models' ability to generate relevant and factually accurate texts, particularly when using a well-crafted prompt. Additionally, a Dynamic Expert Selection (DES) mechanism to optimize the selection of text outputs based on readability and factuality metrics was developed. Out of 54 participants, the WisPerMed team reached the 4th place, measured by readability, factuality, and relevance. Determined by the overall score, our approach improved upon the baseline by approx. 5.5 percentage points and was only approx 1.5 percentage points behind the first place.
WisPerMed at "Discharge Me!": Advancing Text Generation in Healthcare with Large Language Models, Dynamic Expert Selection, and Priming Techniques on MIMIC-IV
May 18, 2024


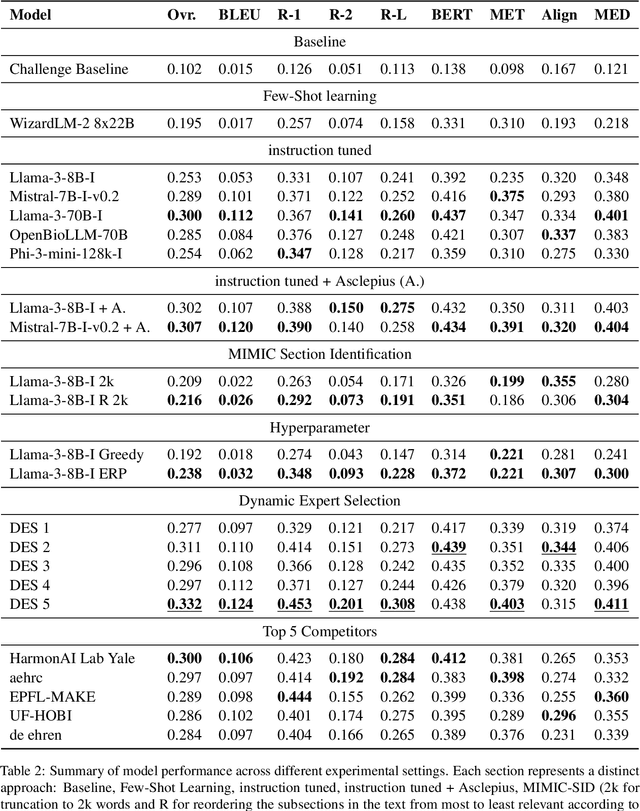
Abstract:This study aims to leverage state of the art language models to automate generating the "Brief Hospital Course" and "Discharge Instructions" sections of Discharge Summaries from the MIMIC-IV dataset, reducing clinicians' administrative workload. We investigate how automation can improve documentation accuracy, alleviate clinician burnout, and enhance operational efficacy in healthcare facilities. This research was conducted within our participation in the Shared Task Discharge Me! at BioNLP @ ACL 2024. Various strategies were employed, including few-shot learning, instruction tuning, and Dynamic Expert Selection (DES), to develop models capable of generating the required text sections. Notably, utilizing an additional clinical domain-specific dataset demonstrated substantial potential to enhance clinical language processing. The DES method, which optimizes the selection of text outputs from multiple predictions, proved to be especially effective. It achieved the highest overall score of 0.332 in the competition, surpassing single-model outputs. This finding suggests that advanced deep learning methods in combination with DES can effectively automate parts of electronic health record documentation. These advancements could enhance patient care by freeing clinician time for patient interactions. The integration of text selection strategies represents a promising avenue for further research.
ROCOv2: Radiology Objects in COntext Version 2, an Updated Multimodal Image Dataset
May 16, 2024Abstract:Automated medical image analysis systems often require large amounts of training data with high quality labels, which are difficult and time consuming to generate. This paper introduces Radiology Object in COntext version 2 (ROCOv2), a multimodal dataset consisting of radiological images and associated medical concepts and captions extracted from the PMC Open Access subset. It is an updated version of the ROCO dataset published in 2018, and adds 35,705 new images added to PMC since 2018. It further provides manually curated concepts for imaging modalities with additional anatomical and directional concepts for X-rays. The dataset consists of 79,789 images and has been used, with minor modifications, in the concept detection and caption prediction tasks of ImageCLEFmedical Caption 2023. The dataset is suitable for training image annotation models based on image-caption pairs, or for multi-label image classification using Unified Medical Language System (UMLS) concepts provided with each image. In addition, it can serve for pre-training of medical domain models, and evaluation of deep learning models for multi-task learning.
Comprehensive Study on German Language Models for Clinical and Biomedical Text Understanding
Apr 08, 2024



Abstract:Recent advances in natural language processing (NLP) can be largely attributed to the advent of pre-trained language models such as BERT and RoBERTa. While these models demonstrate remarkable performance on general datasets, they can struggle in specialized domains such as medicine, where unique domain-specific terminologies, domain-specific abbreviations, and varying document structures are common. This paper explores strategies for adapting these models to domain-specific requirements, primarily through continuous pre-training on domain-specific data. We pre-trained several German medical language models on 2.4B tokens derived from translated public English medical data and 3B tokens of German clinical data. The resulting models were evaluated on various German downstream tasks, including named entity recognition (NER), multi-label classification, and extractive question answering. Our results suggest that models augmented by clinical and translation-based pre-training typically outperform general domain models in medical contexts. We conclude that continuous pre-training has demonstrated the ability to match or even exceed the performance of clinical models trained from scratch. Furthermore, pre-training on clinical data or leveraging translated texts have proven to be reliable methods for domain adaptation in medical NLP tasks.
Domain Adaptation of Transformer-Based Models using Unlabeled Data for Relevance and Polarity Classification of German Customer Feedback
Dec 12, 2022Abstract:Understanding customer feedback is becoming a necessity for companies to identify problems and improve their products and services. Text classification and sentiment analysis can play a major role in analyzing this data by using a variety of machine and deep learning approaches. In this work, different transformer-based models are utilized to explore how efficient these models are when working with a German customer feedback dataset. In addition, these pre-trained models are further analyzed to determine if adapting them to a specific domain using unlabeled data can yield better results than off-the-shelf pre-trained models. To evaluate the models, two downstream tasks from the GermEval 2017 are considered. The experimental results show that transformer-based models can reach significant improvements compared to a fastText baseline and outperform the published scores and previous models. For the subtask Relevance Classification, the best models achieve a micro-averaged $F1$-Score of 96.1 % on the first test set and 95.9 % on the second one, and a score of 85.1 % and 85.3 % for the subtask Polarity Classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge