Anna L. David
Deep Learning-Based Fetal Lung Segmentation from Diffusion-weighted MRI Images and Lung Maturity Evaluation for Fetal Growth Restriction
Jul 17, 2025Abstract:Fetal lung maturity is a critical indicator for predicting neonatal outcomes and the need for post-natal intervention, especially for pregnancies affected by fetal growth restriction. Intra-voxel incoherent motion analysis has shown promising results for non-invasive assessment of fetal lung development, but its reliance on manual segmentation is time-consuming, thus limiting its clinical applicability. In this work, we present an automated lung maturity evaluation pipeline for diffusion-weighted magnetic resonance images that consists of a deep learning-based fetal lung segmentation model and a model-fitting lung maturity assessment. A 3D nnU-Net model was trained on manually segmented images selected from the baseline frames of 4D diffusion-weighted MRI scans. The segmentation model demonstrated robust performance, yielding a mean Dice coefficient of 82.14%. Next, voxel-wise model fitting was performed based on both the nnU-Net-predicted and manual lung segmentations to quantify IVIM parameters reflecting tissue microstructure and perfusion. The results suggested no differences between the two. Our work shows that a fully automated pipeline is possible for supporting fetal lung maturity assessment and clinical decision-making.
Measuring proximity to standard planes during fetal brain ultrasound scanning
Apr 10, 2024



Abstract:This paper introduces a novel pipeline designed to bring ultrasound (US) plane pose estimation closer to clinical use for more effective navigation to the standard planes (SPs) in the fetal brain. We propose a semi-supervised segmentation model utilizing both labeled SPs and unlabeled 3D US volume slices. Our model enables reliable segmentation across a diverse set of fetal brain images. Furthermore, the model incorporates a classification mechanism to identify the fetal brain precisely. Our model not only filters out frames lacking the brain but also generates masks for those containing it, enhancing the relevance of plane pose regression in clinical settings. We focus on fetal brain navigation from 2D ultrasound (US) video analysis and combine this model with a US plane pose regression network to provide sensorless proximity detection to SPs and non-SPs planes; we emphasize the importance of proximity detection to SPs for guiding sonographers, offering a substantial advantage over traditional methods by allowing earlier and more precise adjustments during scanning. We demonstrate the practical applicability of our approach through validation on real fetal scan videos obtained from sonographers of varying expertise levels. Our findings demonstrate the potential of our approach to complement existing fetal US technologies and advance prenatal diagnostic practices.
Learning-Based Keypoint Registration for Fetoscopic Mosaicking
Jul 26, 2022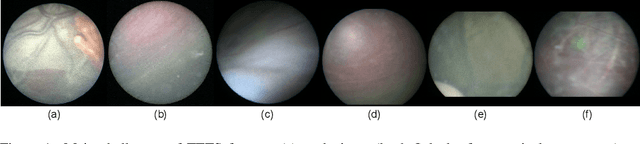
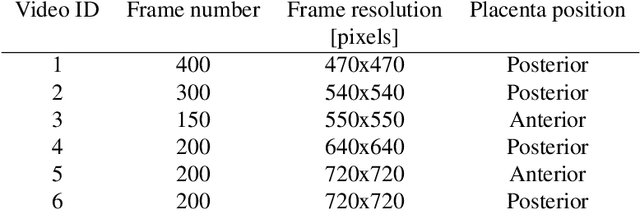
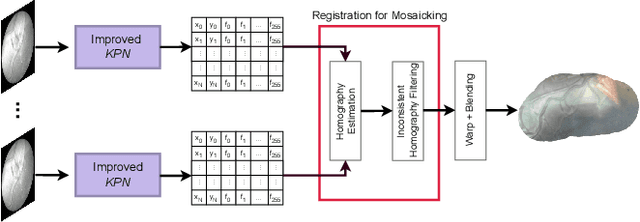

Abstract:In Twin-to-Twin Transfusion Syndrome (TTTS), abnormal vascular anastomoses in the monochorionic placenta can produce uneven blood flow between the two fetuses. In the current practice, TTTS is treated surgically by closing abnormal anastomoses using laser ablation. This surgery is minimally invasive and relies on fetoscopy. Limited field of view makes anastomosis identification a challenging task for the surgeon. To tackle this challenge, we propose a learning-based framework for in-vivo fetoscopy frame registration for field-of-view expansion. The novelties of this framework relies on a learning-based keypoint proposal network and an encoding strategy to filter (i) irrelevant keypoints based on fetoscopic image segmentation and (ii) inconsistent homographies. We validate of our framework on a dataset of 6 intraoperative sequences from 6 TTTS surgeries from 6 different women against the most recent state of the art algorithm, which relies on the segmentation of placenta vessels. The proposed framework achieves higher performance compared to the state of the art, paving the way for robust mosaicking to provide surgeons with context awareness during TTTS surgery.
BiometryNet: Landmark-based Fetal Biometry Estimation from Standard Ultrasound Planes
Jun 29, 2022



Abstract:Fetal growth assessment from ultrasound is based on a few biometric measurements that are performed manually and assessed relative to the expected gestational age. Reliable biometry estimation depends on the precise detection of landmarks in standard ultrasound planes. Manual annotation can be time-consuming and operator dependent task, and may results in high measurements variability. Existing methods for automatic fetal biometry rely on initial automatic fetal structure segmentation followed by geometric landmark detection. However, segmentation annotations are time-consuming and may be inaccurate, and landmark detection requires developing measurement-specific geometric methods. This paper describes BiometryNet, an end-to-end landmark regression framework for fetal biometry estimation that overcomes these limitations. It includes a novel Dynamic Orientation Determination (DOD) method for enforcing measurement-specific orientation consistency during network training. DOD reduces variabilities in network training, increases landmark localization accuracy, thus yields accurate and robust biometric measurements. To validate our method, we assembled a dataset of 3,398 ultrasound images from 1,829 subjects acquired in three clinical sites with seven different ultrasound devices. Comparison and cross-validation of three different biometric measurements on two independent datasets shows that BiometryNet is robust and yields accurate measurements whose errors are lower than the clinically permissible errors, outperforming other existing automated biometry estimation methods. Code is available at https://github.com/netanellavisdris/fetalbiometry.
A Dempster-Shafer approach to trustworthy AI with application to fetal brain MRI segmentation
Apr 05, 2022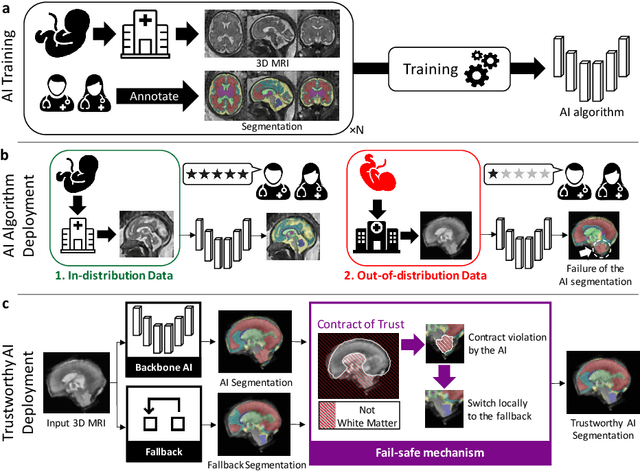
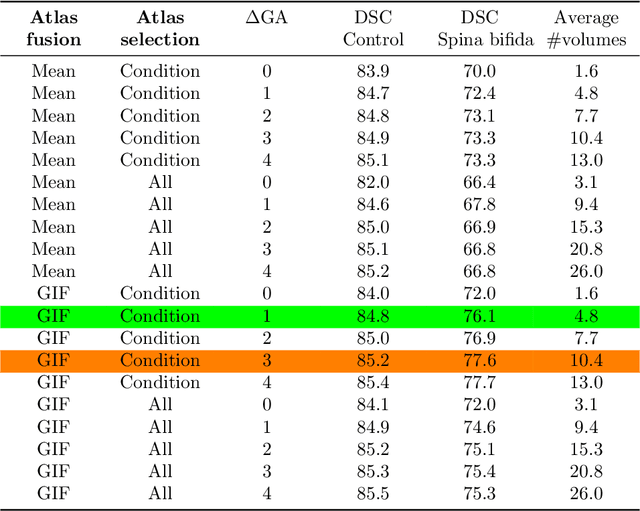
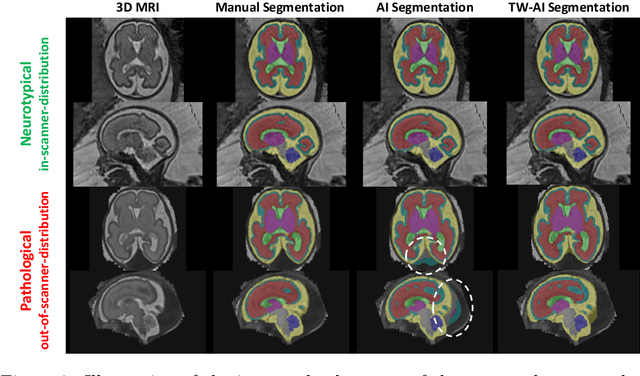
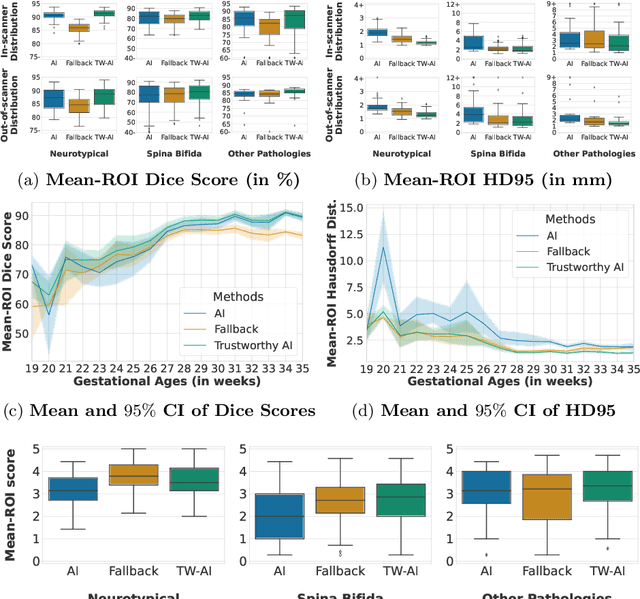
Abstract:Deep learning models for medical image segmentation can fail unexpectedly and spectacularly for pathological cases and for images acquired at different centers than those used for training, with labeling errors that violate expert knowledge about the anatomy and the intensity distribution of the regions to be segmented. Such errors undermine the trustworthiness of deep learning models developed for medical image segmentation. Mechanisms with a fallback method for detecting and correcting such failures are essential for safely translating this technology into clinics and are likely to be a requirement of future regulations on artificial intelligence (AI). Here, we propose a principled trustworthy AI theoretical framework and a practical system that can augment any backbone AI system using a fallback method and a fail-safe mechanism based on Dempster-Shafer theory. Our approach relies on an actionable definition of trustworthy AI. Our method automatically discards the voxel-level labeling predicted by the backbone AI that are likely to violate expert knowledge and relies on a fallback atlas-based segmentation method for those voxels. We demonstrate the effectiveness of the proposed trustworthy AI approach on the largest reported annotated dataset of fetal T2w MRI consisting of 540 manually annotated fetal brain 3D MRIs with neurotypical or abnormal brain development and acquired from 13 sources of data across 6 countries. We show that our trustworthy AI method improves the robustness of a state-of-the-art backbone AI for fetal brain MRI segmentation on MRIs acquired across various centers and for fetuses with various brain abnormalities.
Distributionally Robust Segmentation of Abnormal Fetal Brain 3D MRI
Aug 09, 2021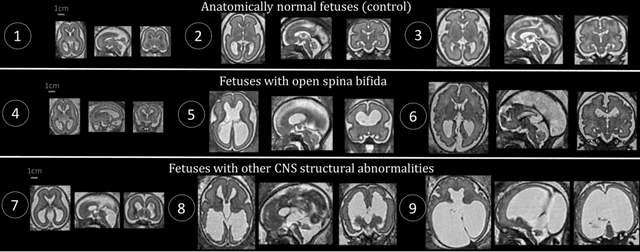
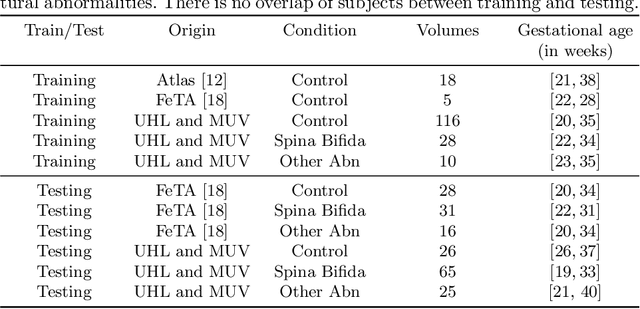
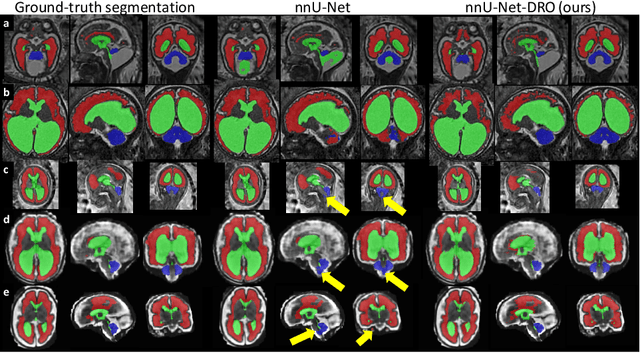
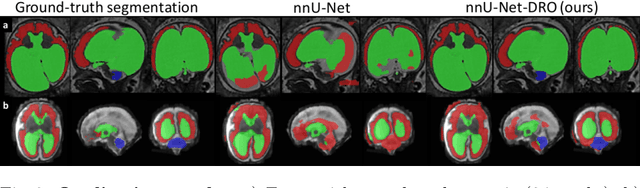
Abstract:The performance of deep neural networks typically increases with the number of training images. However, not all images have the same importance towards improved performance and robustness. In fetal brain MRI, abnormalities exacerbate the variability of the developing brain anatomy compared to non-pathological cases. A small number of abnormal cases, as is typically available in clinical datasets used for training, are unlikely to fairly represent the rich variability of abnormal developing brains. This leads machine learning systems trained by maximizing the average performance to be biased toward non-pathological cases. This problem was recently referred to as hidden stratification. To be suited for clinical use, automatic segmentation methods need to reliably achieve high-quality segmentation outcomes also for pathological cases. In this paper, we show that the state-of-the-art deep learning pipeline nnU-Net has difficulties to generalize to unseen abnormal cases. To mitigate this problem, we propose to train a deep neural network to minimize a percentile of the distribution of per-volume loss over the dataset. We show that this can be achieved by using Distributionally Robust Optimization (DRO). DRO automatically reweights the training samples with lower performance, encouraging nnU-Net to perform more consistently on all cases. We validated our approach using a dataset of 368 fetal brain T2w MRIs, including 124 MRIs of open spina bifida cases and 51 MRIs of cases with other severe abnormalities of brain development.
AutoFB: Automating Fetal Biometry Estimation from Standard Ultrasound Planes
Jul 12, 2021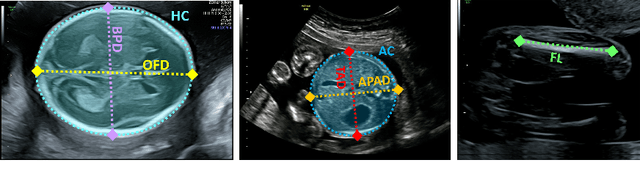

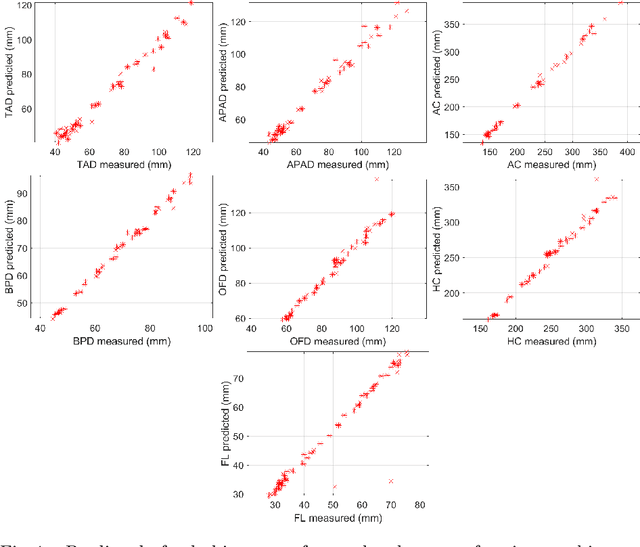
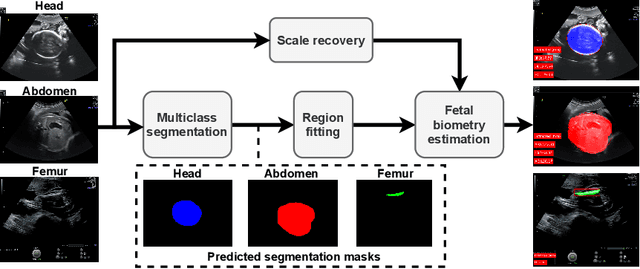
Abstract:During pregnancy, ultrasound examination in the second trimester can assess fetal size according to standardized charts. To achieve a reproducible and accurate measurement, a sonographer needs to identify three standard 2D planes of the fetal anatomy (head, abdomen, femur) and manually mark the key anatomical landmarks on the image for accurate biometry and fetal weight estimation. This can be a time-consuming operator-dependent task, especially for a trainee sonographer. Computer-assisted techniques can help in automating the fetal biometry computation process. In this paper, we present a unified automated framework for estimating all measurements needed for the fetal weight assessment. The proposed framework semantically segments the key fetal anatomies using state-of-the-art segmentation models, followed by region fitting and scale recovery for the biometry estimation. We present an ablation study of segmentation algorithms to show their robustness through 4-fold cross-validation on a dataset of 349 ultrasound standard plane images from 42 pregnancies. Moreover, we show that the network with the best segmentation performance tends to be more accurate for biometry estimation. Furthermore, we demonstrate that the error between clinically measured and predicted fetal biometry is lower than the permissible error during routine clinical measurements.
Label-set Loss Functions for Partial Supervision: Application to Fetal Brain 3D MRI Parcellation
Jul 09, 2021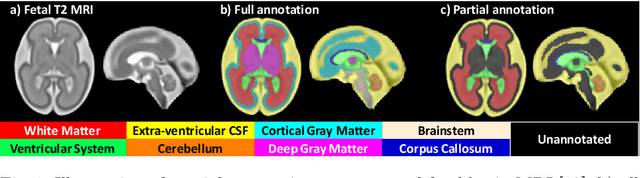
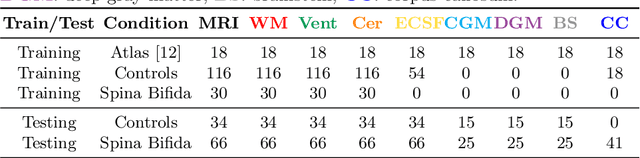
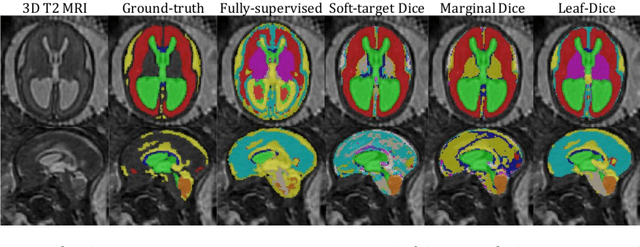
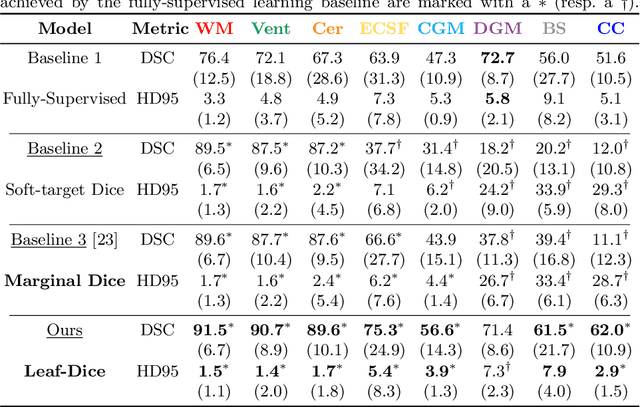
Abstract:Deep neural networks have increased the accuracy of automatic segmentation, however, their accuracy depends on the availability of a large number of fully segmented images. Methods to train deep neural networks using images for which some, but not all, regions of interest are segmented are necessary to make better use of partially annotated datasets. In this paper, we propose the first axiomatic definition of label-set loss functions that are the loss functions that can handle partially segmented images. We prove that there is one and only one method to convert a classical loss function for fully segmented images into a proper label-set loss function. Our theory also allows us to define the leaf-Dice loss, a label-set generalization of the Dice loss particularly suited for partial supervision with only missing labels. Using the leaf-Dice loss, we set a new state of the art in partially supervised learning for fetal brain 3D MRI segmentation. We achieve a deep neural network able to segment white matter, ventricles, cerebellum, extra-ventricular CSF, cortical gray matter, deep gray matter, brainstem, and corpus callosum based on fetal brain 3D MRI of anatomically normal fetuses or with open spina bifida. Our implementation of the proposed label-set loss functions is available at https://github.com/LucasFidon/label-set-loss-functions
FetReg: Placental Vessel Segmentation and Registration in Fetoscopy Challenge Dataset
Jun 16, 2021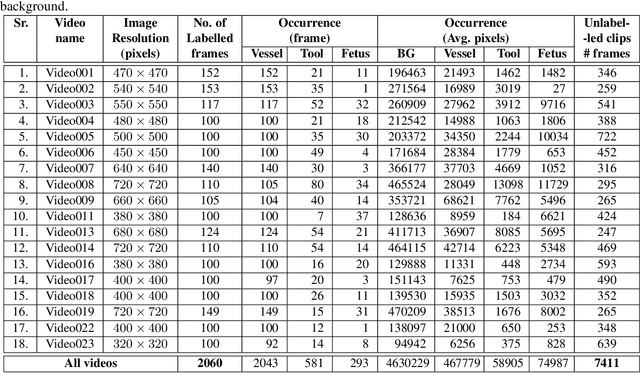
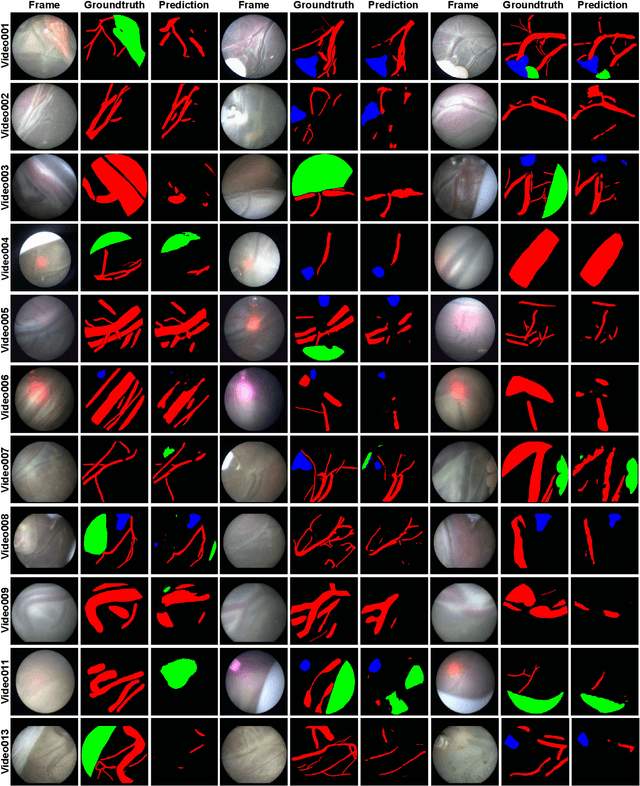

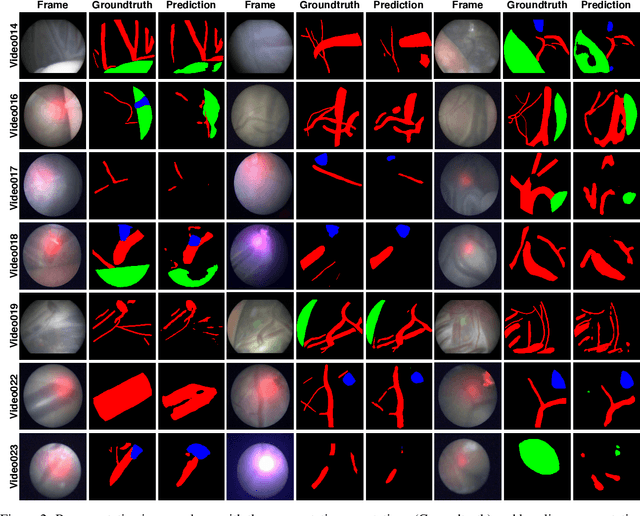
Abstract:Fetoscopy laser photocoagulation is a widely used procedure for the treatment of Twin-to-Twin Transfusion Syndrome (TTTS), that occur in mono-chorionic multiple pregnancies due to placental vascular anastomoses. This procedure is particularly challenging due to limited field of view, poor manoeuvrability of the fetoscope, poor visibility due to fluid turbidity, variability in light source, and unusual position of the placenta. This may lead to increased procedural time and incomplete ablation, resulting in persistent TTTS. Computer-assisted intervention may help overcome these challenges by expanding the fetoscopic field of view through video mosaicking and providing better visualization of the vessel network. However, the research and development in this domain remain limited due to unavailability of high-quality data to encode the intra- and inter-procedure variability. Through the \textit{Fetoscopic Placental Vessel Segmentation and Registration (FetReg)} challenge, we present a large-scale multi-centre dataset for the development of generalized and robust semantic segmentation and video mosaicking algorithms for the fetal environment with a focus on creating drift-free mosaics from long duration fetoscopy videos. In this paper, we provide an overview of the FetReg dataset, challenge tasks, evaluation metrics and baseline methods for both segmentation and registration. Baseline methods results on the FetReg dataset shows that our dataset poses interesting challenges, offering large opportunity for the creation of novel methods and models through a community effort initiative guided by the FetReg challenge.
Deep Placental Vessel Segmentation for Fetoscopic Mosaicking
Jul 08, 2020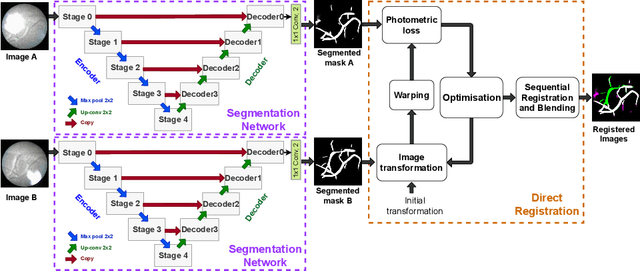
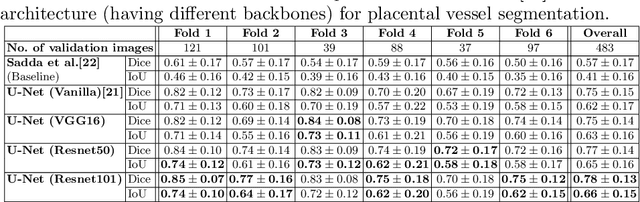
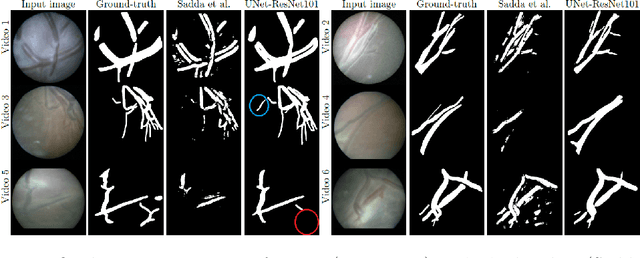
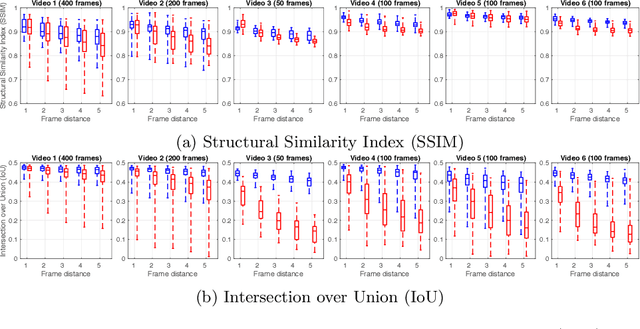
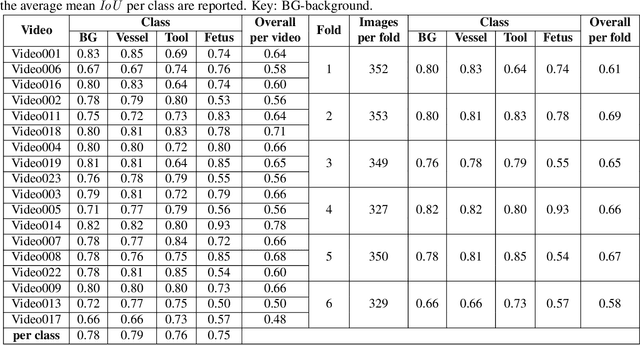
Abstract:During fetoscopic laser photocoagulation, a treatment for twin-to-twin transfusion syndrome (TTTS), the clinician first identifies abnormal placental vascular connections and laser ablates them to regulate blood flow in both fetuses. The procedure is challenging due to the mobility of the environment, poor visibility in amniotic fluid, occasional bleeding, and limitations in the fetoscopic field-of-view and image quality. Ideally, anastomotic placental vessels would be automatically identified, segmented and registered to create expanded vessel maps to guide laser ablation, however, such methods have yet to be clinically adopted. We propose a solution utilising the U-Net architecture for performing placental vessel segmentation in fetoscopic videos. The obtained vessel probability maps provide sufficient cues for mosaicking alignment by registering consecutive vessel maps using the direct intensity-based technique. Experiments on 6 different in vivo fetoscopic videos demonstrate that the vessel intensity-based registration outperformed image intensity-based registration approaches showing better robustness in qualitative and quantitative comparison. We additionally reduce drift accumulation to negligible even for sequences with up to 400 frames and we incorporate a scheme for quantifying drift error in the absence of the ground-truth. Our paper provides a benchmark for fetoscopy placental vessel segmentation and registration by contributing the first in vivo vessel segmentation and fetoscopic videos dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge