Sihang Zeng
A Survey of Reinforcement Learning for Large Reasoning Models
Sep 10, 2025Abstract:In this paper, we survey recent advances in Reinforcement Learning (RL) for reasoning with Large Language Models (LLMs). RL has achieved remarkable success in advancing the frontier of LLM capabilities, particularly in addressing complex logical tasks such as mathematics and coding. As a result, RL has emerged as a foundational methodology for transforming LLMs into LRMs. With the rapid progress of the field, further scaling of RL for LRMs now faces foundational challenges not only in computational resources but also in algorithm design, training data, and infrastructure. To this end, it is timely to revisit the development of this domain, reassess its trajectory, and explore strategies to enhance the scalability of RL toward Artificial SuperIntelligence (ASI). In particular, we examine research applying RL to LLMs and LRMs for reasoning abilities, especially since the release of DeepSeek-R1, including foundational components, core problems, training resources, and downstream applications, to identify future opportunities and directions for this rapidly evolving area. We hope this review will promote future research on RL for broader reasoning models. Github: https://github.com/TsinghuaC3I/Awesome-RL-for-LRMs
ReviewRL: Towards Automated Scientific Review with RL
Aug 14, 2025Abstract:Peer review is essential for scientific progress but faces growing challenges due to increasing submission volumes and reviewer fatigue. Existing automated review approaches struggle with factual accuracy, rating consistency, and analytical depth, often generating superficial or generic feedback lacking the insights characteristic of high-quality human reviews. We introduce ReviewRL, a reinforcement learning framework for generating comprehensive and factually grounded scientific paper reviews. Our approach combines: (1) an ArXiv-MCP retrieval-augmented context generation pipeline that incorporates relevant scientific literature, (2) supervised fine-tuning that establishes foundational reviewing capabilities, and (3) a reinforcement learning procedure with a composite reward function that jointly enhances review quality and rating accuracy. Experiments on ICLR 2025 papers demonstrate that ReviewRL significantly outperforms existing methods across both rule-based metrics and model-based quality assessments. ReviewRL establishes a foundational framework for RL-driven automatic critique generation in scientific discovery, demonstrating promising potential for future development in this domain. The implementation of ReviewRL will be released at GitHub.
TrajSurv: Learning Continuous Latent Trajectories from Electronic Health Records for Trustworthy Survival Prediction
Aug 01, 2025Abstract:Trustworthy survival prediction is essential for clinical decision making. Longitudinal electronic health records (EHRs) provide a uniquely powerful opportunity for the prediction. However, it is challenging to accurately model the continuous clinical progression of patients underlying the irregularly sampled clinical features and to transparently link the progression to survival outcomes. To address these challenges, we develop TrajSurv, a model that learns continuous latent trajectories from longitudinal EHR data for trustworthy survival prediction. TrajSurv employs a neural controlled differential equation (NCDE) to extract continuous-time latent states from the irregularly sampled data, forming continuous latent trajectories. To ensure the latent trajectories reflect the clinical progression, TrajSurv aligns the latent state space with patient state space through a time-aware contrastive learning approach. To transparently link clinical progression to the survival outcome, TrajSurv uses latent trajectories in a two-step divide-and-conquer interpretation process. First, it explains how the changes in clinical features translate into the latent trajectory's evolution using a learned vector field. Second, it clusters these latent trajectories to identify key clinical progression patterns associated with different survival outcomes. Evaluations on two real-world medical datasets, MIMIC-III and eICU, show TrajSurv's competitive accuracy and superior transparency over existing deep learning methods.
Large Language Models as Biomedical Hypothesis Generators: A Comprehensive Evaluation
Jul 12, 2024
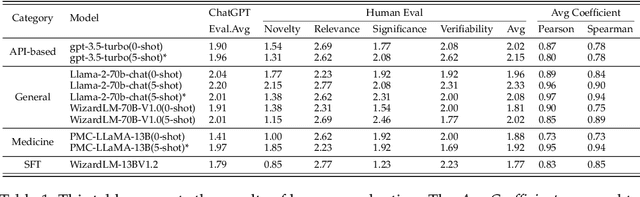
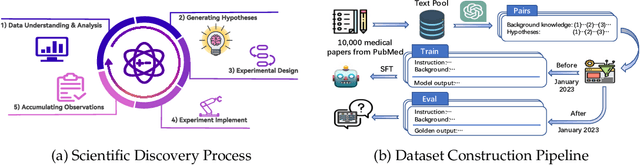

Abstract:The rapid growth of biomedical knowledge has outpaced our ability to efficiently extract insights and generate novel hypotheses. Large language models (LLMs) have emerged as a promising tool to revolutionize knowledge interaction and potentially accelerate biomedical discovery. In this paper, we present a comprehensive evaluation of LLMs as biomedical hypothesis generators. We construct a dataset of background-hypothesis pairs from biomedical literature, carefully partitioned into training, seen, and unseen test sets based on publication date to mitigate data contamination. Using this dataset, we assess the hypothesis generation capabilities of top-tier instructed models in zero-shot, few-shot, and fine-tuning settings. To enhance the exploration of uncertainty, a crucial aspect of scientific discovery, we incorporate tool use and multi-agent interactions in our evaluation framework. Furthermore, we propose four novel metrics grounded in extensive literature review to evaluate the quality of generated hypotheses, considering both LLM-based and human assessments. Our experiments yield two key findings: 1) LLMs can generate novel and validated hypotheses, even when tested on literature unseen during training, and 2) Increasing uncertainty through multi-agent interactions and tool use can facilitate diverse candidate generation and improve zero-shot hypothesis generation performance. However, we also observe that the integration of additional knowledge through few-shot learning and tool use may not always lead to performance gains, highlighting the need for careful consideration of the type and scope of external knowledge incorporated. These findings underscore the potential of LLMs as powerful aids in biomedical hypothesis generation and provide valuable insights to guide further research in this area.
UltraMedical: Building Specialized Generalists in Biomedicine
Jun 06, 2024Abstract:Large Language Models (LLMs) have demonstrated remarkable capabilities across various domains and are moving towards more specialized areas. Recent advanced proprietary models such as GPT-4 and Gemini have achieved significant advancements in biomedicine, which have also raised privacy and security challenges. The construction of specialized generalists hinges largely on high-quality datasets, enhanced by techniques like supervised fine-tuning and reinforcement learning from human or AI feedback, and direct preference optimization. However, these leading technologies (e.g., preference learning) are still significantly limited in the open source community due to the scarcity of specialized data. In this paper, we present the UltraMedical collections, which consist of high-quality manual and synthetic datasets in the biomedicine domain, featuring preference annotations across multiple advanced LLMs. By utilizing these datasets, we fine-tune a suite of specialized medical models based on Llama-3 series, demonstrating breathtaking capabilities across various medical benchmarks. Moreover, we develop powerful reward models skilled in biomedical and general reward benchmark, enhancing further online preference learning within the biomedical LLM community.
CoRTEx: Contrastive Learning for Representing Terms via Explanations with Applications on Constructing Biomedical Knowledge Graphs
Dec 13, 2023

Abstract:Objective: Biomedical Knowledge Graphs play a pivotal role in various biomedical research domains. Concurrently, term clustering emerges as a crucial step in constructing these knowledge graphs, aiming to identify synonymous terms. Due to a lack of knowledge, previous contrastive learning models trained with Unified Medical Language System (UMLS) synonyms struggle at clustering difficult terms and do not generalize well beyond UMLS terms. In this work, we leverage the world knowledge from Large Language Models (LLMs) and propose Contrastive Learning for Representing Terms via Explanations (CoRTEx) to enhance term representation and significantly improves term clustering. Materials and Methods: The model training involves generating explanations for a cleaned subset of UMLS terms using ChatGPT. We employ contrastive learning, considering term and explanation embeddings simultaneously, and progressively introduce hard negative samples. Additionally, a ChatGPT-assisted BIRCH algorithm is designed for efficient clustering of a new ontology. Results: We established a clustering test set and a hard negative test set, where our model consistently achieves the highest F1 score. With CoRTEx embeddings and the modified BIRCH algorithm, we grouped 35,580,932 terms from the Biomedical Informatics Ontology System (BIOS) into 22,104,559 clusters with O(N) queries to ChatGPT. Case studies highlight the model's efficacy in handling challenging samples, aided by information from explanations. Conclusion: By aligning terms to their explanations, CoRTEx demonstrates superior accuracy over benchmark models and robustness beyond its training set, and it is suitable for clustering terms for large-scale biomedical ontologies.
Large Language Models are Zero Shot Hypothesis Proposers
Nov 10, 2023



Abstract:Significant scientific discoveries have driven the progress of human civilisation. The explosion of scientific literature and data has created information barriers across disciplines that have slowed the pace of scientific discovery. Large Language Models (LLMs) hold a wealth of global and interdisciplinary knowledge that promises to break down these information barriers and foster a new wave of scientific discovery. However, the potential of LLMs for scientific discovery has not been formally explored. In this paper, we start from investigating whether LLMs can propose scientific hypotheses. To this end, we construct a dataset consist of background knowledge and hypothesis pairs from biomedical literature. The dataset is divided into training, seen, and unseen test sets based on the publication date to control visibility. We subsequently evaluate the hypothesis generation capabilities of various top-tier instructed models in zero-shot, few-shot, and fine-tuning settings, including both closed and open-source LLMs. Additionally, we introduce an LLM-based multi-agent cooperative framework with different role designs and external tools to enhance the capabilities related to generating hypotheses. We also design four metrics through a comprehensive review to evaluate the generated hypotheses for both ChatGPT-based and human evaluations. Through experiments and analyses, we arrive at the following findings: 1) LLMs surprisingly generate untrained yet validated hypotheses from testing literature. 2) Increasing uncertainty facilitates candidate generation, potentially enhancing zero-shot hypothesis generation capabilities. These findings strongly support the potential of LLMs as catalysts for new scientific discoveries and guide further exploration.
Hierarchical Pretraining for Biomedical Term Embeddings
Jul 01, 2023


Abstract:Electronic health records (EHR) contain narrative notes that provide extensive details on the medical condition and management of patients. Natural language processing (NLP) of clinical notes can use observed frequencies of clinical terms as predictive features for downstream applications such as clinical decision making and patient trajectory prediction. However, due to the vast number of highly similar and related clinical concepts, a more effective modeling strategy is to represent clinical terms as semantic embeddings via representation learning and use the low dimensional embeddings as feature vectors for predictive modeling. To achieve efficient representation, fine-tuning pretrained language models with biomedical knowledge graphs may generate better embeddings for biomedical terms than those from standard language models alone. These embeddings can effectively discriminate synonymous pairs of from those that are unrelated. However, they often fail to capture different degrees of similarity or relatedness for concepts that are hierarchical in nature. To overcome this limitation, we propose HiPrBERT, a novel biomedical term representation model trained on additionally complied data that contains hierarchical structures for various biomedical terms. We modify an existing contrastive loss function to extract information from these hierarchies. Our numerical experiments demonstrate that HiPrBERT effectively learns the pair-wise distance from hierarchical information, resulting in a substantially more informative embeddings for further biomedical applications
Automatic Biomedical Term Clustering by Learning Fine-grained Term Representations
Apr 01, 2022



Abstract:Term clustering is important in biomedical knowledge graph construction. Using similarities between terms embedding is helpful for term clustering. State-of-the-art term embeddings leverage pretrained language models to encode terms, and use synonyms and relation knowledge from knowledge graphs to guide contrastive learning. These embeddings provide close embeddings for terms belonging to the same concept. However, from our probing experiments, these embeddings are not sensitive to minor textual differences which leads to failure for biomedical term clustering. To alleviate this problem, we adjust the sampling strategy in pretraining term embeddings by providing dynamic hard positive and negative samples during contrastive learning to learn fine-grained representations which result in better biomedical term clustering. We name our proposed method as CODER++, and it has been applied in clustering biomedical concepts in the newly released Biomedical Knowledge Graph named BIOS.
BIOS: An Algorithmically Generated Biomedical Knowledge Graph
Mar 18, 2022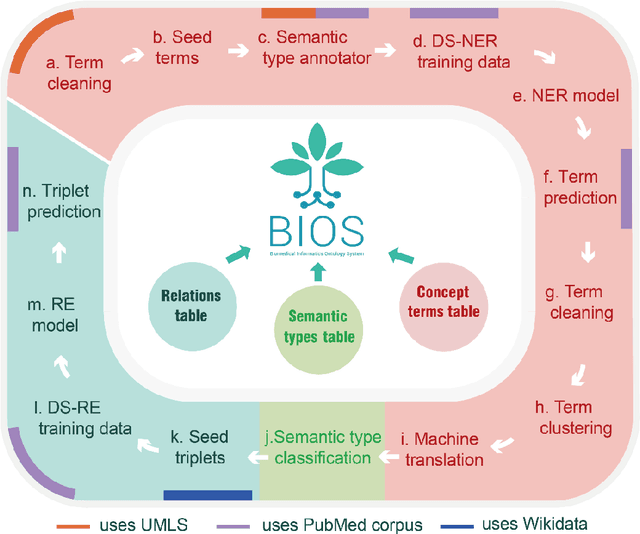
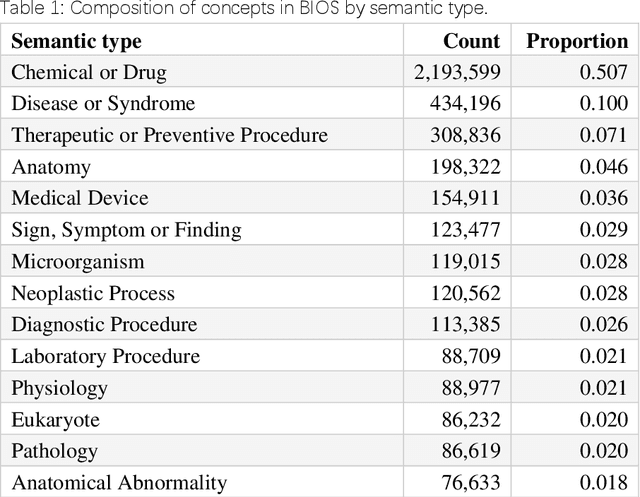
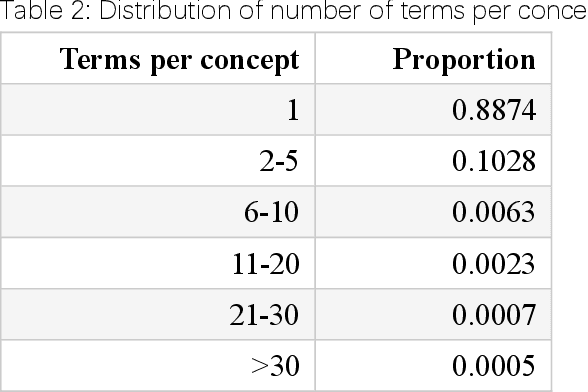

Abstract:Biomedical knowledge graphs (BioMedKGs) are essential infrastructures for biomedical and healthcare big data and artificial intelligence (AI), facilitating natural language processing, model development, and data exchange. For many decades, these knowledge graphs have been built via expert curation, which can no longer catch up with the speed of today's AI development, and a transition to algorithmically generated BioMedKGs is necessary. In this work, we introduce the Biomedical Informatics Ontology System (BIOS), the first large scale publicly available BioMedKG that is fully generated by machine learning algorithms. BIOS currently contains 4.1 million concepts, 7.4 million terms in two languages, and 7.3 million relation triplets. We introduce the methodology for developing BIOS, which covers curation of raw biomedical terms, computationally identifying synonymous terms and aggregating them to create concept nodes, semantic type classification of the concepts, relation identification, and biomedical machine translation. We provide statistics about the current content of BIOS and perform preliminary assessment for term quality, synonym grouping, and relation extraction. Results suggest that machine learning-based BioMedKG development is a totally viable solution for replacing traditional expert curation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge