Piotr Radojewski
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
Apr 25, 2022Abstract:Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data. This is currently addressed by centrally sharing ample, and importantly diverse, data from multiple sites. However, such centralization is challenging to scale (or even not feasible) due to various limitations. Federated ML (FL) provides an alternative to train accurate and generalizable ML models, by only sharing numerical model updates. Here we present findings from the largest FL study to-date, involving data from 71 healthcare institutions across 6 continents, to generate an automatic tumor boundary detector for the rare disease of glioblastoma, utilizing the largest dataset of such patients ever used in the literature (25,256 MRI scans from 6,314 patients). We demonstrate a 33% improvement over a publicly trained model to delineate the surgically targetable tumor, and 23% improvement over the tumor's entire extent. We anticipate our study to: 1) enable more studies in healthcare informed by large and diverse data, ensuring meaningful results for rare diseases and underrepresented populations, 2) facilitate further quantitative analyses for glioblastoma via performance optimization of our consensus model for eventual public release, and 3) demonstrate the effectiveness of FL at such scale and task complexity as a paradigm shift for multi-site collaborations, alleviating the need for data sharing.
QU-BraTS: MICCAI BraTS 2020 Challenge on Quantifying Uncertainty in Brain Tumor Segmentation -- Analysis of Ranking Metrics and Benchmarking Results
Dec 19, 2021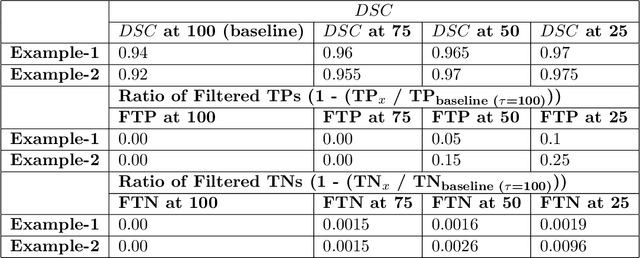
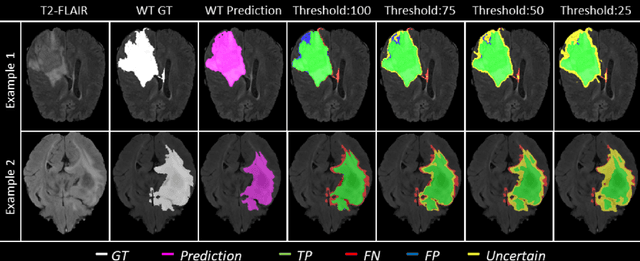
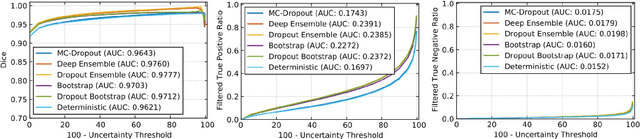
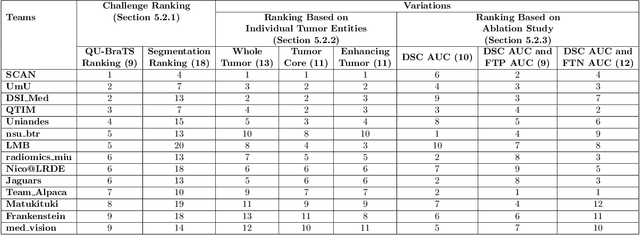
Abstract:Deep learning (DL) models have provided the state-of-the-art performance in a wide variety of medical imaging benchmarking challenges, including the Brain Tumor Segmentation (BraTS) challenges. However, the task of focal pathology multi-compartment segmentation (e.g., tumor and lesion sub-regions) is particularly challenging, and potential errors hinder the translation of DL models into clinical workflows. Quantifying the reliability of DL model predictions in the form of uncertainties, could enable clinical review of the most uncertain regions, thereby building trust and paving the way towards clinical translation. Recently, a number of uncertainty estimation methods have been introduced for DL medical image segmentation tasks. Developing metrics to evaluate and compare the performance of uncertainty measures will assist the end-user in making more informed decisions. In this study, we explore and evaluate a metric developed during the BraTS 2019-2020 task on uncertainty quantification (QU-BraTS), and designed to assess and rank uncertainty estimates for brain tumor multi-compartment segmentation. This metric (1) rewards uncertainty estimates that produce high confidence in correct assertions, and those that assign low confidence levels at incorrect assertions, and (2) penalizes uncertainty measures that lead to a higher percentages of under-confident correct assertions. We further benchmark the segmentation uncertainties generated by 14 independent participating teams of QU-BraTS 2020, all of which also participated in the main BraTS segmentation task. Overall, our findings confirm the importance and complementary value that uncertainty estimates provide to segmentation algorithms, and hence highlight the need for uncertainty quantification in medical image analyses. Our evaluation code is made publicly available at https://github.com/RagMeh11/QU-BraTS.
Uncertainty-driven refinement of tumor-core segmentation using 3D-to-2D networks with label uncertainty
Dec 11, 2020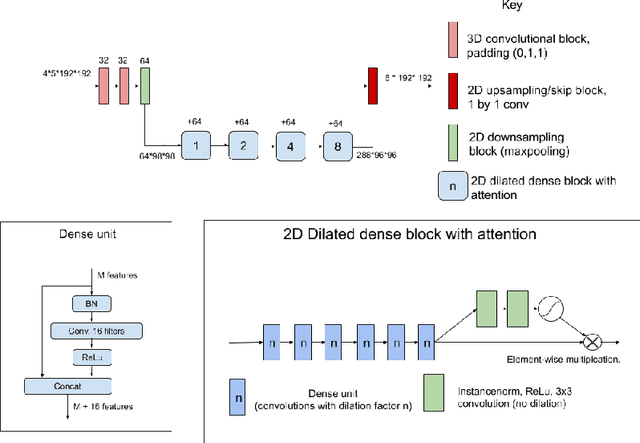
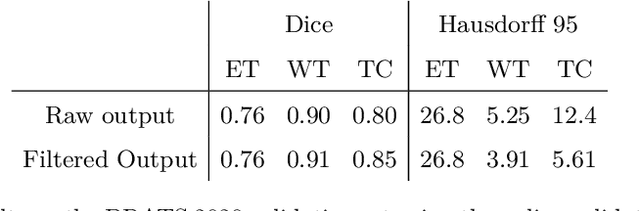
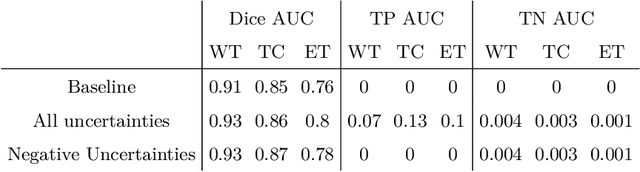
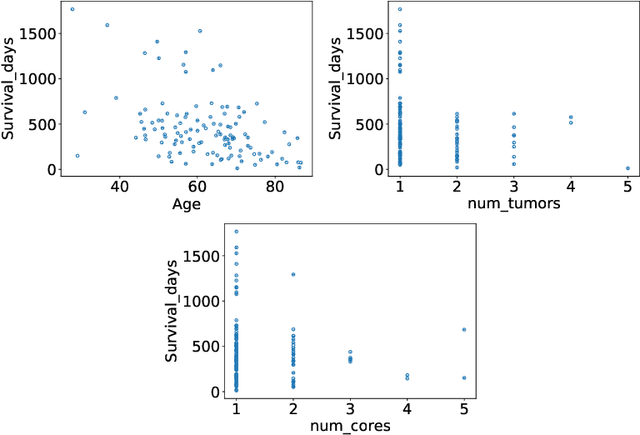
Abstract:The BraTS dataset contains a mixture of high-grade and low-grade gliomas, which have a rather different appearance: previous studies have shown that performance can be improved by separated training on low-grade gliomas (LGGs) and high-grade gliomas (HGGs), but in practice this information is not available at test time to decide which model to use. By contrast with HGGs, LGGs often present no sharp boundary between the tumor core and the surrounding edema, but rather a gradual reduction of tumor-cell density. Utilizing our 3D-to-2D fully convolutional architecture, DeepSCAN, which ranked highly in the 2019 BraTS challenge and was trained using an uncertainty-aware loss, we separate cases into those with a confidently segmented core, and those with a vaguely segmented or missing core. Since by assumption every tumor has a core, we reduce the threshold for classification of core tissue in those cases where the core, as segmented by the classifier, is vaguely defined or missing. We then predict survival of high-grade glioma patients using a fusion of linear regression and random forest classification, based on age, number of distinct tumor components, and number of distinct tumor cores. We present results on the validation dataset of the Multimodal Brain Tumor Segmentation Challenge 2020 (segmentation and uncertainty challenge), and on the testing set, where the method achieved 4th place in Segmentation, 1st place in uncertainty estimation, and 1st place in Survival prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge