Puneet Sharma
LGESynthNet: Controlled Scar Synthesis for Improved Scar Segmentation in Cardiac LGE-MRI Imaging
Mar 18, 2026Abstract:Segmentation of enhancement in LGE cardiac MRI is critical for diagnosing various ischemic and non-ischemic cardiomyopathies. However, creating pixel-level annotations for these images is challenging and labor-intensive, leading to limited availability of annotated data. Generative models, particularly diffusion models, offer promise for synthetic data generation, yet many rely on large training datasets and often struggle with fine-grained conditioning control, especially for small or localized features. We introduce LGESynthNet, a latent diffusion-based framework for controllable enhancement synthesis, enabling explicit control over size, location, and transmural extent. Formulated as inpainting using a ControlNet-based architecture, the model integrates: (a) a reward model for conditioning-specific supervision, (b) a captioning module for anatomically descriptive text prompts, and (c) a biomedical text encoder. Trained on just 429 images (79 patients), it produces realistic, anatomically coherent samples. A quality control filter selects outputs with high conditioning-fidelity, which when used for training augmentation, improve downstream segmentation and detection performance, by up-to 6 and 20 points respectively.
Resilience Meets Autonomy: Governing Embodied AI in Critical Infrastructure
Mar 16, 2026Abstract:Critical infrastructure increasingly incorporates embodied AI for monitoring, predictive maintenance, and decision support. However, AI systems designed to handle statistically representable uncertainty struggle with cascading failures and crisis dynamics that exceed their training assumptions. This paper argues that Embodied AIs resilience depends on bounded autonomy within a hybrid governance architecture. We outline four oversight modes and map them to critical infrastructure sectors based on task complexity, risk level, and consequence severity. Drawing on the EU AI Act, ISO safety standards, and crisis management research, we argue that effective governance requires a structured allocation of machine capability and human judgement.
Hippocampus: An Efficient and Scalable Memory Module for Agentic AI
Feb 14, 2026Abstract:Agentic AI require persistent memory to store user-specific histories beyond the limited context window of LLMs. Existing memory systems use dense vector databases or knowledge-graph traversal (or hybrid), incurring high retrieval latency and poor storage scalability. We introduce Hippocampus, an agentic memory management system that uses compact binary signatures for semantic search and lossless token-ID streams for exact content reconstruction. Its core is a Dynamic Wavelet Matrix (DWM) that compresses and co-indexes both streams to support ultra-fast search in the compressed domain, thus avoiding costly dense-vector or graph computations. This design scales linearly with memory size, making it suitable for long-horizon agentic deployments. Empirically, our evaluation shows that Hippocampus reduces end-to-end retrieval latency by up to 31$\times$ and cuts per-query token footprint by up to 14$\times$, while maintaining accuracy on both LoCoMo and LongMemEval benchmarks.
Using Multi-Instance Learning to Identify Unique Polyps in Colon Capsule Endoscopy Images
Jan 21, 2026Abstract:Identifying unique polyps in colon capsule endoscopy (CCE) images is a critical yet challenging task for medical personnel due to the large volume of images, the cognitive load it creates for clinicians, and the ambiguity in labeling specific frames. This paper formulates this problem as a multi-instance learning (MIL) task, where a query polyp image is compared with a target bag of images to determine uniqueness. We employ a multi-instance verification (MIV) framework that incorporates attention mechanisms, such as variance-excited multi-head attention (VEMA) and distance-based attention (DBA), to enhance the model's ability to extract meaningful representations. Additionally, we investigate the impact of self-supervised learning using SimCLR to generate robust embeddings. Experimental results on a dataset of 1912 polyps from 754 patients demonstrate that attention mechanisms significantly improve performance, with DBA L1 achieving the highest test accuracy of 86.26\% and a test AUC of 0.928 using a ConvNeXt backbone with SimCLR pretraining. This study underscores the potential of MIL and self-supervised learning in advancing automated analysis of Colon Capsule Endoscopy images, with implications for broader medical imaging applications.
Using deep learning for predicting cleansing quality of colon capsule endoscopy images
Jan 19, 2026Abstract:In this study, we explore the application of deep learning techniques for predicting cleansing quality in colon capsule endoscopy (CCE) images. Using a dataset of 500 images labeled by 14 clinicians on the Leighton-Rex scale (Poor, Fair, Good, and Excellent), a ResNet-18 model was trained for classification, leveraging stratified K-fold cross-validation to ensure robust performance. To optimize the model, structured pruning techniques were applied iteratively, achieving significant sparsity while maintaining high accuracy. Explainability of the pruned model was evaluated using Grad-CAM, Grad-CAM++, Eigen-CAM, Ablation-CAM, and Random-CAM, with the ROAD method employed for consistent evaluation. Our results indicate that for a pruned model, we can achieve a cross-validation accuracy of 88% with 79% sparsity, demonstrating the effectiveness of pruning in improving efficiency from 84% without compromising performance. We also highlight the challenges of evaluating cleansing quality of CCE images, emphasize the importance of explainability in clinical applications, and discuss the challenges associated with using the ROAD method for our task. Finally, we employ a variant of adaptive temperature scaling to calibrate the pruned models for an external dataset.
EchoVLM: Measurement-Grounded Multimodal Learning for Echocardiography
Dec 13, 2025Abstract:Echocardiography is the most widely used imaging modality in cardiology, yet its interpretation remains labor-intensive and inherently multimodal, requiring view recognition, quantitative measurements, qualitative assessments, and guideline-based reasoning. While recent vision-language models (VLMs) have achieved broad success in natural images and certain medical domains, their potential in echocardiography has been limited by the lack of large-scale, clinically grounded image-text datasets and the absence of measurement-based reasoning central to echo interpretation. We introduce EchoGround-MIMIC, the first measurement-grounded multimodal echocardiography dataset, comprising 19,065 image-text pairs from 1,572 patients with standardized views, structured measurements, measurement-grounded captions, and guideline-derived disease labels. Building on this resource, we propose EchoVLM, a vision-language model that incorporates two novel pretraining objectives: (i) a view-informed contrastive loss that encodes the view-dependent structure of echocardiographic imaging, and (ii) a negation-aware contrastive loss that distinguishes clinically critical negative from positive findings. Across five types of clinical applications with 36 tasks spanning multimodal disease classification, image-text retrieval, view classification, chamber segmentation, and landmark detection, EchoVLM achieves state-of-the-art performance (86.5% AUC in zero-shot disease classification and 95.1% accuracy in view classification). We demonstrate that clinically grounded multimodal pretraining yields transferable visual representations and establish EchoVLM as a foundation model for end-to-end echocardiography interpretation. We will release EchoGround-MIMIC and the data curation code, enabling reproducibility and further research in multimodal echocardiography interpretation.
Towards Easy and Realistic Network Infrastructure Testing for Large-scale Machine Learning
Apr 29, 2025


Abstract:This paper lays the foundation for Genie, a testing framework that captures the impact of real hardware network behavior on ML workload performance, without requiring expensive GPUs. Genie uses CPU-initiated traffic over a hardware testbed to emulate GPU to GPU communication, and adapts the ASTRA-sim simulator to model interaction between the network and the ML workload.
Fake It Till You Make It: Using Synthetic Data and Domain Knowledge for Improved Text-Based Learning for LGE Detection
Feb 18, 2025



Abstract:Detection of hyperenhancement from cardiac LGE MRI images is a complex task requiring significant clinical expertise. Although deep learning-based models have shown promising results for the task, they require large amounts of data with fine-grained annotations. Clinical reports generated for cardiac MR studies contain rich, clinically relevant information, including the location, extent and etiology of any scars present. Although recently developed CLIP-based training enables pretraining models with image-text pairs, it requires large amounts of data and further finetuning strategies on downstream tasks. In this study, we use various strategies rooted in domain knowledge to train a model for LGE detection solely using text from clinical reports, on a relatively small clinical cohort of 965 patients. We improve performance through the use of synthetic data augmentation, by systematically creating scar images and associated text. In addition, we standardize the orientation of the images in an anatomy-informed way to enable better alignment of spatial and text features. We also use a captioning loss to enable fine-grained supervision and explore the effect of pretraining of the vision encoder on performance. Finally, ablation studies are carried out to elucidate the contributions of each design component to the overall performance of the model.
EchoApex: A General-Purpose Vision Foundation Model for Echocardiography
Oct 14, 2024



Abstract:Quantitative evaluation of echocardiography is essential for precise assessment of cardiac condition, monitoring disease progression, and guiding treatment decisions. The diverse nature of echo images, including variations in probe types, manufacturers, and pathologies, poses challenges for developing artificial intelligent models that can generalize across different clinical practice. We introduce EchoApex, the first general-purpose vision foundation model echocardiography with applications on a variety of clinical practice. Leveraging self-supervised learning, EchoApex is pretrained on over 20 million echo images from 11 clinical centres. By incorporating task-specific decoders and adapter modules, we demonstrate the effectiveness of EchoApex on 4 different kind of clinical applications with 28 sub-tasks, including view classification, interactive structure segmentation, left ventricle hypertrophy detection and automated ejection fraction estimation from view sequences. Compared to state-of-the-art task-specific models, EchoApex attains improved performance with a unified image encoding architecture, demonstrating the benefits of model pretraining at scale with in-domain data. Furthermore, EchoApex illustrates the potential for developing a general-purpose vision foundation model tailored specifically for echocardiography, capable of addressing a diverse range of clinical applications with high efficiency and efficacy.
Towards a vision foundation model for comprehensive assessment of Cardiac MRI
Oct 02, 2024


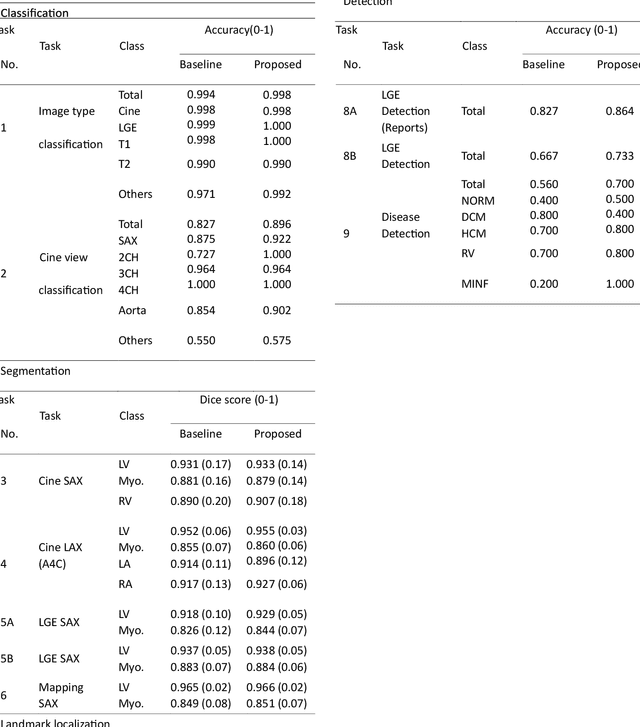
Abstract:Cardiac magnetic resonance imaging (CMR), considered the gold standard for noninvasive cardiac assessment, is a diverse and complex modality requiring a wide variety of image processing tasks for comprehensive assessment of cardiac morphology and function. Advances in deep learning have enabled the development of state-of-the-art (SoTA) models for these tasks. However, model training is challenging due to data and label scarcity, especially in the less common imaging sequences. Moreover, each model is often trained for a specific task, with no connection between related tasks. In this work, we introduce a vision foundation model trained for CMR assessment, that is trained in a self-supervised fashion on 36 million CMR images. We then finetune the model in supervised way for 9 clinical tasks typical to a CMR workflow, across classification, segmentation, landmark localization, and pathology detection. We demonstrate improved accuracy and robustness across all tasks, over a range of available labeled dataset sizes. We also demonstrate improved few-shot learning with fewer labeled samples, a common challenge in medical image analyses. We achieve an out-of-box performance comparable to SoTA for most clinical tasks. The proposed method thus presents a resource-efficient, unified framework for CMR assessment, with the potential to accelerate the development of deep learning-based solutions for image analysis tasks, even with few annotated data available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge