Winnie Pang
Towards Robust and Reliable Concept Representations: Reliability-Enhanced Concept Embedding Model
Feb 03, 2025



Abstract:Concept Bottleneck Models (CBMs) aim to enhance interpretability by predicting human-understandable concepts as intermediates for decision-making. However, these models often face challenges in ensuring reliable concept representations, which can propagate to downstream tasks and undermine robustness, especially under distribution shifts. Two inherent issues contribute to concept unreliability: sensitivity to concept-irrelevant features (e.g., background variations) and lack of semantic consistency for the same concept across different samples. To address these limitations, we propose the Reliability-Enhanced Concept Embedding Model (RECEM), which introduces a two-fold strategy: Concept-Level Disentanglement to separate irrelevant features from concept-relevant information and a Concept Mixup mechanism to ensure semantic alignment across samples. These mechanisms work together to improve concept reliability, enabling the model to focus on meaningful object attributes and generate faithful concept representations. Experimental results demonstrate that RECEM consistently outperforms existing baselines across multiple datasets, showing superior performance under background and domain shifts. These findings highlight the effectiveness of disentanglement and alignment strategies in enhancing both reliability and robustness in CBMs.
Integrating Clinical Knowledge into Concept Bottleneck Models
Jul 09, 2024Abstract:Concept bottleneck models (CBMs), which predict human-interpretable concepts (e.g., nucleus shapes in cell images) before predicting the final output (e.g., cell type), provide insights into the decision-making processes of the model. However, training CBMs solely in a data-driven manner can introduce undesirable biases, which may compromise prediction performance, especially when the trained models are evaluated on out-of-domain images (e.g., those acquired using different devices). To mitigate this challenge, we propose integrating clinical knowledge to refine CBMs, better aligning them with clinicians' decision-making processes. Specifically, we guide the model to prioritize the concepts that clinicians also prioritize. We validate our approach on two datasets of medical images: white blood cell and skin images. Empirical validation demonstrates that incorporating medical guidance enhances the model's classification performance on unseen datasets with varying preparation methods, thereby increasing its real-world applicability.
WBCAtt: A White Blood Cell Dataset Annotated with Detailed Morphological Attributes
Jun 23, 2023



Abstract:The examination of blood samples at a microscopic level plays a fundamental role in clinical diagnostics, influencing a wide range of medical conditions. For instance, an in-depth study of White Blood Cells (WBCs), a crucial component of our blood, is essential for diagnosing blood-related diseases such as leukemia and anemia. While multiple datasets containing WBC images have been proposed, they mostly focus on cell categorization, often lacking the necessary morphological details to explain such categorizations, despite the importance of explainable artificial intelligence (XAI) in medical domains. This paper seeks to address this limitation by introducing comprehensive annotations for WBC images. Through collaboration with pathologists, a thorough literature review, and manual inspection of microscopic images, we have identified 11 morphological attributes associated with the cell and its components (nucleus, cytoplasm, and granules). We then annotated ten thousand WBC images with these attributes. Moreover, we conduct experiments to predict these attributes from images, providing insights beyond basic WBC classification. As the first public dataset to offer such extensive annotations, we also illustrate specific applications that can benefit from our attribute annotations. Overall, our dataset paves the way for interpreting WBC recognition models, further advancing XAI in the fields of pathology and hematology.
Surgical tool classification and localization: results and methods from the MICCAI 2022 SurgToolLoc challenge
May 11, 2023



Abstract:The ability to automatically detect and track surgical instruments in endoscopic videos can enable transformational interventions. Assessing surgical performance and efficiency, identifying skilled tool use and choreography, and planning operational and logistical aspects of OR resources are just a few of the applications that could benefit. Unfortunately, obtaining the annotations needed to train machine learning models to identify and localize surgical tools is a difficult task. Annotating bounding boxes frame-by-frame is tedious and time-consuming, yet large amounts of data with a wide variety of surgical tools and surgeries must be captured for robust training. Moreover, ongoing annotator training is needed to stay up to date with surgical instrument innovation. In robotic-assisted surgery, however, potentially informative data like timestamps of instrument installation and removal can be programmatically harvested. The ability to rely on tool installation data alone would significantly reduce the workload to train robust tool-tracking models. With this motivation in mind we invited the surgical data science community to participate in the challenge, SurgToolLoc 2022. The goal was to leverage tool presence data as weak labels for machine learning models trained to detect tools and localize them in video frames with bounding boxes. We present the results of this challenge along with many of the team's efforts. We conclude by discussing these results in the broader context of machine learning and surgical data science. The training data used for this challenge consisting of 24,695 video clips with tool presence labels is also being released publicly and can be accessed at https://console.cloud.google.com/storage/browser/isi-surgtoolloc-2022.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
CholecTriplet2021: A benchmark challenge for surgical action triplet recognition
Apr 10, 2022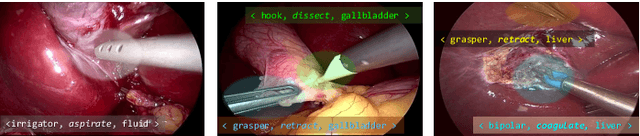
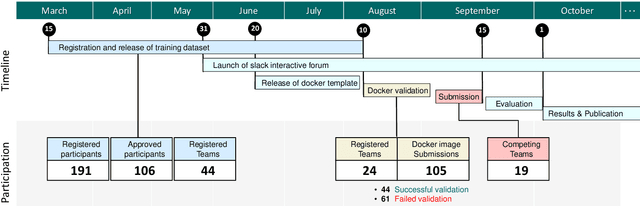
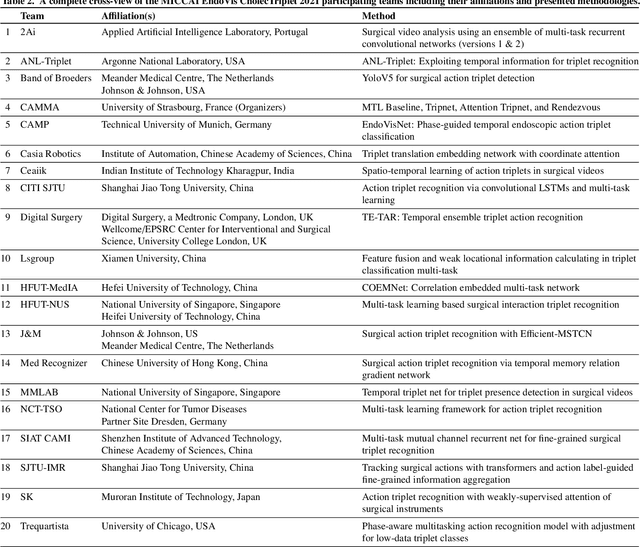
Abstract:Context-aware decision support in the operating room can foster surgical safety and efficiency by leveraging real-time feedback from surgical workflow analysis. Most existing works recognize surgical activities at a coarse-grained level, such as phases, steps or events, leaving out fine-grained interaction details about the surgical activity; yet those are needed for more helpful AI assistance in the operating room. Recognizing surgical actions as triplets of <instrument, verb, target> combination delivers comprehensive details about the activities taking place in surgical videos. This paper presents CholecTriplet2021: an endoscopic vision challenge organized at MICCAI 2021 for the recognition of surgical action triplets in laparoscopic videos. The challenge granted private access to the large-scale CholecT50 dataset, which is annotated with action triplet information. In this paper, we present the challenge setup and assessment of the state-of-the-art deep learning methods proposed by the participants during the challenge. A total of 4 baseline methods from the challenge organizers and 19 new deep learning algorithms by competing teams are presented to recognize surgical action triplets directly from surgical videos, achieving mean average precision (mAP) ranging from 4.2% to 38.1%. This study also analyzes the significance of the results obtained by the presented approaches, performs a thorough methodological comparison between them, in-depth result analysis, and proposes a novel ensemble method for enhanced recognition. Our analysis shows that surgical workflow analysis is not yet solved, and also highlights interesting directions for future research on fine-grained surgical activity recognition which is of utmost importance for the development of AI in surgery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge