Yinghao Zhu
ExperienceWeaver: Optimizing Small-sample Experience Learning for LLM-based Clinical Text Improvement
Jan 31, 2026Abstract:Clinical text improvement is vital for healthcare efficiency but remains difficult due to limited high-quality data and the complex constraints of medical documentation. While Large Language Models (LLMs) show promise, current approaches struggle in small-sample settings: supervised fine-tuning is data-intensive and costly, while retrieval-augmented generation often provides superficial corrections without capturing the reasoning behind revisions. To address these limitations, we propose ExperienceWeaver, a hierarchical framework that shifts the focus from data retrieval to experience learning. Instead of simply recalling past examples, ExperienceWeaver distills noisy, multi-dimensional feedback into structured, actionable knowledge. Specifically, error-specific Tips and high-level Strategies. By injecting this distilled experience into an agentic pipeline, the model learns "how to revise" rather than just "what to revise". Extensive evaluations across four clinical datasets demonstrate that ExperienceWeaver consistently improves performance, surpassing state-of-the-art models such as Gemini-3 Pro in small-sample settings.
Augmenting Clinical Decision-Making with an Interactive and Interpretable AI Copilot: A Real-World User Study with Clinicians in Nephrology and Obstetrics
Jan 31, 2026Abstract:Clinician skepticism toward opaque AI hinders adoption in high-stakes healthcare. We present AICare, an interactive and interpretable AI copilot for collaborative clinical decision-making. By analyzing longitudinal electronic health records, AICare grounds dynamic risk predictions in scrutable visualizations and LLM-driven diagnostic recommendations. Through a within-subjects counterbalanced study with 16 clinicians across nephrology and obstetrics, we comprehensively evaluated AICare using objective measures (task completion time and error rate), subjective assessments (NASA-TLX, SUS, and confidence ratings), and semi-structured interviews. Our findings indicate AICare's reduced cognitive workload. Beyond performance metrics, qualitative analysis reveals that trust is actively constructed through verification, with interaction strategies diverging by expertise: junior clinicians used the system as cognitive scaffolding to structure their analysis, while experts engaged in adversarial verification to challenge the AI's logic. This work offers design implications for creating AI systems that function as transparent partners, accommodating diverse reasoning styles to augment rather than replace clinical judgment.
SearchGym: Bootstrapping Real-World Search Agents via Cost-Effective and High-Fidelity Environment Simulation
Jan 21, 2026Abstract:Search agents have emerged as a pivotal paradigm for solving open-ended, knowledge-intensive reasoning tasks. However, training these agents via Reinforcement Learning (RL) faces a critical dilemma: interacting with live commercial Web APIs is prohibitively expensive, while relying on static data snapshots often introduces noise due to data misalignment. This misalignment generates corrupted reward signals that destabilize training by penalizing correct reasoning or rewarding hallucination. To address this, we propose SearchGym, a simulation environment designed to bootstrap robust search agents. SearchGym employs a rigorous generative pipeline to construct a verifiable knowledge graph and an aligned document corpus, ensuring that every reasoning task is factually grounded and strictly solvable. Building on this controllable environment, we introduce SearchGym-RL, a curriculum learning methodology that progressively optimizes agent policies through purified feedback, evolving from basic interactions to complex, long-horizon planning. Extensive experiments across the Llama and Qwen families demonstrate strong Sim-to-Real generalization. Notably, our Qwen2.5-7B-Base model trained within SearchGym surpasses the web-enhanced ASearcher baseline across nine diverse benchmarks by an average relative margin of 10.6%. Our results validate that high-fidelity simulation serves as a scalable and highly cost-effective methodology for developing capable search agents.
SecureReviewer: Enhancing Large Language Models for Secure Code Review through Secure-aware Fine-tuning
Oct 30, 2025Abstract:Identifying and addressing security issues during the early phase of the development lifecycle is critical for mitigating the long-term negative impacts on software systems. Code review serves as an effective practice that enables developers to check their teammates' code before integration into the codebase. To streamline the generation of review comments, various automated code review approaches have been proposed, where LLM-based methods have significantly advanced the capabilities of automated review generation. However, existing models primarily focus on general-purpose code review, their effectiveness in identifying and addressing security-related issues remains underexplored. Moreover, adapting existing code review approaches to target security issues faces substantial challenges, including data scarcity and inadequate evaluation metrics. To address these limitations, we propose SecureReviewer, a new approach designed for enhancing LLMs' ability to identify and resolve security-related issues during code review. Specifically, we first construct a dataset tailored for training and evaluating secure code review capabilities. Leveraging this dataset, we fine-tune LLMs to generate code review comments that can effectively identify security issues and provide fix suggestions with our proposed secure-aware fine-tuning strategy. To mitigate hallucination in LLMs and enhance the reliability of their outputs, we integrate the RAG technique, which grounds the generated comments in domain-specific security knowledge. Additionally, we introduce SecureBLEU, a new evaluation metric designed to assess the effectiveness of review comments in addressing security issues. Experimental results demonstrate that SecureReviewer outperforms state-of-the-art baselines in both security issue detection accuracy and the overall quality and practical utility of generated review comments.
Scaf-GRPO: Scaffolded Group Relative Policy Optimization for Enhancing LLM Reasoning
Oct 22, 2025Abstract:Reinforcement learning from verifiable rewards has emerged as a powerful technique for enhancing the complex reasoning abilities of Large Language Models (LLMs). However, these methods are fundamentally constrained by the ''learning cliff'' phenomenon: when faced with problems far beyond their current capabilities, models consistently fail, yielding a persistent zero-reward signal. In policy optimization algorithms like GRPO, this collapses the advantage calculation to zero, rendering these difficult problems invisible to the learning gradient and stalling progress. To overcome this, we introduce Scaf-GRPO (Scaffolded Group Relative Policy Optimization), a progressive training framework that strategically provides minimal guidance only when a model's independent learning has plateaued. The framework first diagnoses learning stagnation and then intervenes by injecting tiered in-prompt hints, ranging from abstract concepts to concrete steps, enabling the model to construct a valid solution by itself. Extensive experiments on challenging mathematics benchmarks demonstrate Scaf-GRPO's effectiveness, boosting the pass@1 score of the Qwen2.5-Math-7B model on the AIME24 benchmark by a relative 44.3% over a vanilla GRPO baseline. This result demonstrates our framework provides a robust and effective methodology for unlocking a model's ability to solve problems previously beyond its reach, a critical step towards extending the frontier of autonomous reasoning in LLM.
An Empirical Study on Failures in Automated Issue Solving
Sep 17, 2025Abstract:Automated issue solving seeks to autonomously identify and repair defective code snippets across an entire codebase. SWE-Bench has emerged as the most widely adopted benchmark for evaluating progress in this area. While LLM-based agentic tools show great promise, they still fail on a substantial portion of tasks. Moreover, current evaluations primarily report aggregate issue-solving rates, which obscure the underlying causes of success and failure, making it challenging to diagnose model weaknesses or guide targeted improvements. To bridge this gap, we first analyze the performance and efficiency of three SOTA tools, spanning both pipeline-based and agentic architectures, in automated issue solving tasks of SWE-Bench-Verified under varying task characteristics. Furthermore, to move from high-level performance metrics to underlying cause analysis, we conducted a systematic manual analysis of 150 failed instances. From this analysis, we developed a comprehensive taxonomy of failure modes comprising 3 primary phases, 9 main categories, and 25 fine-grained subcategories. Then we systematically analyze the distribution of the identified failure modes, the results reveal distinct failure fingerprints between the two architectural paradigms, with the majority of agentic failures stemming from flawed reasoning and cognitive deadlocks. Motivated by these insights, we propose a collaborative Expert-Executor framework. It introduces a supervisory Expert agent tasked with providing strategic oversight and course-correction for a primary Executor agent. This architecture is designed to correct flawed reasoning and break the cognitive deadlocks that frequently lead to failure. Experiments show that our framework solves 22.2% of previously intractable issues for a leading single agent. These findings pave the way for building more robust agents through diagnostic evaluation and collaborative design.
Magical: Medical Lay Language Generation via Semantic Invariance and Layperson-tailored Adaptation
Aug 12, 2025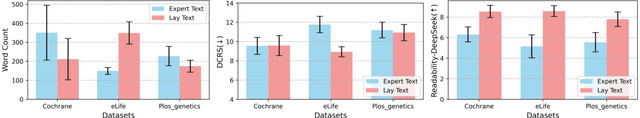
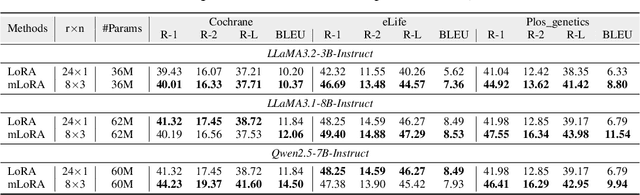
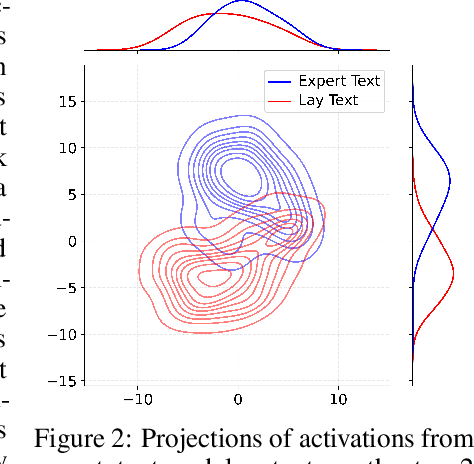
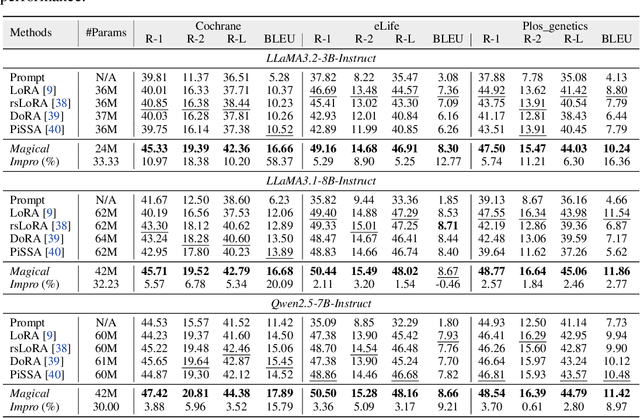
Abstract:Medical Lay Language Generation (MLLG) plays a vital role in improving the accessibility of complex scientific content for broader audiences. Recent literature to MLLG commonly employ parameter-efficient fine-tuning methods such as Low-Rank Adaptation (LoRA) to fine-tuning large language models (LLMs) using paired expert-lay language datasets. However, LoRA struggles with the challenges posed by multi-source heterogeneous MLLG datasets. Specifically, through a series of exploratory experiments, we reveal that standard LoRA fail to meet the requirement for semantic fidelity and diverse lay-style generation in MLLG task. To address these limitations, we propose Magical, an asymmetric LoRA architecture tailored for MLLG under heterogeneous data scenarios. Magical employs a shared matrix $A$ for abstractive summarization, along with multiple isolated matrices $B$ for diverse lay-style generation. To preserve semantic fidelity during the lay language generation process, Magical introduces a Semantic Invariance Constraint to mitigate semantic subspace shifts on matrix $A$. Furthermore, to better adapt to diverse lay-style generation, Magical incorporates the Recommendation-guided Switch, an externally interface to prompt the LLM to switch between different matrices $B$. Experimental results on three real-world lay language generation datasets demonstrate that Magical consistently outperforms prompt-based methods, vanilla LoRA, and its recent variants, while also reducing trainable parameters by 31.66%.
ConfAgents: A Conformal-Guided Multi-Agent Framework for Cost-Efficient Medical Diagnosis
Aug 06, 2025



Abstract:The efficacy of AI agents in healthcare research is hindered by their reliance on static, predefined strategies. This creates a critical limitation: agents can become better tool-users but cannot learn to become better strategic planners, a crucial skill for complex domains like healthcare. We introduce HealthFlow, a self-evolving AI agent that overcomes this limitation through a novel meta-level evolution mechanism. HealthFlow autonomously refines its own high-level problem-solving policies by distilling procedural successes and failures into a durable, strategic knowledge base. To anchor our research and facilitate reproducible evaluation, we introduce EHRFlowBench, a new benchmark featuring complex, realistic health data analysis tasks derived from peer-reviewed clinical research. Our comprehensive experiments demonstrate that HealthFlow's self-evolving approach significantly outperforms state-of-the-art agent frameworks. This work marks a necessary shift from building better tool-users to designing smarter, self-evolving task-managers, paving the way for more autonomous and effective AI for scientific discovery.
SuperGS: Consistent and Detailed 3D Super-Resolution Scene Reconstruction via Gaussian Splatting
May 24, 2025Abstract:Recently, 3D Gaussian Splatting (3DGS) has excelled in novel view synthesis (NVS) with its real-time rendering capabilities and superior quality. However, it encounters challenges for high-resolution novel view synthesis (HRNVS) due to the coarse nature of primitives derived from low-resolution input views. To address this issue, we propose SuperGS, an expansion of Scaffold-GS designed with a two-stage coarse-to-fine training framework. In the low-resolution stage, we introduce a latent feature field to represent the low-resolution scene, which serves as both the initialization and foundational information for super-resolution optimization. In the high-resolution stage, we propose a multi-view consistent densification strategy that backprojects high-resolution depth maps based on error maps and employs a multi-view voting mechanism, mitigating ambiguities caused by multi-view inconsistencies in the pseudo labels provided by 2D prior models while avoiding Gaussian redundancy. Furthermore, we model uncertainty through variational feature learning and use it to guide further scene representation refinement and adjust the supervisory effect of pseudo-labels, ensuring consistent and detailed scene reconstruction. Extensive experiments demonstrate that SuperGS outperforms state-of-the-art HRNVS methods on both forward-facing and 360-degree datasets.
MedAgentBoard: Benchmarking Multi-Agent Collaboration with Conventional Methods for Diverse Medical Tasks
May 18, 2025Abstract:The rapid advancement of Large Language Models (LLMs) has stimulated interest in multi-agent collaboration for addressing complex medical tasks. However, the practical advantages of multi-agent collaboration approaches remain insufficiently understood. Existing evaluations often lack generalizability, failing to cover diverse tasks reflective of real-world clinical practice, and frequently omit rigorous comparisons against both single-LLM-based and established conventional methods. To address this critical gap, we introduce MedAgentBoard, a comprehensive benchmark for the systematic evaluation of multi-agent collaboration, single-LLM, and conventional approaches. MedAgentBoard encompasses four diverse medical task categories: (1) medical (visual) question answering, (2) lay summary generation, (3) structured Electronic Health Record (EHR) predictive modeling, and (4) clinical workflow automation, across text, medical images, and structured EHR data. Our extensive experiments reveal a nuanced landscape: while multi-agent collaboration demonstrates benefits in specific scenarios, such as enhancing task completeness in clinical workflow automation, it does not consistently outperform advanced single LLMs (e.g., in textual medical QA) or, critically, specialized conventional methods that generally maintain better performance in tasks like medical VQA and EHR-based prediction. MedAgentBoard offers a vital resource and actionable insights, emphasizing the necessity of a task-specific, evidence-based approach to selecting and developing AI solutions in medicine. It underscores that the inherent complexity and overhead of multi-agent collaboration must be carefully weighed against tangible performance gains. All code, datasets, detailed prompts, and experimental results are open-sourced at https://medagentboard.netlify.app/.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge