Johannes Stegmaier
Unconsciously Forget: Mitigating Memorization; Without Knowing What is being Memorized
Dec 12, 2025Abstract:Recent advances in generative models have demonstrated an exceptional ability to produce highly realistic images. However, previous studies show that generated images often resemble the training data, and this problem becomes more severe as the model size increases. Memorizing training data can lead to legal challenges, including copyright infringement, violations of portrait rights, and trademark violations. Existing approaches to mitigating memorization mainly focus on manipulating the denoising sampling process to steer image embeddings away from the memorized embedding space or employ unlearning methods that require training on datasets containing specific sets of memorized concepts. However, existing methods often incur substantial computational overhead during sampling, or focus narrowly on removing one or more groups of target concepts, imposing a significant limitation on their scalability. To understand and mitigate these problems, our work, UniForget, offers a new perspective on understanding the root cause of memorization. Our work demonstrates that specific parts of the model are responsible for copyrighted content generation. By applying model pruning, we can effectively suppress the probability of generating copyrighted content without targeting specific concepts while preserving the general generative capabilities of the model. Additionally, we show that our approach is both orthogonal and complementary to existing unlearning methods, thereby highlighting its potential to improve current unlearning and de-memorization techniques.
Diffusion-based Sinogram Interpolation for Limited Angle PET
Nov 12, 2025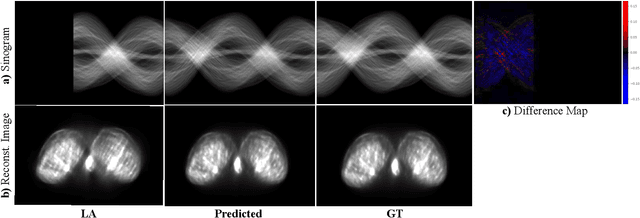
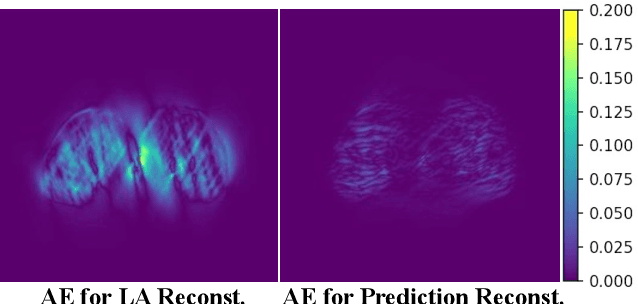

Abstract:Accurate PET imaging increasingly requires methods that support unconstrained detector layouts from walk-through designs to long-axial rings where gaps and open sides lead to severely undersampled sinograms. Instead of constraining the hardware to form complete cylinders, we propose treating the missing lines-of-responses as a learnable prior. Data-driven approaches, particularly generative models, offer a promising pathway to recover this missing information. In this work, we explore the use of conditional diffusion models to interpolate sparsely sampled sinograms, paving the way for novel, cost-efficient, and patient-friendly PET geometries in real clinical settings.
Towards Robust and Generalizable Gerchberg Saxton based Physics Inspired Neural Networks for Computer Generated Holography: A Sensitivity Analysis Framework
Apr 30, 2025Abstract:Computer-generated holography (CGH) enables applications in holographic augmented reality (AR), 3D displays, systems neuroscience, and optical trapping. The fundamental challenge in CGH is solving the inverse problem of phase retrieval from intensity measurements. Physics-inspired neural networks (PINNs), especially Gerchberg-Saxton-based PINNs (GS-PINNs), have advanced phase retrieval capabilities. However, their performance strongly depends on forward models (FMs) and their hyperparameters (FMHs), limiting generalization, complicating benchmarking, and hindering hardware optimization. We present a systematic sensitivity analysis framework based on Saltelli's extension of Sobol's method to quantify FMH impacts on GS-PINN performance. Our analysis demonstrates that SLM pixel-resolution is the primary factor affecting neural network sensitivity, followed by pixel-pitch, propagation distance, and wavelength. Free space propagation forward models demonstrate superior neural network performance compared to Fourier holography, providing enhanced parameterization and generalization. We introduce a composite evaluation metric combining performance consistency, generalization capability, and hyperparameter perturbation resilience, establishing a unified benchmarking standard across CGH configurations. Our research connects physics-inspired deep learning theory with practical CGH implementations through concrete guidelines for forward model selection, neural network architecture, and performance evaluation. Our contributions advance the development of robust, interpretable, and generalizable neural networks for diverse holographic applications, supporting evidence-based decisions in CGH research and implementation.
Comparative Analysis of Unsupervised and Supervised Autoencoders for Nuclei Classification in Clear Cell Renal Cell Carcinoma Images
Apr 04, 2025Abstract:This study explores the application of supervised and unsupervised autoencoders (AEs) to automate nuclei classification in clear cell renal cell carcinoma (ccRCC) images, a diagnostic task traditionally reliant on subjective visual grading by pathologists. We evaluate various AE architectures, including standard AEs, contractive AEs (CAEs), and discriminative AEs (DAEs), as well as a classifier-based discriminative AE (CDAE), optimized using the hyperparameter tuning tool Optuna. Bhattacharyya distance is selected from several metrics to assess class separability in the latent space, revealing challenges in distinguishing adjacent grades using unsupervised models. CDAE, integrating a supervised classifier branch, demonstrated superior performance in both latent space separation and classification accuracy. Given that CDAE-CNN achieved notable improvements in classification metrics, affirming the value of supervised learning for class-specific feature extraction, F1 score was incorporated into the tuning process to optimize classification performance. Results show significant improvements in identifying aggressive ccRCC grades by leveraging the classification capability of AE through latent clustering followed by fine-grained classification. Our model outperforms the current state of the art, CHR-Network, across all evaluated metrics. These findings suggest that integrating a classifier branch in AEs, combined with neural architecture search and contrastive learning, enhances grading automation in ccRCC pathology, particularly in detecting aggressive tumor grades, and may improve diagnostic accuracy.
Eye on the Target: Eye Tracking Meets Rodent Tracking
Mar 13, 2025Abstract:Analyzing animal behavior from video recordings is crucial for scientific research, yet manual annotation remains labor-intensive and prone to subjectivity. Efficient segmentation methods are needed to automate this process while maintaining high accuracy. In this work, we propose a novel pipeline that utilizes eye-tracking data from Aria glasses to generate prompt points, which are then used to produce segmentation masks via a fast zero-shot segmentation model. Additionally, we apply post-processing to refine the prompts, leading to improved segmentation quality. Through our approach, we demonstrate that combining eye-tracking-based annotation with smart prompt refinement can enhance segmentation accuracy, achieving an improvement of 70.6% from 38.8 to 66.2 in the Jaccard Index for segmentation results in the rats dataset.
CellStyle: Improved Zero-Shot Cell Segmentation via Style Transfer
Mar 11, 2025Abstract:Cell microscopy data are abundant; however, corresponding segmentation annotations remain scarce. Moreover, variations in cell types, imaging devices, and staining techniques introduce significant domain gaps between datasets. As a result, even large, pretrained segmentation models trained on diverse datasets (source datasets) struggle to generalize to unseen datasets (target datasets). To overcome this generalization problem, we propose CellStyle, which improves the segmentation quality of such models without requiring labels for the target dataset, thereby enabling zero-shot adaptation. CellStyle transfers the attributes of an unannotated target dataset, such as texture, color, and noise, to the annotated source dataset. This transfer is performed while preserving the cell shapes of the source images, ensuring that the existing source annotations can still be used while maintaining the visual characteristics of the target dataset. The styled synthetic images with the existing annotations enable the finetuning of a generalist segmentation model for application to the unannotated target data. We demonstrate that CellStyle significantly improves zero-shot cell segmentation performance across diverse datasets by finetuning multiple segmentation models on the style-transferred data. The code will be made publicly available.
A Pragmatic Note on Evaluating Generative Models with Fréchet Inception Distance for Retinal Image Synthesis
Feb 26, 2025Abstract:Fr\'echet Inception Distance (FID), computed with an ImageNet pretrained Inception-v3 network, is widely used as a state-of-the-art evaluation metric for generative models. It assumes that feature vectors from Inception-v3 follow a multivariate Gaussian distribution and calculates the 2-Wasserstein distance based on their means and covariances. While FID effectively measures how closely synthetic data match real data in many image synthesis tasks, the primary goal in biomedical generative models is often to enrich training datasets ideally with corresponding annotations. For this purpose, the gold standard for evaluating generative models is to incorporate synthetic data into downstream task training, such as classification and segmentation, to pragmatically assess its performance. In this paper, we examine cases from retinal imaging modalities, including color fundus photography and optical coherence tomography, where FID and its related metrics misalign with task-specific evaluation goals in classification and segmentation. We highlight the limitations of using various metrics, represented by FID and its variants, as evaluation criteria for these applications and address their potential caveats in broader biomedical imaging modalities and downstream tasks.
Unsupervised Learning for Feature Extraction and Temporal Alignment of 3D+t Point Clouds of Zebrafish Embryos
Feb 10, 2025Abstract:Zebrafish are widely used in biomedical research and developmental stages of their embryos often need to be synchronized for further analysis. We present an unsupervised approach to extract descriptive features from 3D+t point clouds of zebrafish embryos and subsequently use those features to temporally align corresponding developmental stages. An autoencoder architecture is proposed to learn a descriptive representation of the point clouds and we designed a deep regression network for their temporal alignment. We achieve a high alignment accuracy with an average mismatch of only 3.83 minutes over an experimental duration of 5.3 hours. As a fully-unsupervised approach, there is no manual labeling effort required and unlike manual analyses the method easily scales. Besides, the alignment without human annotation of the data also avoids any influence caused by subjective bias.
No Free Lunch in Annotation either: An objective evaluation of foundation models for streamlining annotation in animal tracking
Feb 06, 2025Abstract:We analyze the capabilities of foundation models addressing the tedious task of generating annotations for animal tracking. Annotating a large amount of data is vital and can be a make-or-break factor for the robustness of a tracking model. Robustness is particularly crucial in animal tracking, as accurate tracking over long time horizons is essential for capturing the behavior of animals. However, generating additional annotations using foundation models can be counterproductive, as the quality of the annotations is just as important. Poorly annotated data can introduce noise and inaccuracies, ultimately compromising the performance and accuracy of the trained model. Over-reliance on automated annotations without ensuring precision can lead to diminished results, making careful oversight and quality control essential in the annotation process. Ultimately, we demonstrate that a thoughtful combination of automated annotations and manually annotated data is a valuable strategy, yielding an IDF1 score of 80.8 against blind usage of SAM2 video with an IDF1 score of 65.6.
LogicAD: Explainable Anomaly Detection via VLM-based Text Feature Extraction
Jan 08, 2025

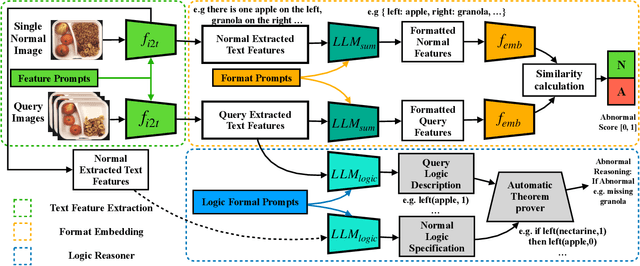
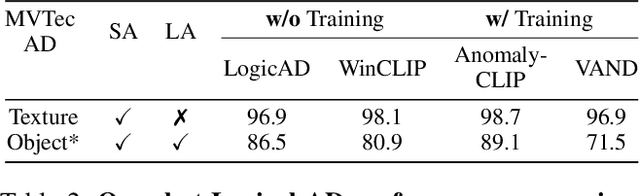
Abstract:Logical image understanding involves interpreting and reasoning about the relationships and consistency within an image's visual content. This capability is essential in applications such as industrial inspection, where logical anomaly detection is critical for maintaining high-quality standards and minimizing costly recalls. Previous research in anomaly detection (AD) has relied on prior knowledge for designing algorithms, which often requires extensive manual annotations, significant computing power, and large amounts of data for training. Autoregressive, multimodal Vision Language Models (AVLMs) offer a promising alternative due to their exceptional performance in visual reasoning across various domains. Despite this, their application to logical AD remains unexplored. In this work, we investigate using AVLMs for logical AD and demonstrate that they are well-suited to the task. Combining AVLMs with format embedding and a logic reasoner, we achieve SOTA performance on public benchmarks, MVTec LOCO AD, with an AUROC of 86.0% and F1-max of 83.7%, along with explanations of anomalies. This significantly outperforms the existing SOTA method by a large margin.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge