Jonathan Shapey
School of Biomedical Engineering and Imaging Sciences, Kings College London, Department of Neurosurgery, Kings College Hospital NHS Foundation Trust
X-RAFT: Cross-Modal Non-Rigid Registration of Blue and White Light Neurosurgical Hyperspectral Images
Jul 10, 2025Abstract:Integration of hyperspectral imaging into fluorescence-guided neurosurgery has the potential to improve surgical decision making by providing quantitative fluorescence measurements in real-time. Quantitative fluorescence requires paired spectral data in fluorescence (blue light) and reflectance (white light) mode. Blue and white image acquisition needs to be performed sequentially in a potentially dynamic surgical environment. A key component to the fluorescence quantification process is therefore the ability to find dense cross-modal image correspondences between two hyperspectral images taken under these drastically different lighting conditions. We address this challenge with the introduction of X-RAFT, a Recurrent All-Pairs Field Transforms (RAFT) optical flow model modified for cross-modal inputs. We propose using distinct image encoders for each modality pair, and fine-tune these in a self-supervised manner using flow-cycle-consistency on our neurosurgical hyperspectral data. We show an error reduction of 36.6% across our evaluation metrics when comparing to a naive baseline and 27.83% reduction compared to an existing cross-modal optical flow method (CrossRAFT). Our code and models will be made publicly available after the review process.
Systematic Review of Pituitary Gland and Pituitary Adenoma Automatic Segmentation Techniques in Magnetic Resonance Imaging
Jun 24, 2025Abstract:Purpose: Accurate segmentation of both the pituitary gland and adenomas from magnetic resonance imaging (MRI) is essential for diagnosis and treatment of pituitary adenomas. This systematic review evaluates automatic segmentation methods for improving the accuracy and efficiency of MRI-based segmentation of pituitary adenomas and the gland itself. Methods: We reviewed 34 studies that employed automatic and semi-automatic segmentation methods. We extracted and synthesized data on segmentation techniques and performance metrics (such as Dice overlap scores). Results: The majority of reviewed studies utilized deep learning approaches, with U-Net-based models being the most prevalent. Automatic methods yielded Dice scores of 0.19--89.00\% for pituitary gland and 4.60--96.41\% for adenoma segmentation. Semi-automatic methods reported 80.00--92.10\% for pituitary gland and 75.90--88.36\% for adenoma segmentation. Conclusion: Most studies did not report important metrics such as MR field strength, age and adenoma size. Automated segmentation techniques such as U-Net-based models show promise, especially for adenoma segmentation, but further improvements are needed to achieve consistently good performance in small structures like the normal pituitary gland. Continued innovation and larger, diverse datasets are likely critical to enhancing clinical applicability.
crossMoDA Challenge: Evolution of Cross-Modality Domain Adaptation Techniques for Vestibular Schwannoma and Cochlea Segmentation from 2021 to 2023
Jun 13, 2025Abstract:The cross-Modality Domain Adaptation (crossMoDA) challenge series, initiated in 2021 in conjunction with the International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI), focuses on unsupervised cross-modality segmentation, learning from contrast-enhanced T1 (ceT1) and transferring to T2 MRI. The task is an extreme example of domain shift chosen to serve as a meaningful and illustrative benchmark. From a clinical application perspective, it aims to automate Vestibular Schwannoma (VS) and cochlea segmentation on T2 scans for more cost-effective VS management. Over time, the challenge objectives have evolved to enhance its clinical relevance. The challenge evolved from using single-institutional data and basic segmentation in 2021 to incorporating multi-institutional data and Koos grading in 2022, and by 2023, it included heterogeneous routine data and sub-segmentation of intra- and extra-meatal tumour components. In this work, we report the findings of the 2022 and 2023 editions and perform a retrospective analysis of the challenge progression over the years. The observations from the successive challenge contributions indicate that the number of outliers decreases with an expanding dataset. This is notable since the diversity of scanning protocols of the datasets concurrently increased. The winning approach of the 2023 edition reduced the number of outliers on the 2021 and 2022 testing data, demonstrating how increased data heterogeneity can enhance segmentation performance even on homogeneous data. However, the cochlea Dice score declined in 2023, likely due to the added complexity from tumour sub-annotations affecting overall segmentation performance. While progress is still needed for clinically acceptable VS segmentation, the plateauing performance suggests that a more challenging cross-modal task may better serve future benchmarking.
A generalisable head MRI defacing pipeline: Evaluation on 2,566 meningioma scans
May 19, 2025Abstract:Reliable MRI defacing techniques to safeguard patient privacy while preserving brain anatomy are critical for research collaboration. Existing methods often struggle with incomplete defacing or degradation of brain tissue regions. We present a robust, generalisable defacing pipeline for high-resolution MRI that integrates atlas-based registration with brain masking. Our method was evaluated on 2,566 heterogeneous clinical scans for meningioma and achieved a 99.92 per cent success rate (2,564/2,566) upon visual inspection. Excellent anatomical preservation is demonstrated with a Dice similarity coefficient of 0.9975 plus or minus 0.0023 between brain masks automatically extracted from the original and defaced volumes. Source code is available at https://github.com/cai4cai/defacing_pipeline.
UltraFlwr -- An Efficient Federated Medical and Surgical Object Detection Framework
Mar 19, 2025Abstract:Object detection shows promise for medical and surgical applications such as cell counting and tool tracking. However, its faces multiple real-world edge deployment challenges including limited high-quality annotated data, data sharing restrictions, and computational constraints. In this work, we introduce UltraFlwr, a framework for federated medical and surgical object detection. By leveraging Federated Learning (FL), UltraFlwr enables decentralized model training across multiple sites without sharing raw data. To further enhance UltraFlwr's efficiency, we propose YOLO-PA, a set of novel Partial Aggregation (PA) strategies specifically designed for YOLO models in FL. YOLO-PA significantly reduces communication overhead by up to 83% per round while maintaining performance comparable to Full Aggregation (FA) strategies. Our extensive experiments on BCCD and m2cai16-tool-locations datasets demonstrate that YOLO-PA not only provides better client models compared to client-wise centralized training and FA strategies, but also facilitates efficient training and deployment across resource-constrained edge devices. Further, we also establish one of the first benchmarks in federated medical and surgical object detection. This paper advances the feasibility of training and deploying detection models on the edge, making federated object detection more practical for time-critical and resource-constrained medical and surgical applications. UltraFlwr is publicly available at https://github.com/KCL-BMEIS/UltraFlwr.
OOD-SEG: Out-Of-Distribution detection for image SEGmentation with sparse multi-class positive-only annotations
Nov 14, 2024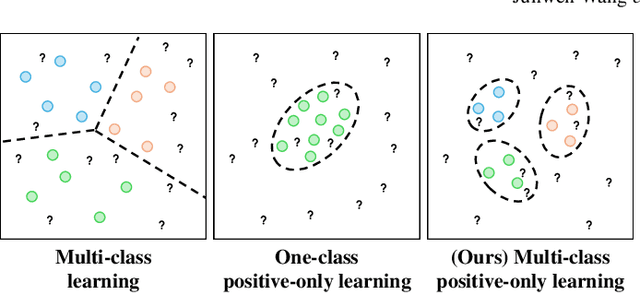


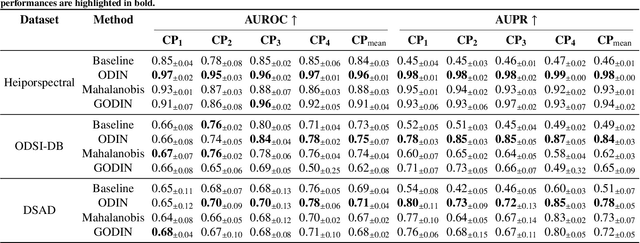
Abstract:Despite significant advancements, segmentation based on deep neural networks in medical and surgical imaging faces several challenges, two of which we aim to address in this work. First, acquiring complete pixel-level segmentation labels for medical images is time-consuming and requires domain expertise. Second, typical segmentation pipelines cannot detect out-of-distribution (OOD) pixels, leaving them prone to spurious outputs during deployment. In this work, we propose a novel segmentation approach exploiting OOD detection that learns only from sparsely annotated pixels from multiple positive-only classes. %but \emph{no background class} annotation. These multi-class positive annotations naturally fall within the in-distribution (ID) set. Unlabelled pixels may contain positive classes but also negative ones, including what is typically referred to as \emph{background} in standard segmentation formulations. Here, we forgo the need for background annotation and consider these together with any other unseen classes as part of the OOD set. Our framework can integrate, at a pixel-level, any OOD detection approaches designed for classification tasks. To address the lack of existing OOD datasets and established evaluation metric for medical image segmentation, we propose a cross-validation strategy that treats held-out labelled classes as OOD. Extensive experiments on both multi-class hyperspectral and RGB surgical imaging datasets demonstrate the robustness and generalisation capability of our proposed framework.
Scribble-Based Interactive Segmentation of Medical Hyperspectral Images
Aug 05, 2024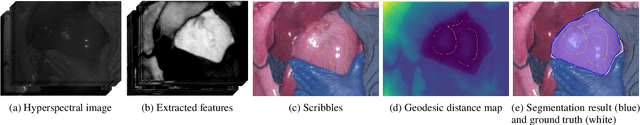

Abstract:Hyperspectral imaging (HSI) is an advanced medical imaging modality that captures optical data across a broad spectral range, providing novel insights into the biochemical composition of tissues. HSI may enable precise differentiation between various tissue types and pathologies, making it particularly valuable for tumour detection, tissue classification, and disease diagnosis. Deep learning-based segmentation methods have shown considerable advancements, offering automated and accurate results. However, these methods face challenges with HSI datasets due to limited annotated data and discrepancies from hardware and acquisition techniques~\cite{clancy2020surgical,studier2023heiporspectral}. Variability in clinical protocols also leads to different definitions of structure boundaries. Interactive segmentation methods, utilizing user knowledge and clinical insights, can overcome these issues and achieve precise segmentation results \cite{zhao2013overview}. This work introduces a scribble-based interactive segmentation framework for medical hyperspectral images. The proposed method utilizes deep learning for feature extraction and a geodesic distance map generated from user-provided scribbles to obtain the segmentation results. The experiment results show that utilising the geodesic distance maps based on deep learning-extracted features achieved better segmentation results than geodesic distance maps directly generated from hyperspectral images, reconstructed RGB images, or Euclidean distance maps.
A self-supervised and adversarial approach to hyperspectral demosaicking and RGB reconstruction in surgical imaging
Jul 27, 2024



Abstract:Hyperspectral imaging holds promises in surgical imaging by offering biological tissue differentiation capabilities with detailed information that is invisible to the naked eye. For intra-operative guidance, real-time spectral data capture and display is mandated. Snapshot mosaic hyperspectral cameras are currently seen as the most suitable technology given this requirement. However, snapshot mosaic imaging requires a demosaicking algorithm to fully restore the spatial and spectral details in the images. Modern demosaicking approaches typically rely on synthetic datasets to develop supervised learning methods, as it is practically impossible to simultaneously capture both snapshot and high-resolution spectral images of the exact same surgical scene. In this work, we present a self-supervised demosaicking and RGB reconstruction method that does not depend on paired high-resolution data as ground truth. We leverage unpaired standard high-resolution surgical microscopy images, which only provide RGB data but can be collected during routine surgeries. Adversarial learning complemented by self-supervised approaches are used to drive our hyperspectral-based RGB reconstruction into resembling surgical microscopy images and increasing the spatial resolution of our demosaicking. The spatial and spectral fidelity of the reconstructed hyperspectral images have been evaluated quantitatively. Moreover, a user study was conducted to evaluate the RGB visualisation generated from these spectral images. Both spatial detail and colour accuracy were assessed by neurosurgical experts. Our proposed self-supervised demosaicking method demonstrates improved results compared to existing methods, demonstrating its potential for seamless integration into intra-operative workflows.
Brain Tumor Segmentation (BraTS) Challenge 2024: Meningioma Radiotherapy Planning Automated Segmentation
May 28, 2024Abstract:The 2024 Brain Tumor Segmentation Meningioma Radiotherapy (BraTS-MEN-RT) challenge aims to advance automated segmentation algorithms using the largest known multi-institutional dataset of radiotherapy planning brain MRIs with expert-annotated target labels for patients with intact or post-operative meningioma that underwent either conventional external beam radiotherapy or stereotactic radiosurgery. Each case includes a defaced 3D post-contrast T1-weighted radiotherapy planning MRI in its native acquisition space, accompanied by a single-label "target volume" representing the gross tumor volume (GTV) and any at-risk post-operative site. Target volume annotations adhere to established radiotherapy planning protocols, ensuring consistency across cases and institutions. For pre-operative meningiomas, the target volume encompasses the entire GTV and associated nodular dural tail, while for post-operative cases, it includes at-risk resection cavity margins as determined by the treating institution. Case annotations were reviewed and approved by expert neuroradiologists and radiation oncologists. Participating teams will develop, containerize, and evaluate automated segmentation models using this comprehensive dataset. Model performance will be assessed using the lesion-wise Dice Similarity Coefficient and the 95% Hausdorff distance. The top-performing teams will be recognized at the Medical Image Computing and Computer Assisted Intervention Conference in October 2024. BraTS-MEN-RT is expected to significantly advance automated radiotherapy planning by enabling precise tumor segmentation and facilitating tailored treatment, ultimately improving patient outcomes.
A Clinical Guideline Driven Automated Linear Feature Extraction for Vestibular Schwannoma
Oct 30, 2023Abstract:Vestibular Schwannoma is a benign brain tumour that grows from one of the balance nerves. Patients may be treated by surgery, radiosurgery or with a conservative "wait-and-scan" strategy. Clinicians typically use manually extracted linear measurements to aid clinical decision making. This work aims to automate and improve this process by using deep learning based segmentation to extract relevant clinical features through computational algorithms. To the best of our knowledge, our study is the first to propose an automated approach to replicate local clinical guidelines. Our deep learning based segmentation provided Dice-scores of 0.8124 +- 0.2343 and 0.8969 +- 0.0521 for extrameatal and whole tumour regions respectively for T2 weighted MRI, whereas 0.8222 +- 0.2108 and 0.9049 +- 0.0646 were obtained for T1 weighted MRI. We propose a novel algorithm to choose and extract the most appropriate maximum linear measurement from the segmented regions based on the size of the extrameatal portion of the tumour. Using this tool, clinicians will be provided with a visual guide and related metrics relating to tumour progression that will function as a clinical decision aid. In this study, we utilize 187 scans obtained from 50 patients referred to a tertiary specialist neurosurgical service in the United Kingdom. The measurements extracted manually by an expert neuroradiologist indicated a significant correlation with the automated measurements (p < 0.0001).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge