Danielle S. Bitterman
Proof of Time: A Benchmark for Evaluating Scientific Idea Judgments
Jan 12, 2026Abstract:Large language models are increasingly being used to assess and forecast research ideas, yet we lack scalable ways to evaluate the quality of models' judgments about these scientific ideas. Towards this goal, we introduce PoT, a semi-verifiable benchmarking framework that links scientific idea judgments to downstream signals that become observable later (e.g., citations and shifts in researchers' agendas). PoT freezes a pre-cutoff snapshot of evidence in an offline sandbox and asks models to forecast post-cutoff outcomes, enabling verifiable evaluation when ground truth arrives, scalable benchmarking without exhaustive expert annotation, and analysis of human-model misalignment against signals such as peer-review awards. In addition, PoT provides a controlled testbed for agent-based research judgments that evaluate scientific ideas, comparing tool-using agents to non-agent baselines under prompt ablations and budget scaling. Across 30,000+ instances spanning four benchmark domains, we find that, compared with non-agent baselines, higher interaction budgets generally improve agent performance, while the benefit of tool use is strongly task-dependent. By combining time-partitioned, future-verifiable targets with an offline sandbox for tool use, PoT supports scalable evaluation of agents on future-facing scientific idea judgment tasks.
Equitable Survival Prediction: A Fairness-Aware Survival Modeling (FASM) Approach
Oct 23, 2025Abstract:As machine learning models become increasingly integrated into healthcare, structural inequities and social biases embedded in clinical data can be perpetuated or even amplified by data-driven models. In survival analysis, censoring and time dynamics can further add complexity to fair model development. Additionally, algorithmic fairness approaches often overlook disparities in cross-group rankings, e.g., high-risk Black patients may be ranked below lower-risk White patients who do not experience the event of mortality. Such misranking can reinforce biological essentialism and undermine equitable care. We propose a Fairness-Aware Survival Modeling (FASM), designed to mitigate algorithmic bias regarding both intra-group and cross-group risk rankings over time. Using breast cancer prognosis as a representative case and applying FASM to SEER breast cancer data, we show that FASM substantially improves fairness while preserving discrimination performance comparable to fairness-unaware survival models. Time-stratified evaluations show that FASM maintains stable fairness over a 10-year horizon, with the greatest improvements observed during the mid-term of follow-up. Our approach enables the development of survival models that prioritize both accuracy and equity in clinical decision-making, advancing fairness as a core principle in clinical care.
Foundation Artificial Intelligence Models for Health Recognition Using Face Photographs (FAHR-Face)
Jun 17, 2025Abstract:Background: Facial appearance offers a noninvasive window into health. We built FAHR-Face, a foundation model trained on >40 million facial images and fine-tuned it for two distinct tasks: biological age estimation (FAHR-FaceAge) and survival risk prediction (FAHR-FaceSurvival). Methods: FAHR-FaceAge underwent a two-stage, age-balanced fine-tuning on 749,935 public images; FAHR-FaceSurvival was fine-tuned on 34,389 photos of cancer patients. Model robustness (cosmetic surgery, makeup, pose, lighting) and independence (saliency mapping) was tested extensively. Both models were clinically tested in two independent cancer patient datasets with survival analyzed by multivariable Cox models and adjusted for clinical prognostic factors. Findings: For age estimation, FAHR-FaceAge had the lowest mean absolute error of 5.1 years on public datasets, outperforming benchmark models and maintaining accuracy across the full human lifespan. In cancer patients, FAHR-FaceAge outperformed a prior facial age estimation model in survival prognostication. FAHR-FaceSurvival demonstrated robust prediction of mortality, and the highest-risk quartile had more than triple the mortality of the lowest (adjusted hazard ratio 3.22; P<0.001). These findings were validated in the independent cohort and both models showed generalizability across age, sex, race and cancer subgroups. The two algorithms provided distinct, complementary prognostic information; saliency mapping revealed each model relied on distinct facial regions. The combination of FAHR-FaceAge and FAHR-FaceSurvival improved prognostic accuracy. Interpretation: A single foundation model can generate inexpensive, scalable facial biomarkers that capture both biological ageing and disease-related mortality risk. The foundation model enabled effective training using relatively small clinical datasets.
When Models Reason in Your Language: Controlling Thinking Trace Language Comes at the Cost of Accuracy
May 28, 2025Abstract:Recent Large Reasoning Models (LRMs) with thinking traces have shown strong performance on English reasoning tasks. However, their ability to think in other languages is less studied. This capability is as important as answer accuracy for real world applications because users may find the reasoning trace useful for oversight only when it is expressed in their own language. We comprehensively evaluate two leading families of LRMs on our XReasoning benchmark and find that even the most advanced models often revert to English or produce fragmented reasoning in other languages, revealing a substantial gap in multilingual reasoning. Prompt based interventions that force models to reason in the users language improve readability and oversight but reduce answer accuracy, exposing an important trade off. We further show that targeted post training on just 100 examples mitigates this mismatch, though some accuracy loss remains. Our results highlight the limited multilingual reasoning capabilities of current LRMs and outline directions for future work. Code and data are available at https://github.com/Betswish/mCoT-XReasoning.
MedBrowseComp: Benchmarking Medical Deep Research and Computer Use
May 20, 2025Abstract:Large language models (LLMs) are increasingly envisioned as decision-support tools in clinical practice, yet safe clinical reasoning demands integrating heterogeneous knowledge bases -- trials, primary studies, regulatory documents, and cost data -- under strict accuracy constraints. Existing evaluations often rely on synthetic prompts, reduce the task to single-hop factoid queries, or conflate reasoning with open-ended generation, leaving their real-world utility unclear. To close this gap, we present MedBrowseComp, the first benchmark that systematically tests an agent's ability to reliably retrieve and synthesize multi-hop medical facts from live, domain-specific knowledge bases. MedBrowseComp contains more than 1,000 human-curated questions that mirror clinical scenarios where practitioners must reconcile fragmented or conflicting information to reach an up-to-date conclusion. Applying MedBrowseComp to frontier agentic systems reveals performance shortfalls as low as ten percent, exposing a critical gap between current LLM capabilities and the rigor demanded in clinical settings. MedBrowseComp therefore offers a clear testbed for reliable medical information seeking and sets concrete goals for future model and toolchain upgrades. You can visit our project page at: https://moreirap12.github.io/mbc-browse-app/
Sparse Autoencoder Features for Classifications and Transferability
Feb 17, 2025Abstract:Sparse Autoencoders (SAEs) provide potentials for uncovering structured, human-interpretable representations in Large Language Models (LLMs), making them a crucial tool for transparent and controllable AI systems. We systematically analyze SAE for interpretable feature extraction from LLMs in safety-critical classification tasks. Our framework evaluates (1) model-layer selection and scaling properties, (2) SAE architectural configurations, including width and pooling strategies, and (3) the effect of binarizing continuous SAE activations. SAE-derived features achieve macro F1 > 0.8, outperforming hidden-state and BoW baselines while demonstrating cross-model transfer from Gemma 2 2B to 9B-IT models. These features generalize in a zero-shot manner to cross-lingual toxicity detection and visual classification tasks. Our analysis highlights the significant impact of pooling strategies and binarization thresholds, showing that binarization offers an efficient alternative to traditional feature selection while maintaining or improving performance. These findings establish new best practices for SAE-based interpretability and enable scalable, transparent deployment of LLMs in real-world applications. Full repo: https://github.com/shan23chen/MOSAIC.
The use of large language models to enhance cancer clinical trial educational materials
Dec 02, 2024Abstract:Cancer clinical trials often face challenges in recruitment and engagement due to a lack of participant-facing informational and educational resources. This study investigated the potential of Large Language Models (LLMs), specifically GPT4, in generating patient-friendly educational content from clinical trial informed consent forms. Using data from ClinicalTrials.gov, we employed zero-shot learning for creating trial summaries and one-shot learning for developing multiple-choice questions, evaluating their effectiveness through patient surveys and crowdsourced annotation. Results showed that GPT4-generated summaries were both readable and comprehensive, and may improve patients' understanding and interest in clinical trials. The multiple-choice questions demonstrated high accuracy and agreement with crowdsourced annotators. For both resource types, hallucinations were identified that require ongoing human oversight. The findings demonstrate the potential of LLMs "out-of-the-box" to support the generation of clinical trial education materials with minimal trial-specific engineering, but implementation with a human-in-the-loop is still needed to avoid misinformation risks.
WorldMedQA-V: a multilingual, multimodal medical examination dataset for multimodal language models evaluation
Oct 16, 2024Abstract:Multimodal/vision language models (VLMs) are increasingly being deployed in healthcare settings worldwide, necessitating robust benchmarks to ensure their safety, efficacy, and fairness. Multiple-choice question and answer (QA) datasets derived from national medical examinations have long served as valuable evaluation tools, but existing datasets are largely text-only and available in a limited subset of languages and countries. To address these challenges, we present WorldMedQA-V, an updated multilingual, multimodal benchmarking dataset designed to evaluate VLMs in healthcare. WorldMedQA-V includes 568 labeled multiple-choice QAs paired with 568 medical images from four countries (Brazil, Israel, Japan, and Spain), covering original languages and validated English translations by native clinicians, respectively. Baseline performance for common open- and closed-source models are provided in the local language and English translations, and with and without images provided to the model. The WorldMedQA-V benchmark aims to better match AI systems to the diverse healthcare environments in which they are deployed, fostering more equitable, effective, and representative applications.
AIPatient: Simulating Patients with EHRs and LLM Powered Agentic Workflow
Sep 27, 2024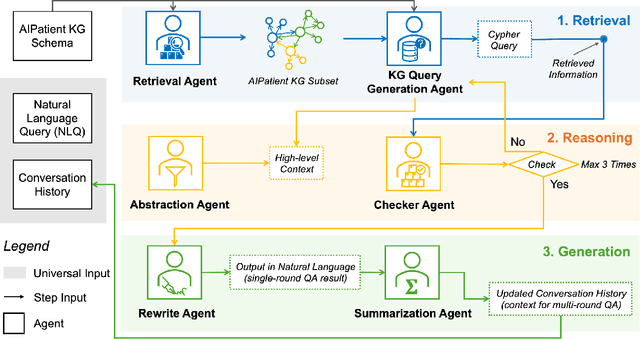
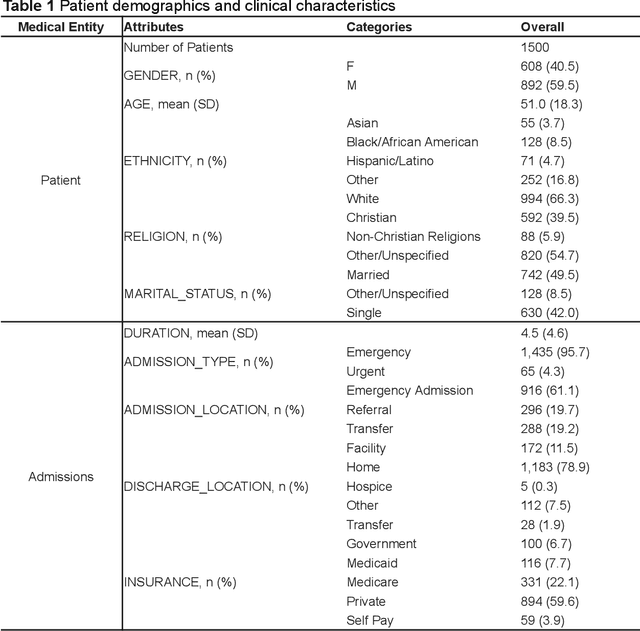
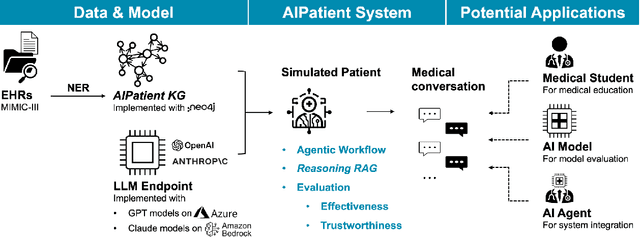
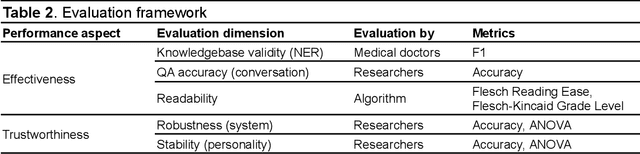
Abstract:Simulated patient systems play a crucial role in modern medical education and research, providing safe, integrative learning environments and enabling clinical decision-making simulations. Large Language Models (LLM) could advance simulated patient systems by replicating medical conditions and patient-doctor interactions with high fidelity and low cost. However, ensuring the effectiveness and trustworthiness of these systems remains a challenge, as they require a large, diverse, and precise patient knowledgebase, along with a robust and stable knowledge diffusion to users. Here, we developed AIPatient, an advanced simulated patient system with AIPatient Knowledge Graph (AIPatient KG) as the input and the Reasoning Retrieval-Augmented Generation (Reasoning RAG) agentic workflow as the generation backbone. AIPatient KG samples data from Electronic Health Records (EHRs) in the Medical Information Mart for Intensive Care (MIMIC)-III database, producing a clinically diverse and relevant cohort of 1,495 patients with high knowledgebase validity (F1 0.89). Reasoning RAG leverages six LLM powered agents spanning tasks including retrieval, KG query generation, abstraction, checker, rewrite, and summarization. This agentic framework reaches an overall accuracy of 94.15% in EHR-based medical Question Answering (QA), outperforming benchmarks that use either no agent or only partial agent integration. Our system also presents high readability (median Flesch Reading Ease 77.23; median Flesch Kincaid Grade 5.6), robustness (ANOVA F-value 0.6126, p<0.1), and stability (ANOVA F-value 0.782, p<0.1). The promising performance of the AIPatient system highlights its potential to support a wide range of applications, including medical education, model evaluation, and system integration.
Cross-Care: Assessing the Healthcare Implications of Pre-training Data on Language Model Bias
May 09, 2024



Abstract:Large language models (LLMs) are increasingly essential in processing natural languages, yet their application is frequently compromised by biases and inaccuracies originating in their training data. In this study, we introduce Cross-Care, the first benchmark framework dedicated to assessing biases and real world knowledge in LLMs, specifically focusing on the representation of disease prevalence across diverse demographic groups. We systematically evaluate how demographic biases embedded in pre-training corpora like $ThePile$ influence the outputs of LLMs. We expose and quantify discrepancies by juxtaposing these biases against actual disease prevalences in various U.S. demographic groups. Our results highlight substantial misalignment between LLM representation of disease prevalence and real disease prevalence rates across demographic subgroups, indicating a pronounced risk of bias propagation and a lack of real-world grounding for medical applications of LLMs. Furthermore, we observe that various alignment methods minimally resolve inconsistencies in the models' representation of disease prevalence across different languages. For further exploration and analysis, we make all data and a data visualization tool available at: www.crosscare.net.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge