Fakai Wang
Liver Tumor Screening and Diagnosis in CT with Pixel-Lesion-Patient Network
Jul 17, 2023Abstract:Liver tumor segmentation and classification are important tasks in computer aided diagnosis. We aim to address three problems: liver tumor screening and preliminary diagnosis in non-contrast computed tomography (CT), and differential diagnosis in dynamic contrast-enhanced CT. A novel framework named Pixel-Lesion-pAtient Network (PLAN) is proposed. It uses a mask transformer to jointly segment and classify each lesion with improved anchor queries and a foreground-enhanced sampling loss. It also has an image-wise classifier to effectively aggregate global information and predict patient-level diagnosis. A large-scale multi-phase dataset is collected containing 939 tumor patients and 810 normal subjects. 4010 tumor instances of eight types are extensively annotated. On the non-contrast tumor screening task, PLAN achieves 95% and 96% in patient-level sensitivity and specificity. On contrast-enhanced CT, our lesion-level detection precision, recall, and classification accuracy are 92%, 89%, and 86%, outperforming widely used CNN and transformers for lesion segmentation. We also conduct a reader study on a holdout set of 250 cases. PLAN is on par with a senior human radiologist, showing the clinical significance of our results.
A Cascaded Approach for ultraly High Performance Lesion Detection and False Positive Removal in Liver CT Scans
Jun 28, 2023Abstract:Liver cancer has high morbidity and mortality rates in the world. Multi-phase CT is a main medical imaging modality for detecting/identifying and diagnosing liver tumors. Automatically detecting and classifying liver lesions in CT images have the potential to improve the clinical workflow. This task remains challenging due to liver lesions' large variations in size, appearance, image contrast, and the complexities of tumor types or subtypes. In this work, we customize a multi-object labeling tool for multi-phase CT images, which is used to curate a large-scale dataset containing 1,631 patients with four-phase CT images, multi-organ masks, and multi-lesion (six major types of liver lesions confirmed by pathology) masks. We develop a two-stage liver lesion detection pipeline, where the high-sensitivity detecting algorithms in the first stage discover as many lesion proposals as possible, and the lesion-reclassification algorithms in the second stage remove as many false alarms as possible. The multi-sensitivity lesion detection algorithm maximizes the information utilization of the individual probability maps of segmentation, and the lesion-shuffle augmentation effectively explores the texture contrast between lesions and the liver. Independently tested on 331 patient cases, the proposed model achieves high sensitivity and specificity for malignancy classification in the multi-phase contrast-enhanced CT (99.2%, 97.1%, diagnosis setting) and in the noncontrast CT (97.3%, 95.7%, screening setting).
Towards a Single Unified Model for Effective Detection, Segmentation, and Diagnosis of Eight Major Cancers Using a Large Collection of CT Scans
Jan 28, 2023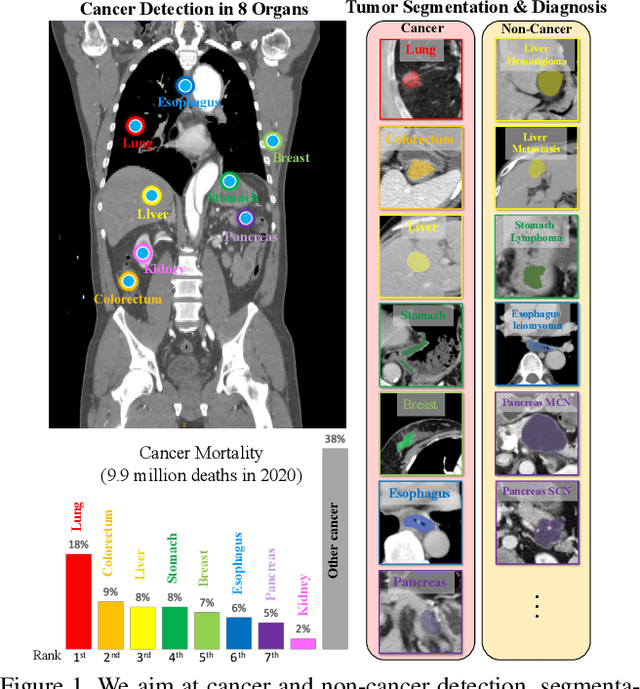
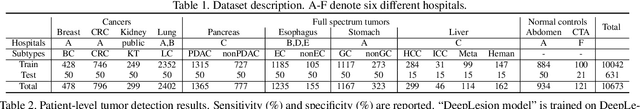
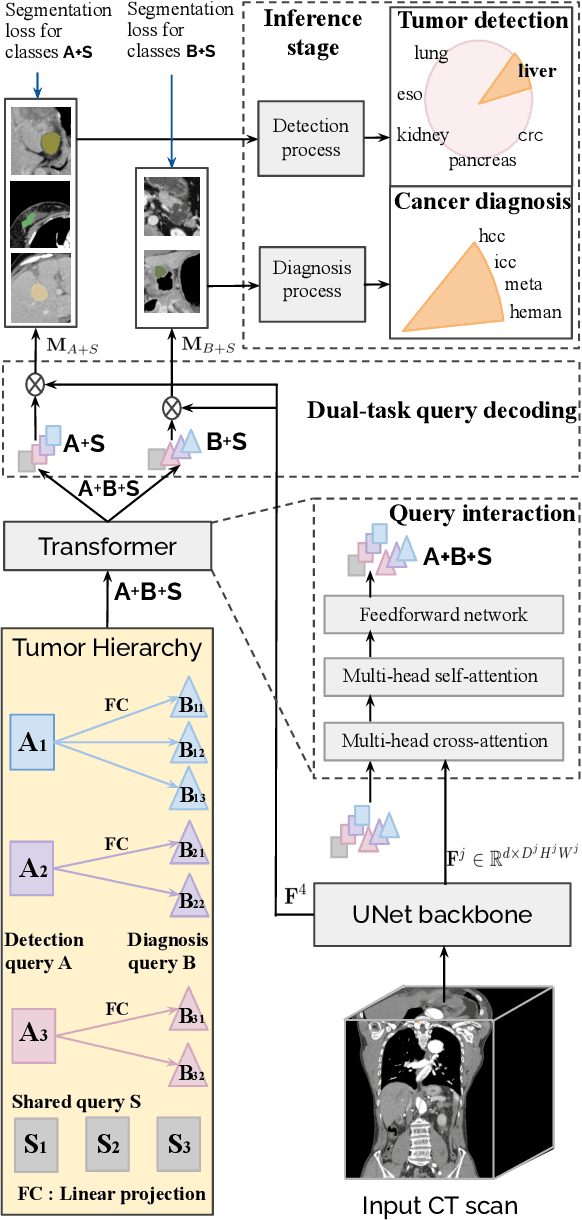
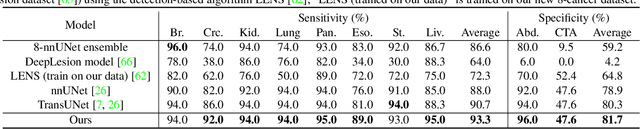
Abstract:Human readers or radiologists routinely perform full-body multi-organ multi-disease detection and diagnosis in clinical practice, while most medical AI systems are built to focus on single organs with a narrow list of a few diseases. This might severely limit AI's clinical adoption. A certain number of AI models need to be assembled non-trivially to match the diagnostic process of a human reading a CT scan. In this paper, we construct a Unified Tumor Transformer (UniT) model to detect (tumor existence and location) and diagnose (tumor characteristics) eight major cancer-prevalent organs in CT scans. UniT is a query-based Mask Transformer model with the output of multi-organ and multi-tumor semantic segmentation. We decouple the object queries into organ queries, detection queries and diagnosis queries, and further establish hierarchical relationships among the three groups. This clinically-inspired architecture effectively assists inter- and intra-organ representation learning of tumors and facilitates the resolution of these complex, anatomically related multi-organ cancer image reading tasks. UniT is trained end-to-end using a curated large-scale CT images of 10,042 patients including eight major types of cancers and occurring non-cancer tumors (all are pathology-confirmed with 3D tumor masks annotated by radiologists). On the test set of 631 patients, UniT has demonstrated strong performance under a set of clinically relevant evaluation metrics, substantially outperforming both multi-organ segmentation methods and an assembly of eight single-organ expert models in tumor detection, segmentation, and diagnosis. Such a unified multi-cancer image reading model (UniT) can significantly reduce the number of false positives produced by combined multi-system models. This moves one step closer towards a universal high-performance cancer screening tool.
Lumbar Bone Mineral Density Estimation from Chest X-ray Images: Anatomy-aware Attentive Multi-ROI Modeling
Jan 05, 2022


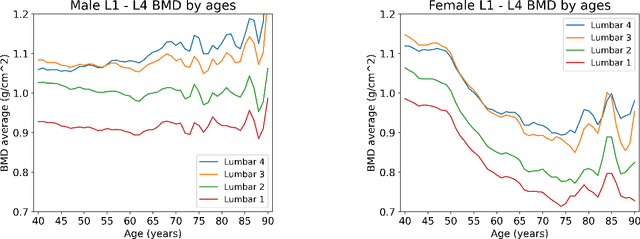
Abstract:Osteoporosis is a common chronic metabolic bone disease that is often under-diagnosed and under-treated due to the limited access to bone mineral density (BMD) examinations, e.g. via Dual-energy X-ray Absorptiometry (DXA). In this paper, we propose a method to predict BMD from Chest X-ray (CXR), one of the most commonly accessible and low-cost medical imaging examinations. Our method first automatically detects Regions of Interest (ROIs) of local and global bone structures from the CXR. Then a multi-ROI deep model with transformer encoder is developed to exploit both local and global information in the chest X-ray image for accurate BMD estimation. Our method is evaluated on 13719 CXR patient cases with their ground truth BMD scores measured by gold-standard DXA. The model predicted BMD has a strong correlation with the ground truth (Pearson correlation coefficient 0.889 on lumbar 1). When applied for osteoporosis screening, it achieves a high classification performance (AUC 0.963 on lumbar 1). As the first effort in the field using CXR scans to predict the BMD, the proposed algorithm holds strong potential in early osteoporosis screening and public health promotion.
Scalable Semi-supervised Landmark Localization for X-ray Images using Few-shot Deep Adaptive Graph
Apr 29, 2021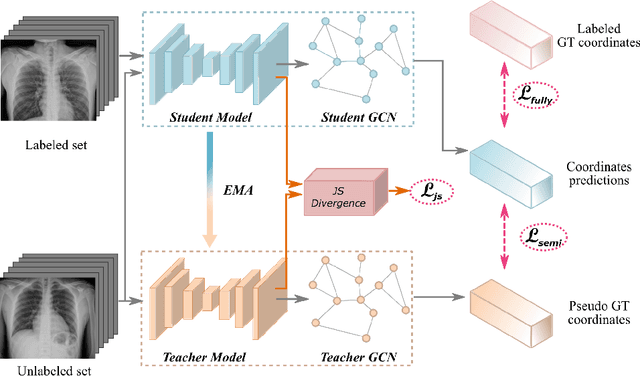
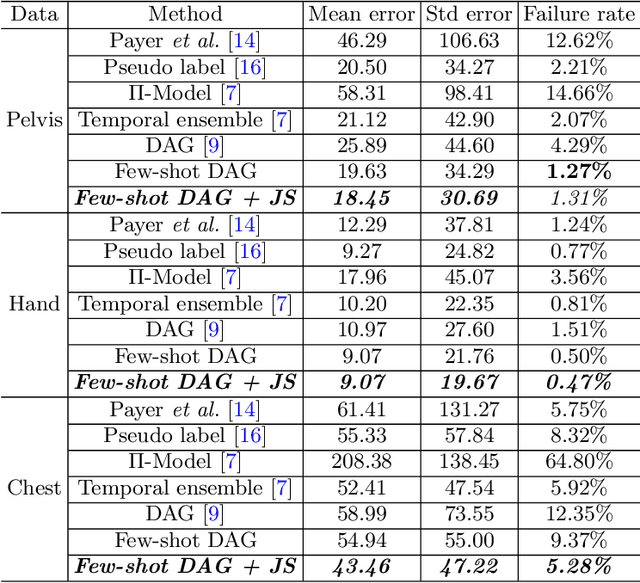

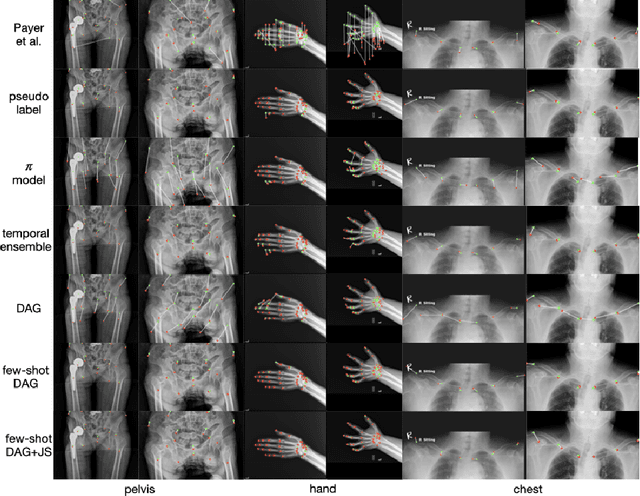
Abstract:Landmark localization plays an important role in medical image analysis. Learning based methods, including CNN and GCN, have demonstrated the state-of-the-art performance. However, most of these methods are fully-supervised and heavily rely on manual labeling of a large training dataset. In this paper, based on a fully-supervised graph-based method, DAG, we proposed a semi-supervised extension of it, termed few-shot DAG, \ie five-shot DAG. It first trains a DAG model on the labeled data and then fine-tunes the pre-trained model on the unlabeled data with a teacher-student SSL mechanism. In addition to the semi-supervised loss, we propose another loss using JS divergence to regulate the consistency of the intermediate feature maps. We extensively evaluated our method on pelvis, hand and chest landmark detection tasks. Our experiment results demonstrate consistent and significant improvements over previous methods.
Opportunistic Screening of Osteoporosis Using Plain Film Chest X-ray
Apr 05, 2021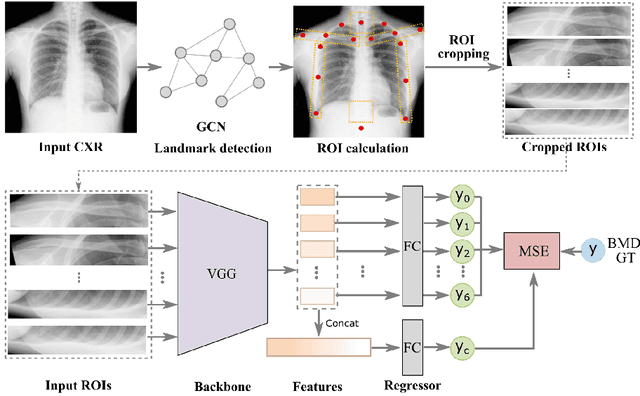



Abstract:Osteoporosis is a common chronic metabolic bone disease that is often under-diagnosed and under-treated due to the limited access to bone mineral density (BMD) examinations, Dual-energy X-ray Absorptiometry (DXA). In this paper, we propose a method to predict BMD from Chest X-ray (CXR), one of the most common, accessible, and low-cost medical image examinations. Our method first automatically detects Regions of Interest (ROIs) of local and global bone structures from the CXR. Then a multi-ROI model is developed to exploit both local and global information in the chest X-ray image for accurate BMD estimation. Our method is evaluated on 329 CXR cases with ground truth BMD measured by DXA. The model predicted BMD has a strong correlation with the gold standard DXA BMD (Pearson correlation coefficient 0.840). When applied for osteoporosis screening, it achieves a high classification performance (AUC 0.936). As the first effort in the field to use CXR scans to predict the spine BMD, the proposed algorithm holds strong potential in enabling early osteoporosis screening through routine chest X-rays and contributing to the enhancement of public health.
Semi-Supervised Learning for Bone Mineral Density Estimation in Hip X-ray Images
Mar 24, 2021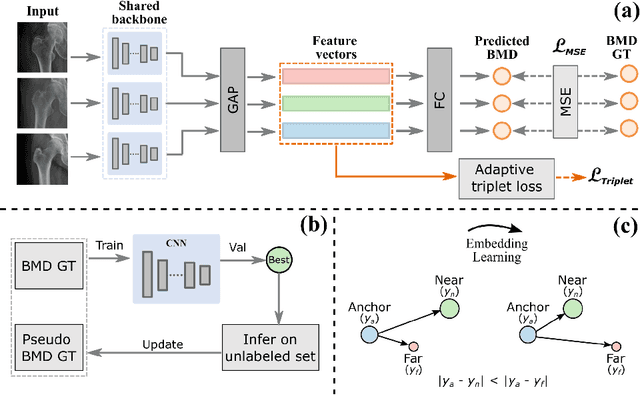

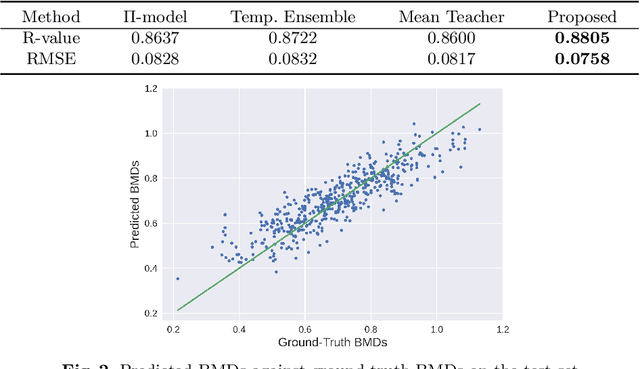
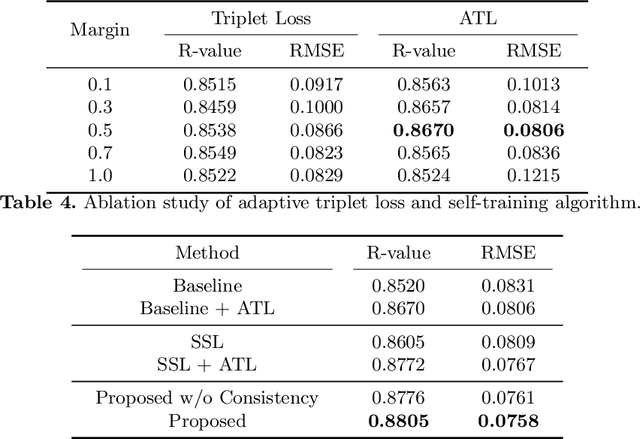
Abstract:Bone mineral density (BMD) is a clinically critical indicator of osteoporosis, usually measured by dual-energy X-ray absorptiometry (DEXA). Due to the limited accessibility of DEXA machines and examinations, osteoporosis is often under-diagnosed and under-treated, leading to increased fragility fracture risks. Thus it is highly desirable to obtain BMDs with alternative cost-effective and more accessible medical imaging examinations such as X-ray plain films. In this work, we formulate the BMD estimation from plain hip X-ray images as a regression problem. Specifically, we propose a new semi-supervised self-training algorithm to train the BMD regression model using images coupled with DEXA measured BMDs and unlabeled images with pseudo BMDs. Pseudo BMDs are generated and refined iteratively for unlabeled images during self-training. We also present a novel adaptive triplet loss to improve the model's regression accuracy. On an in-house dataset of 1,090 images (819 unique patients), our BMD estimation method achieves a high Pearson correlation coefficient of 0.8805 to ground-truth BMDs. It offers good feasibility to use the more accessible and cheaper X-ray imaging for opportunistic osteoporosis screening.
Automatic Vertebra Localization and Identification in CT by Spine Rectification and Anatomically-constrained Optimization
Dec 14, 2020



Abstract:Accurate vertebra localization and identification are required in many clinical applications of spine disorder diagnosis and surgery planning. However, significant challenges are posed in this task by highly varying pathologies (such as vertebral compression fracture, scoliosis, and vertebral fixation) and imaging conditions (such as limited field of view and metal streak artifacts). This paper proposes a robust and accurate method that effectively exploits the anatomical knowledge of the spine to facilitate vertebra localization and identification. A key point localization model is trained to produce activation maps of vertebra centers. They are then re-sampled along the spine centerline to produce spine-rectified activation maps, which are further aggregated into 1-D activation signals. Following this, an anatomically-constrained optimization module is introduced to jointly search for the optimal vertebra centers under a soft constraint that regulates the distance between vertebrae and a hard constraint on the consecutive vertebra indices. When being evaluated on a major public benchmark of 302 highly pathological CT images, the proposed method reports the state of the art identification (id.) rate of 97.4%, and outperforms the best competing method of 94.7% id. rate by reducing the relative id. error rate by half.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge