Philippe Demaerel
Partial supervision for the FeTA challenge 2021
Nov 03, 2021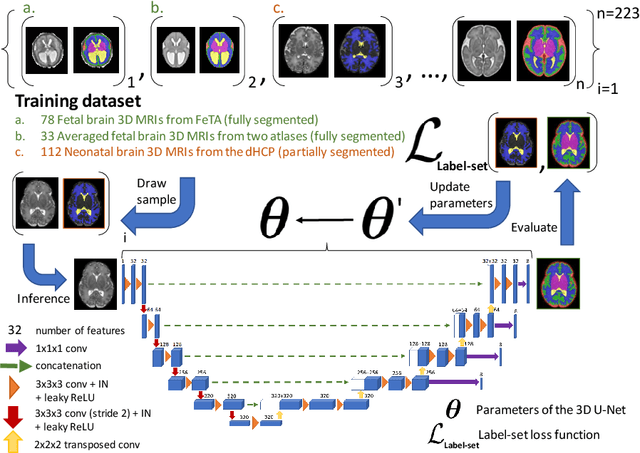
Abstract:This paper describes our method for our participation in the FeTA challenge2021 (team name: TRABIT). The performance of convolutional neural networks for medical image segmentation is thought to correlate positively with the number of training data. The FeTA challenge does not restrict participants to using only the provided training data but also allows for using other publicly available sources. Yet, open access fetal brain data remains limited. An advantageous strategy could thus be to expand the training data to cover broader perinatal brain imaging sources. Perinatal brain MRIs, other than the FeTA challenge data, that are currently publicly available, span normal and pathological fetal atlases as well as neonatal scans. However, perinatal brain MRIs segmented in different datasets typically come with different annotation protocols. This makes it challenging to combine those datasets to train a deep neural network. We recently proposed a family of loss functions, the label-set loss functions, for partially supervised learning. Label-set loss functions allow to train deep neural networks with partially segmented images, i.e. segmentations in which some classes may be grouped into super-classes. We propose to use label-set loss functions to improve the segmentation performance of a state-of-the-art deep learning pipeline for multi-class fetal brain segmentation by merging several publicly available datasets. To promote generalisability, our approach does not introduce any additional hyper-parameters tuning.
Label-set Loss Functions for Partial Supervision: Application to Fetal Brain 3D MRI Parcellation
Jul 09, 2021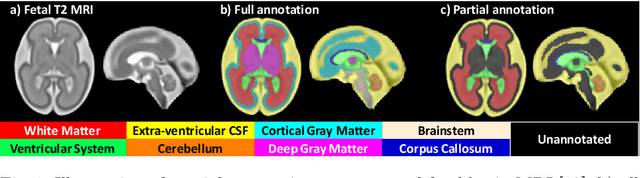
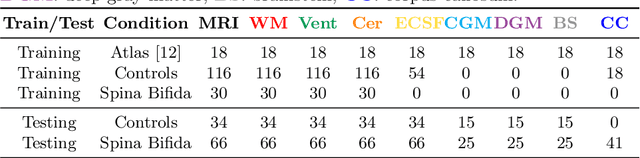
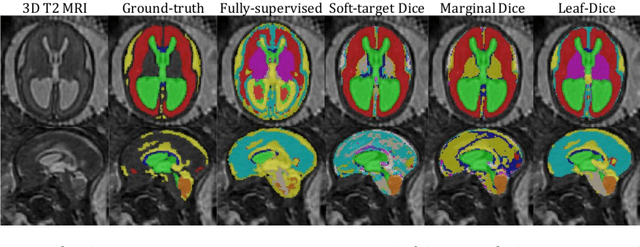
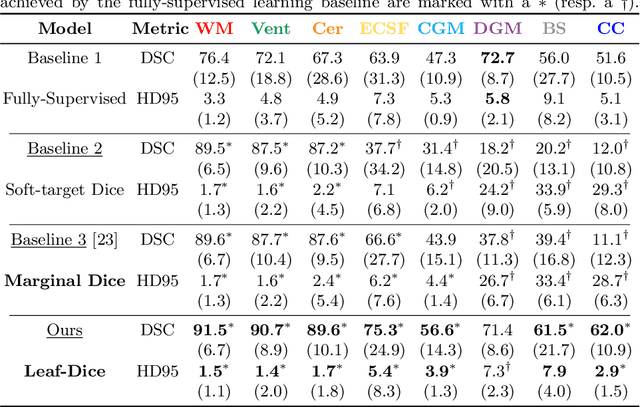
Abstract:Deep neural networks have increased the accuracy of automatic segmentation, however, their accuracy depends on the availability of a large number of fully segmented images. Methods to train deep neural networks using images for which some, but not all, regions of interest are segmented are necessary to make better use of partially annotated datasets. In this paper, we propose the first axiomatic definition of label-set loss functions that are the loss functions that can handle partially segmented images. We prove that there is one and only one method to convert a classical loss function for fully segmented images into a proper label-set loss function. Our theory also allows us to define the leaf-Dice loss, a label-set generalization of the Dice loss particularly suited for partial supervision with only missing labels. Using the leaf-Dice loss, we set a new state of the art in partially supervised learning for fetal brain 3D MRI segmentation. We achieve a deep neural network able to segment white matter, ventricles, cerebellum, extra-ventricular CSF, cortical gray matter, deep gray matter, brainstem, and corpus callosum based on fetal brain 3D MRI of anatomically normal fetuses or with open spina bifida. Our implementation of the proposed label-set loss functions is available at https://github.com/LucasFidon/label-set-loss-functions
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge