Thomas Deprest
A Dempster-Shafer approach to trustworthy AI with application to fetal brain MRI segmentation
Apr 05, 2022
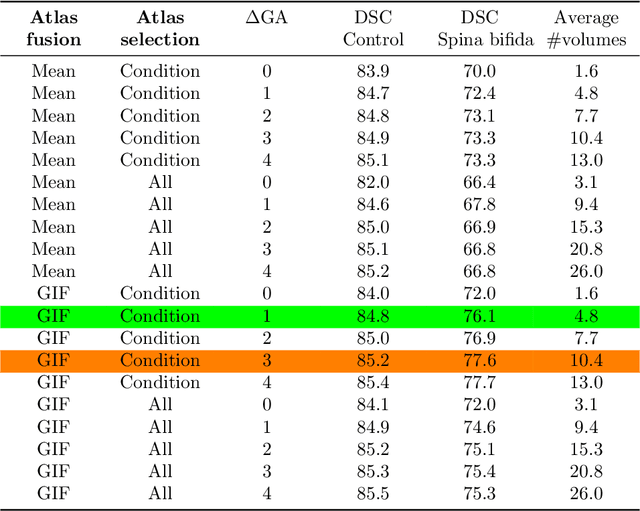
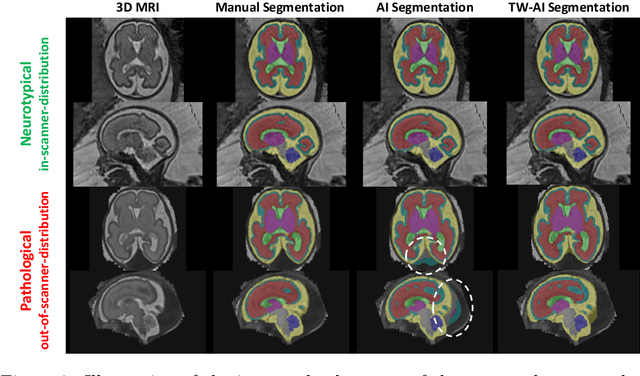
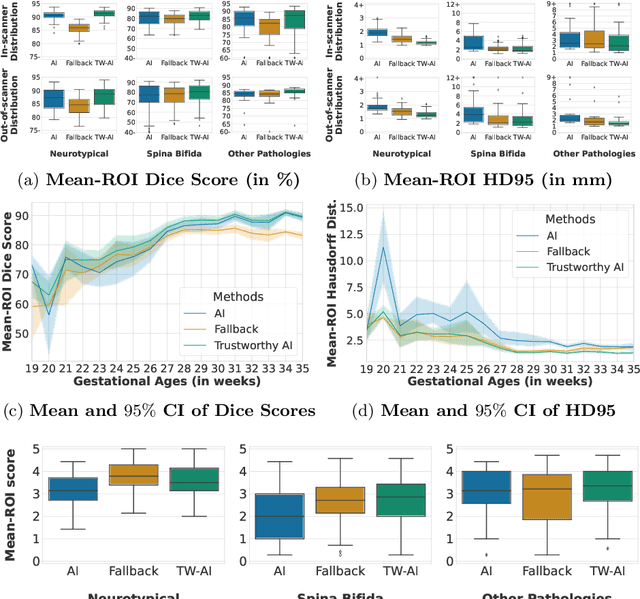
Abstract:Deep learning models for medical image segmentation can fail unexpectedly and spectacularly for pathological cases and for images acquired at different centers than those used for training, with labeling errors that violate expert knowledge about the anatomy and the intensity distribution of the regions to be segmented. Such errors undermine the trustworthiness of deep learning models developed for medical image segmentation. Mechanisms with a fallback method for detecting and correcting such failures are essential for safely translating this technology into clinics and are likely to be a requirement of future regulations on artificial intelligence (AI). Here, we propose a principled trustworthy AI theoretical framework and a practical system that can augment any backbone AI system using a fallback method and a fail-safe mechanism based on Dempster-Shafer theory. Our approach relies on an actionable definition of trustworthy AI. Our method automatically discards the voxel-level labeling predicted by the backbone AI that are likely to violate expert knowledge and relies on a fallback atlas-based segmentation method for those voxels. We demonstrate the effectiveness of the proposed trustworthy AI approach on the largest reported annotated dataset of fetal T2w MRI consisting of 540 manually annotated fetal brain 3D MRIs with neurotypical or abnormal brain development and acquired from 13 sources of data across 6 countries. We show that our trustworthy AI method improves the robustness of a state-of-the-art backbone AI for fetal brain MRI segmentation on MRIs acquired across various centers and for fetuses with various brain abnormalities.
Distributionally Robust Segmentation of Abnormal Fetal Brain 3D MRI
Aug 09, 2021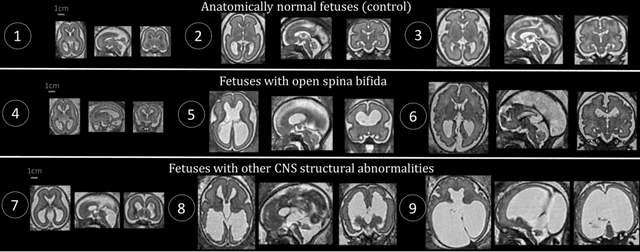
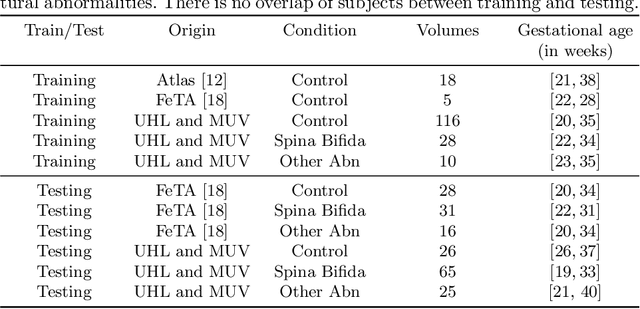
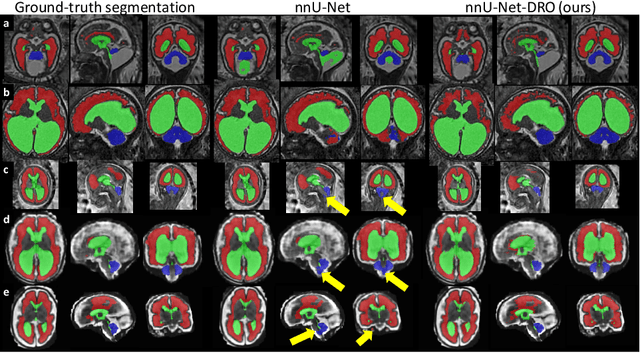
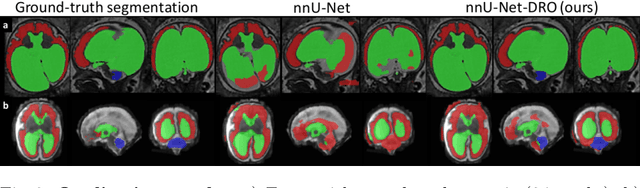
Abstract:The performance of deep neural networks typically increases with the number of training images. However, not all images have the same importance towards improved performance and robustness. In fetal brain MRI, abnormalities exacerbate the variability of the developing brain anatomy compared to non-pathological cases. A small number of abnormal cases, as is typically available in clinical datasets used for training, are unlikely to fairly represent the rich variability of abnormal developing brains. This leads machine learning systems trained by maximizing the average performance to be biased toward non-pathological cases. This problem was recently referred to as hidden stratification. To be suited for clinical use, automatic segmentation methods need to reliably achieve high-quality segmentation outcomes also for pathological cases. In this paper, we show that the state-of-the-art deep learning pipeline nnU-Net has difficulties to generalize to unseen abnormal cases. To mitigate this problem, we propose to train a deep neural network to minimize a percentile of the distribution of per-volume loss over the dataset. We show that this can be achieved by using Distributionally Robust Optimization (DRO). DRO automatically reweights the training samples with lower performance, encouraging nnU-Net to perform more consistently on all cases. We validated our approach using a dataset of 368 fetal brain T2w MRIs, including 124 MRIs of open spina bifida cases and 51 MRIs of cases with other severe abnormalities of brain development.
Label-set Loss Functions for Partial Supervision: Application to Fetal Brain 3D MRI Parcellation
Jul 09, 2021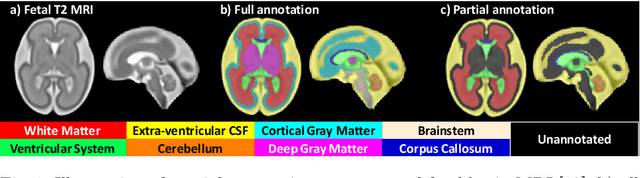
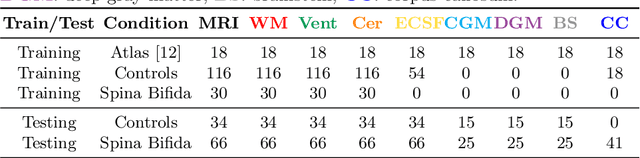
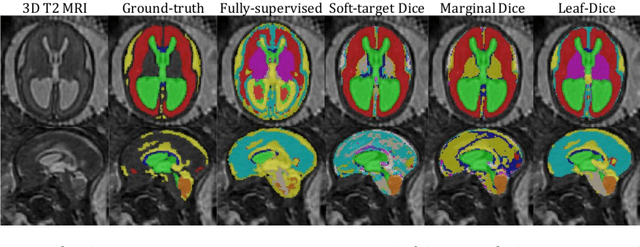
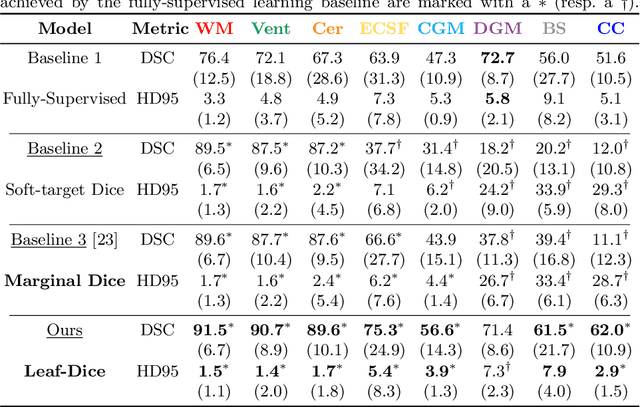
Abstract:Deep neural networks have increased the accuracy of automatic segmentation, however, their accuracy depends on the availability of a large number of fully segmented images. Methods to train deep neural networks using images for which some, but not all, regions of interest are segmented are necessary to make better use of partially annotated datasets. In this paper, we propose the first axiomatic definition of label-set loss functions that are the loss functions that can handle partially segmented images. We prove that there is one and only one method to convert a classical loss function for fully segmented images into a proper label-set loss function. Our theory also allows us to define the leaf-Dice loss, a label-set generalization of the Dice loss particularly suited for partial supervision with only missing labels. Using the leaf-Dice loss, we set a new state of the art in partially supervised learning for fetal brain 3D MRI segmentation. We achieve a deep neural network able to segment white matter, ventricles, cerebellum, extra-ventricular CSF, cortical gray matter, deep gray matter, brainstem, and corpus callosum based on fetal brain 3D MRI of anatomically normal fetuses or with open spina bifida. Our implementation of the proposed label-set loss functions is available at https://github.com/LucasFidon/label-set-loss-functions
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge