Jay Patel
Diagnosis of diabetic retinopathy using machine learning & deep learning technique
Nov 25, 2024



Abstract:Fundus images are widely used for diagnosing various eye diseases, such as diabetic retinopathy, glaucoma, and age-related macular degeneration. However, manual analysis of fundus images is time-consuming and prone to errors. In this report, we propose a novel method for fundus detection using object detection and machine learning classification techniques. We use a YOLO_V8 to perform object detection on fundus images and locate the regions of interest (ROIs) such as optic disc, optic cup and lesions. We then use machine learning SVM classification algorithms to classify the ROIs into different DR stages based on the presence or absence of pathological signs such as exudates, microaneurysms, and haemorrhages etc. Our method achieves 84% accuracy and efficiency for fundus detection and can be applied for retinal fundus disease triage, especially in remote areas around the world.
* 9 pages, 11 figures, Journal Paper
iDRAMA-Scored-2024: A Dataset of the Scored Social Media Platform from 2020 to 2023
May 16, 2024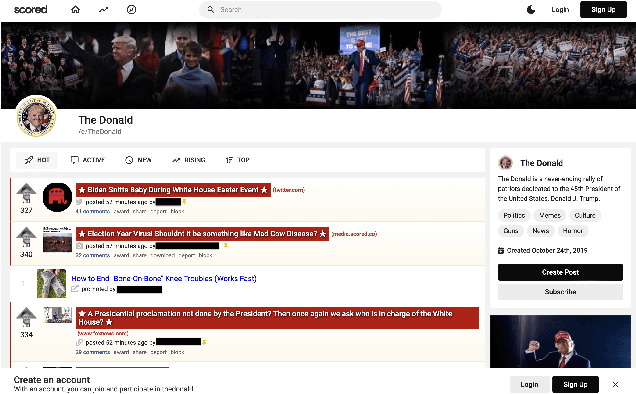
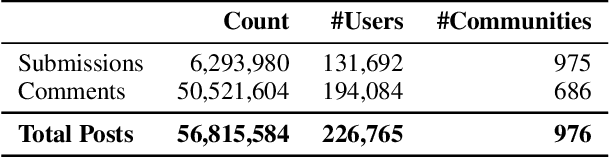
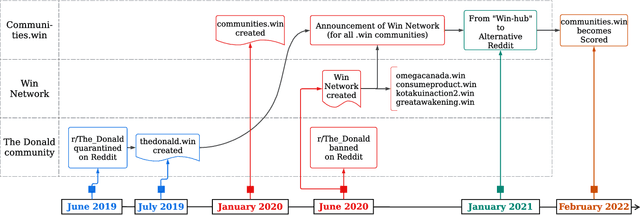

Abstract:Online web communities often face bans for violating platform policies, encouraging their migration to alternative platforms. This migration, however, can result in increased toxicity and unforeseen consequences on the new platform. In recent years, researchers have collected data from many alternative platforms, indicating coordinated efforts leading to offline events, conspiracy movements, hate speech propagation, and harassment. Thus, it becomes crucial to characterize and understand these alternative platforms. To advance research in this direction, we collect and release a large-scale dataset from Scored -- an alternative Reddit platform that sheltered banned fringe communities, for example, c/TheDonald (a prominent right-wing community) and c/GreatAwakening (a conspiratorial community). Over four years, we collected approximately 57M posts from Scored, with at least 58 communities identified as migrating from Reddit and over 950 communities created since the platform's inception. Furthermore, we provide sentence embeddings of all posts in our dataset, generated through a state-of-the-art model, to further advance the field in characterizing the discussions within these communities. We aim to provide these resources to facilitate their investigations without the need for extensive data collection and processing efforts.
Aya Dataset: An Open-Access Collection for Multilingual Instruction Tuning
Feb 09, 2024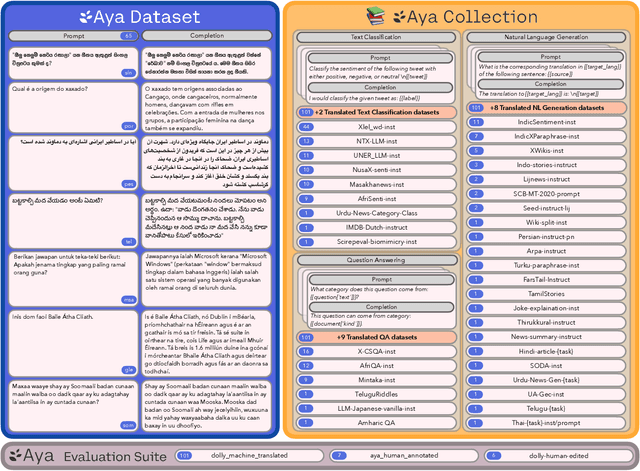
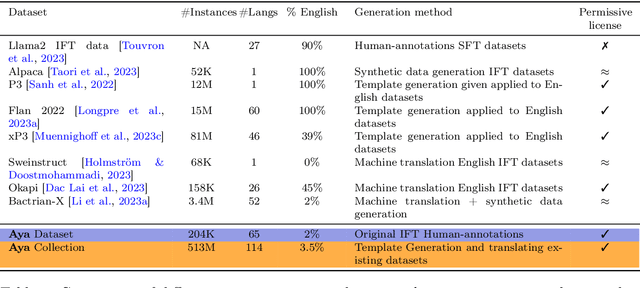
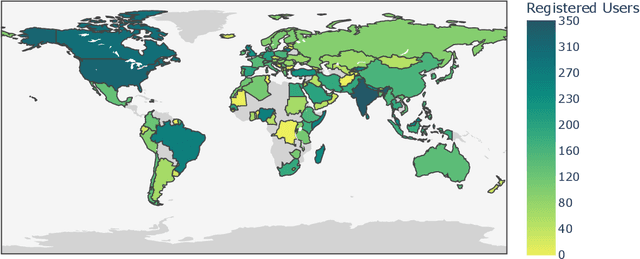
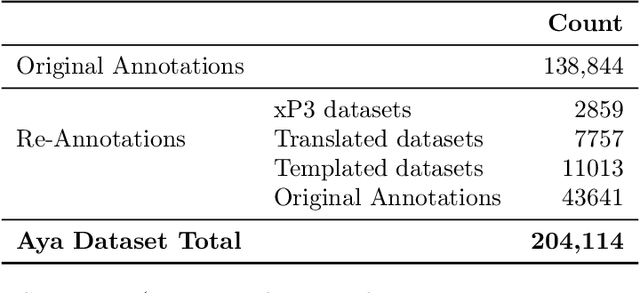
Abstract:Datasets are foundational to many breakthroughs in modern artificial intelligence. Many recent achievements in the space of natural language processing (NLP) can be attributed to the finetuning of pre-trained models on a diverse set of tasks that enables a large language model (LLM) to respond to instructions. Instruction fine-tuning (IFT) requires specifically constructed and annotated datasets. However, existing datasets are almost all in the English language. In this work, our primary goal is to bridge the language gap by building a human-curated instruction-following dataset spanning 65 languages. We worked with fluent speakers of languages from around the world to collect natural instances of instructions and completions. Furthermore, we create the most extensive multilingual collection to date, comprising 513 million instances through templating and translating existing datasets across 114 languages. In total, we contribute four key resources: we develop and open-source the Aya Annotation Platform, the Aya Dataset, the Aya Collection, and the Aya Evaluation Suite. The Aya initiative also serves as a valuable case study in participatory research, involving collaborators from 119 countries. We see this as a valuable framework for future research collaborations that aim to bridge gaps in resources.
From Unstable Contacts to Stable Control: A Deep Learning Paradigm for HD-sEMG in Neurorobotics
Sep 20, 2023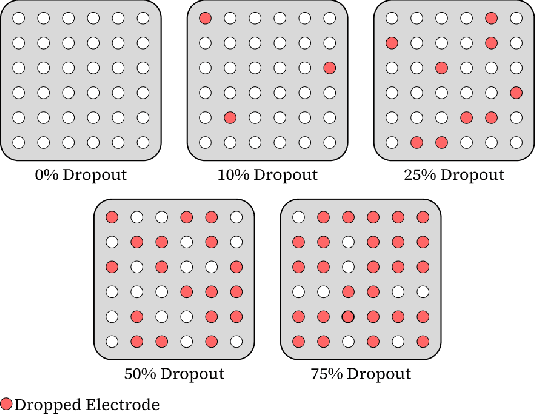
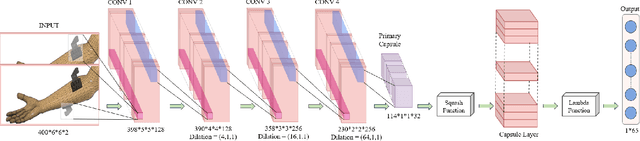
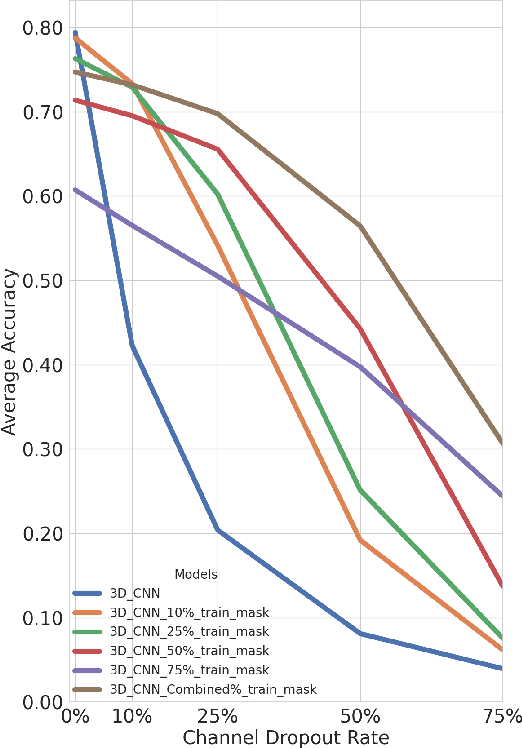
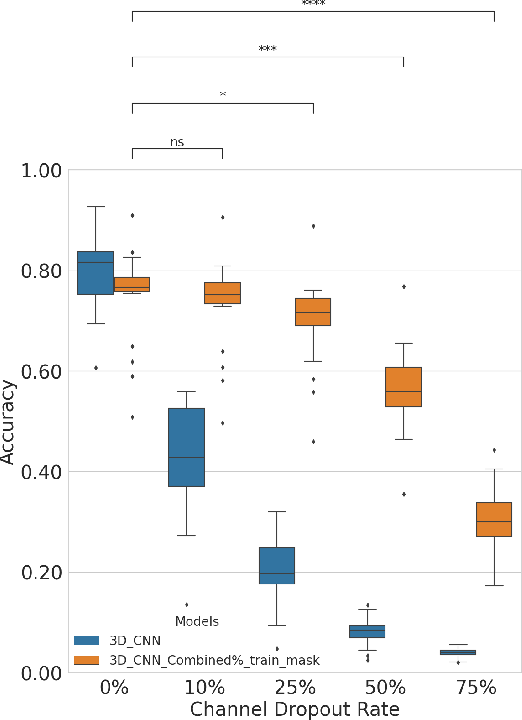
Abstract:In the past decade, there has been significant advancement in designing wearable neural interfaces for controlling neurorobotic systems, particularly bionic limbs. These interfaces function by decoding signals captured non-invasively from the skin's surface. Portable high-density surface electromyography (HD-sEMG) modules combined with deep learning decoding have attracted interest by achieving excellent gesture prediction and myoelectric control of prosthetic systems and neurorobots. However, factors like pixel-shape electrode size and unstable skin contact make HD-sEMG susceptible to pixel electrode drops. The sparse electrode-skin disconnections rooted in issues such as low adhesion, sweating, hair blockage, and skin stretch challenge the reliability and scalability of these modules as the perception unit for neurorobotic systems. This paper proposes a novel deep-learning model providing resiliency for HD-sEMG modules, which can be used in the wearable interfaces of neurorobots. The proposed 3D Dilated Efficient CapsNet model trains on an augmented input space to computationally `force' the network to learn channel dropout variations and thus learn robustness to channel dropout. The proposed framework maintained high performance under a sensor dropout reliability study conducted. Results show conventional models' performance significantly degrades with dropout and is recovered using the proposed architecture and the training paradigm.
AnthroNet: Conditional Generation of Humans via Anthropometrics
Sep 07, 2023Abstract:We present a novel human body model formulated by an extensive set of anthropocentric measurements, which is capable of generating a wide range of human body shapes and poses. The proposed model enables direct modeling of specific human identities through a deep generative architecture, which can produce humans in any arbitrary pose. It is the first of its kind to have been trained end-to-end using only synthetically generated data, which not only provides highly accurate human mesh representations but also allows for precise anthropometry of the body. Moreover, using a highly diverse animation library, we articulated our synthetic humans' body and hands to maximize the diversity of the learnable priors for model training. Our model was trained on a dataset of $100k$ procedurally-generated posed human meshes and their corresponding anthropometric measurements. Our synthetic data generator can be used to generate millions of unique human identities and poses for non-commercial academic research purposes.
Pressmatch: Automated journalist recommendation for media coverage with Nearest Neighbor search
Sep 02, 2023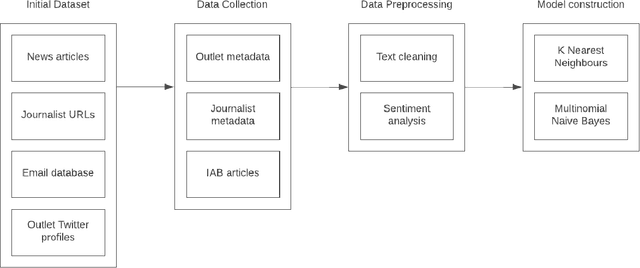
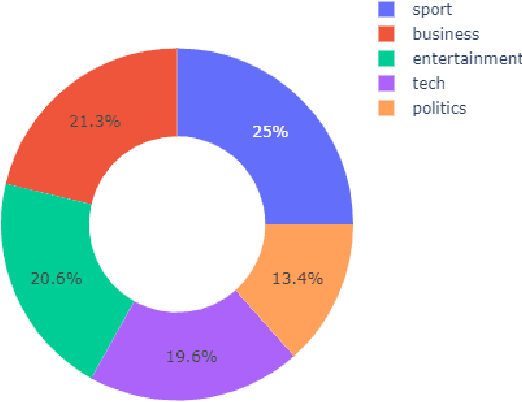
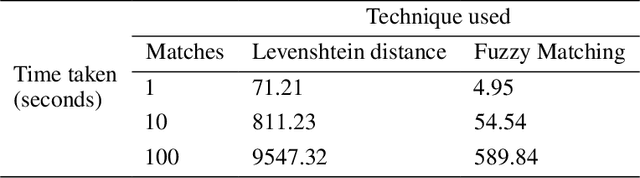
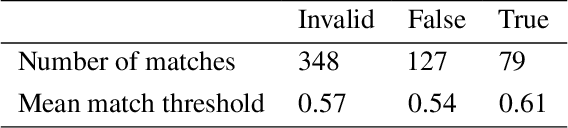
Abstract:Slating a product for release often involves pitching journalists to run stories on your press release. Good media coverage often ensures greater product reach and drives audience engagement for those products. Hence, ensuring that those releases are pitched to the right journalists with relevant interests is crucial, since they receive several pitches daily. Keeping up with journalist beats and curating a media contacts list is often a huge and time-consuming task. This study proposes a model to automate and expedite the process by recommending suitable journalists to run media coverage on the press releases provided by the user.
QU-BraTS: MICCAI BraTS 2020 Challenge on Quantifying Uncertainty in Brain Tumor Segmentation -- Analysis of Ranking Metrics and Benchmarking Results
Dec 19, 2021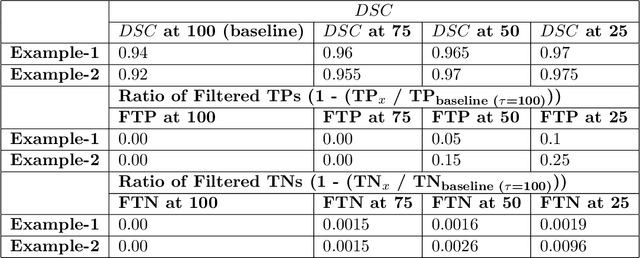
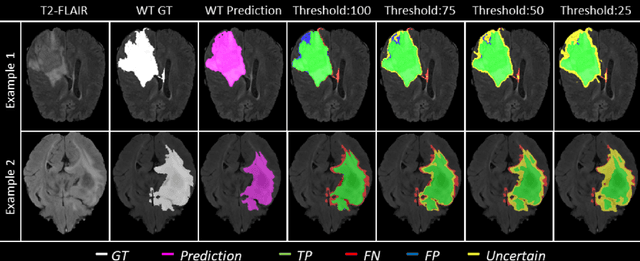
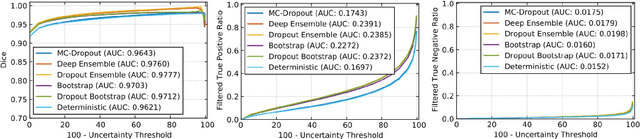
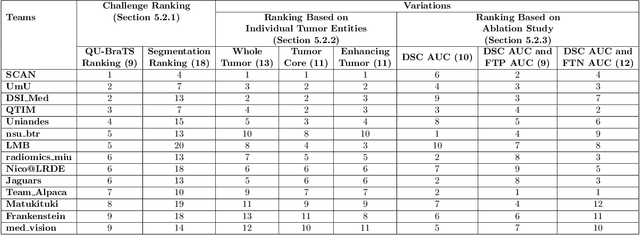
Abstract:Deep learning (DL) models have provided the state-of-the-art performance in a wide variety of medical imaging benchmarking challenges, including the Brain Tumor Segmentation (BraTS) challenges. However, the task of focal pathology multi-compartment segmentation (e.g., tumor and lesion sub-regions) is particularly challenging, and potential errors hinder the translation of DL models into clinical workflows. Quantifying the reliability of DL model predictions in the form of uncertainties, could enable clinical review of the most uncertain regions, thereby building trust and paving the way towards clinical translation. Recently, a number of uncertainty estimation methods have been introduced for DL medical image segmentation tasks. Developing metrics to evaluate and compare the performance of uncertainty measures will assist the end-user in making more informed decisions. In this study, we explore and evaluate a metric developed during the BraTS 2019-2020 task on uncertainty quantification (QU-BraTS), and designed to assess and rank uncertainty estimates for brain tumor multi-compartment segmentation. This metric (1) rewards uncertainty estimates that produce high confidence in correct assertions, and those that assign low confidence levels at incorrect assertions, and (2) penalizes uncertainty measures that lead to a higher percentages of under-confident correct assertions. We further benchmark the segmentation uncertainties generated by 14 independent participating teams of QU-BraTS 2020, all of which also participated in the main BraTS segmentation task. Overall, our findings confirm the importance and complementary value that uncertainty estimates provide to segmentation algorithms, and hence highlight the need for uncertainty quantification in medical image analyses. Our evaluation code is made publicly available at https://github.com/RagMeh11/QU-BraTS.
Addressing catastrophic forgetting for medical domain expansion
Mar 24, 2021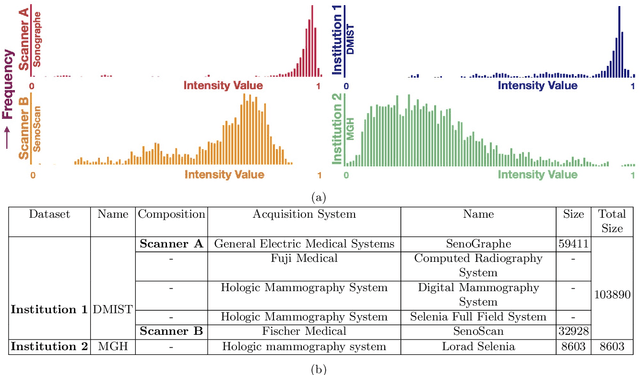
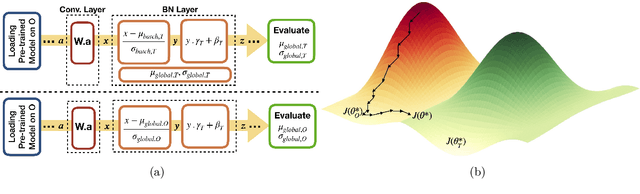
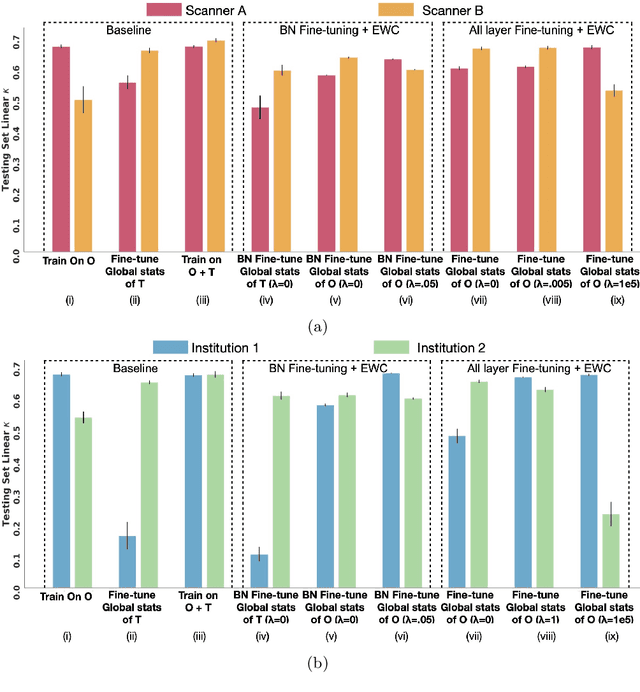
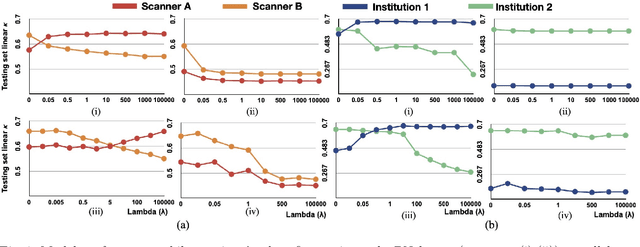
Abstract:Model brittleness is a key concern when deploying deep learning models in real-world medical settings. A model that has high performance at one institution may suffer a significant decline in performance when tested at other institutions. While pooling datasets from multiple institutions and retraining may provide a straightforward solution, it is often infeasible and may compromise patient privacy. An alternative approach is to fine-tune the model on subsequent institutions after training on the original institution. Notably, this approach degrades model performance at the original institution, a phenomenon known as catastrophic forgetting. In this paper, we develop an approach to address catastrophic forget-ting based on elastic weight consolidation combined with modulation of batch normalization statistics under two scenarios: first, for expanding the domain from one imaging system's data to another imaging system's, and second, for expanding the domain from a large multi-institutional dataset to another single institution dataset. We show that our approach outperforms several other state-of-the-art approaches and provide theoretical justification for the efficacy of batch normalization modulation. The results of this study are generally applicable to the deployment of any clinical deep learning model which requires domain expansion.
The unreasonable effectiveness of Batch-Norm statistics in addressing catastrophic forgetting across medical institutions
Nov 16, 2020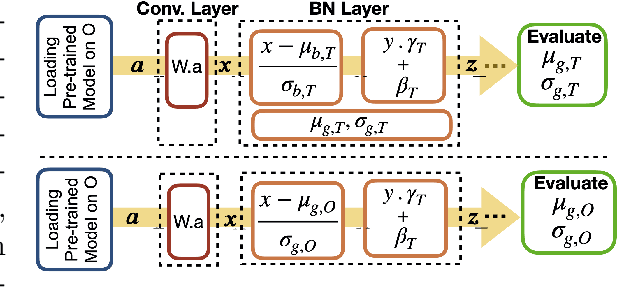
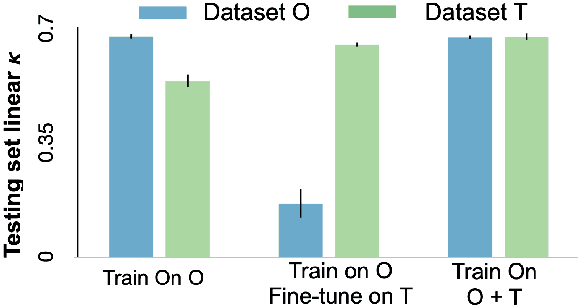
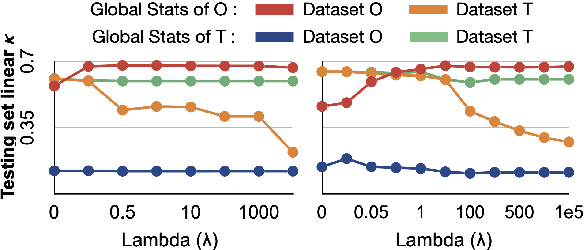
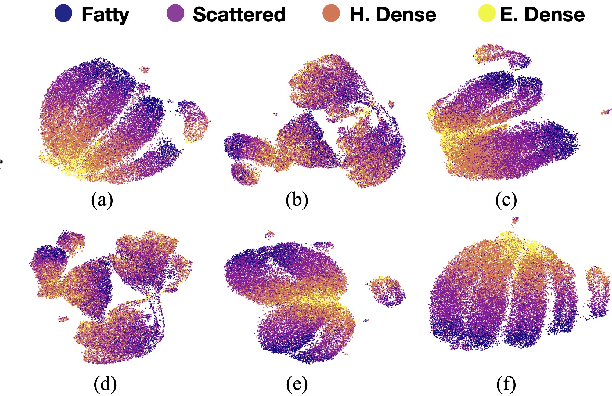
Abstract:Model brittleness is a primary concern when deploying deep learning models in medical settings owing to inter-institution variations, like patient demographics and intra-institution variation, such as multiple scanner types. While simply training on the combined datasets is fraught with data privacy limitations, fine-tuning the model on subsequent institutions after training it on the original institution results in a decrease in performance on the original dataset, a phenomenon called catastrophic forgetting. In this paper, we investigate trade-off between model refinement and retention of previously learned knowledge and subsequently address catastrophic forgetting for the assessment of mammographic breast density. More specifically, we propose a simple yet effective approach, adapting Elastic weight consolidation (EWC) using the global batch normalization (BN) statistics of the original dataset. The results of this study provide guidance for the deployment of clinical deep learning models where continuous learning is needed for domain expansion.
Towards Trainable Saliency Maps in Medical Imaging
Nov 15, 2020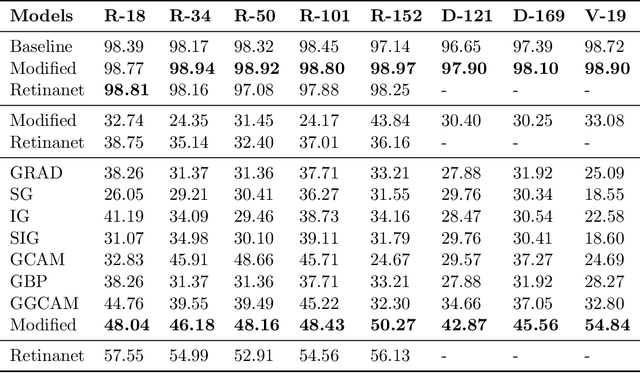
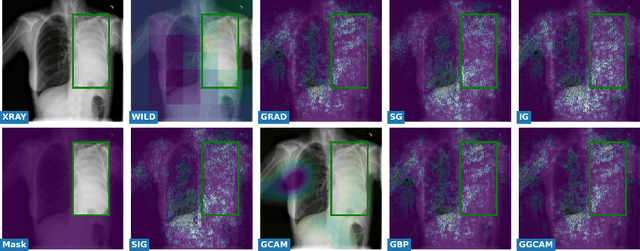
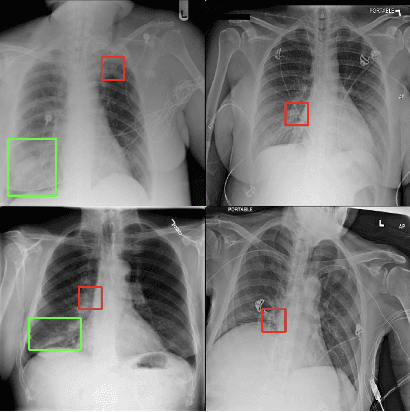
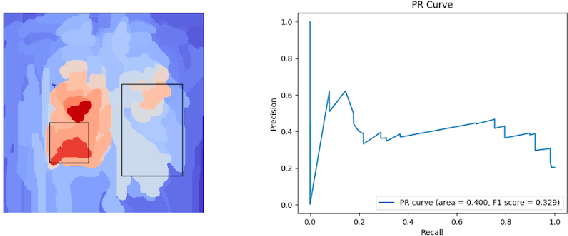
Abstract:While success of Deep Learning (DL) in automated diagnosis can be transformative to the medicinal practice especially for people with little or no access to doctors, its widespread acceptability is severely limited by inherent black-box decision making and unsafe failure modes. While saliency methods attempt to tackle this problem in non-medical contexts, their apriori explanations do not transfer well to medical usecases. With this study we validate a model design element agnostic to both architecture complexity and model task, and show how introducing this element gives an inherently self-explanatory model. We compare our results with state of the art non-trainable saliency maps on RSNA Pneumonia Dataset and demonstrate a much higher localization efficacy using our adopted technique. We also compare, with a fully supervised baseline and provide a reasonable alternative to it's high data labelling overhead. We further investigate the validity of our claims through qualitative evaluation from an expert reader.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge