James R. Clough
A persistent homology-based topological loss function for multi-class CNN segmentation of cardiac MRI
Aug 21, 2020
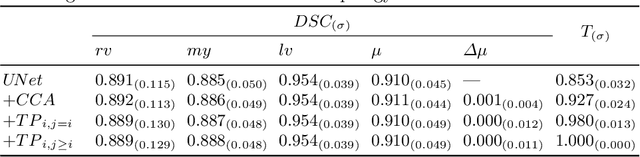
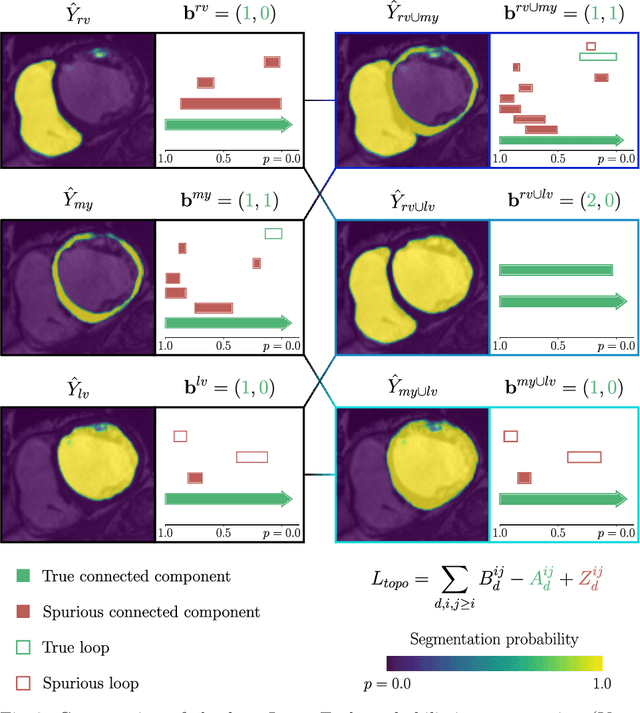
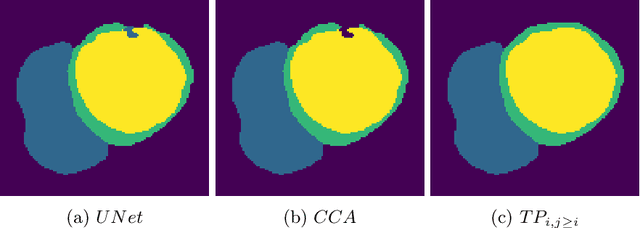
Abstract:With respect to spatial overlap, CNN-based segmentation of short axis cardiovascular magnetic resonance (CMR) images has achieved a level of performance consistent with inter observer variation. However, conventional training procedures frequently depend on pixel-wise loss functions, limiting optimisation with respect to extended or global features. As a result, inferred segmentations can lack spatial coherence, including spurious connected components or holes. Such results are implausible, violating the anticipated topology of image segments, which is frequently known a priori. Addressing this challenge, published work has employed persistent homology, constructing topological loss functions for the evaluation of image segments against an explicit prior. Building a richer description of segmentation topology by considering all possible labels and label pairs, we extend these losses to the task of multi-class segmentation. These topological priors allow us to resolve all topological errors in a subset of 150 examples from the ACDC short axis CMR training data set, without sacrificing overlap performance.
Interpretable Deep Models for Cardiac Resynchronisation Therapy Response Prediction
Jul 09, 2020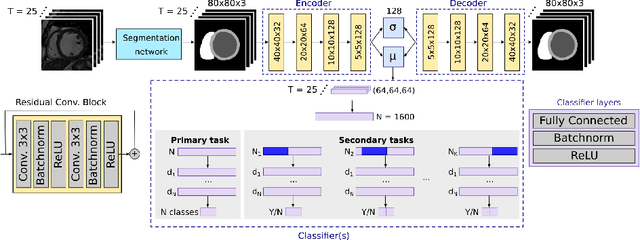
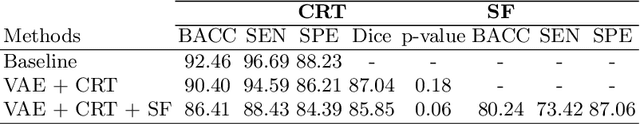
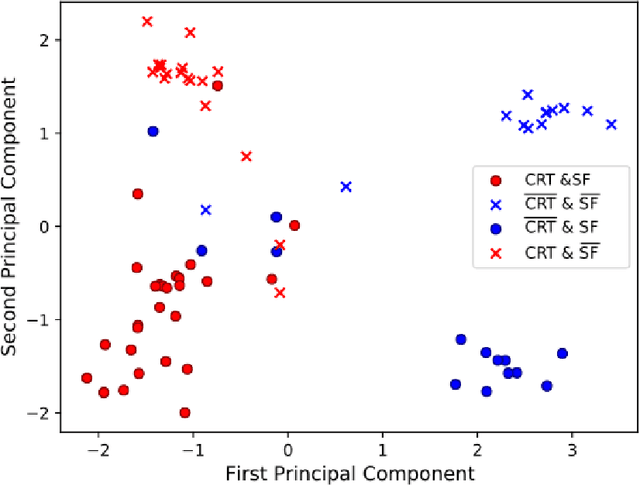

Abstract:Advances in deep learning (DL) have resulted in impressive accuracy in some medical image classification tasks, but often deep models lack interpretability. The ability of these models to explain their decisions is important for fostering clinical trust and facilitating clinical translation. Furthermore, for many problems in medicine there is a wealth of existing clinical knowledge to draw upon, which may be useful in generating explanations, but it is not obvious how this knowledge can be encoded into DL models - most models are learnt either from scratch or using transfer learning from a different domain. In this paper we address both of these issues. We propose a novel DL framework for image-based classification based on a variational autoencoder (VAE). The framework allows prediction of the output of interest from the latent space of the autoencoder, as well as visualisation (in the image domain) of the effects of crossing the decision boundary, thus enhancing the interpretability of the classifier. Our key contribution is that the VAE disentangles the latent space based on `explanations' drawn from existing clinical knowledge. The framework can predict outputs as well as explanations for these outputs, and also raises the possibility of discovering new biomarkers that are separate (or disentangled) from the existing knowledge. We demonstrate our framework on the problem of predicting response of patients with cardiomyopathy to cardiac resynchronization therapy (CRT) from cine cardiac magnetic resonance images. The sensitivity and specificity of the proposed model on the task of CRT response prediction are 88.43% and 84.39% respectively, and we showcase the potential of our model in enhancing understanding of the factors contributing to CRT response.
Deep Learning Based Detection and Correction of Cardiac MR Motion Artefacts During Reconstruction for High-Quality Segmentation
Oct 21, 2019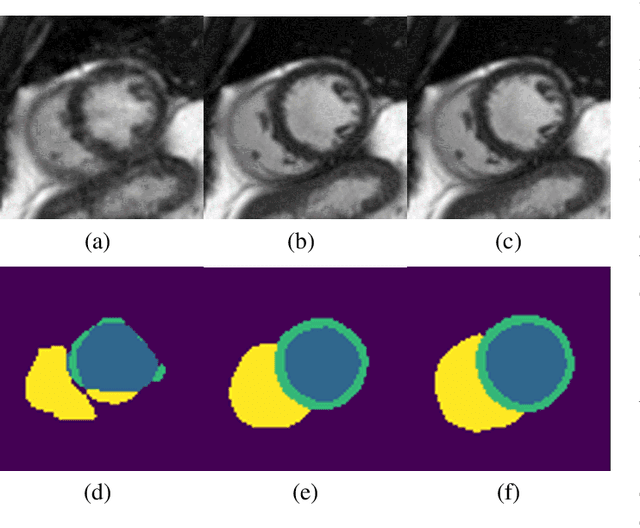
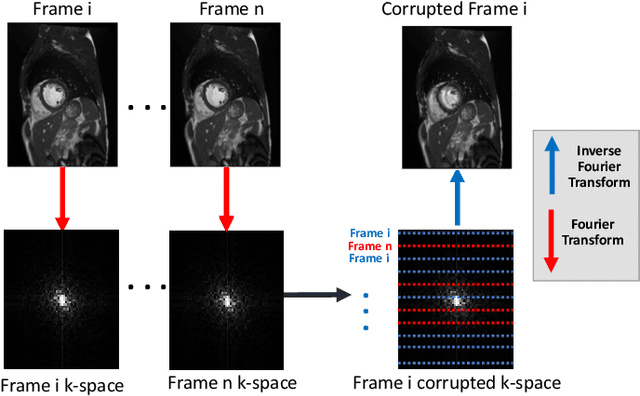
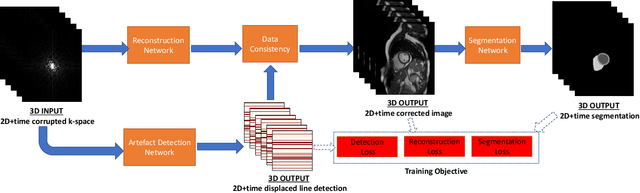
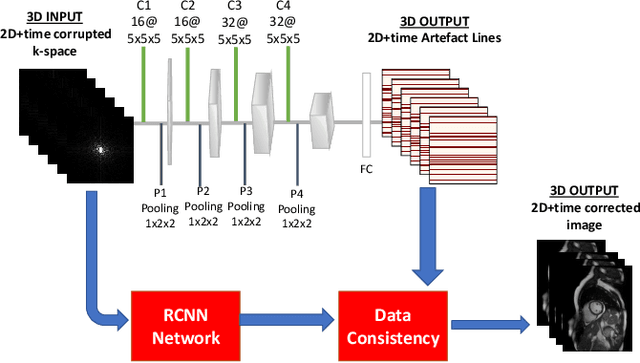
Abstract:Segmenting anatomical structures in medical images has been successfully addressed with deep learning methods for a range of applications. However, this success is heavily dependent on the quality of the image that is being segmented. A commonly neglected point in the medical image analysis community is the vast amount of clinical images that have severe image artefacts due to organ motion, movement of the patient and/or image acquisition related issues. In this paper, we discuss the implications of image motion artefacts on cardiac MR segmentation and compare a variety of approaches for jointly correcting for artefacts and segmenting the cardiac cavity. We propose to use a segmentation network coupled with this in an end-to-end framework. Our training optimises three different tasks: 1) image artefact detection, 2) artefact correction and 3) image segmentation. We train the reconstruction network to automatically correct for motion-related artefacts using synthetically corrupted cardiac MR k-space data and uncorrected reconstructed images. Using a test set of 500 2D+time cine MR acquisitions from the UK Biobank data set, we achieve demonstrably good image quality and high segmentation accuracy in the presence of synthetic motion artefacts. We quantitatively compare our method with a variety of techniques for jointly recovering image quality and performing image segmentation. We showcase better performance compared to state-of-the-art image correction techniques. Moreover, our method preserves the quality of uncorrupted images and therefore can be utilised as a global image reconstruction algorithm.
A Topological Loss Function for Deep-Learning based Image Segmentation using Persistent Homology
Oct 04, 2019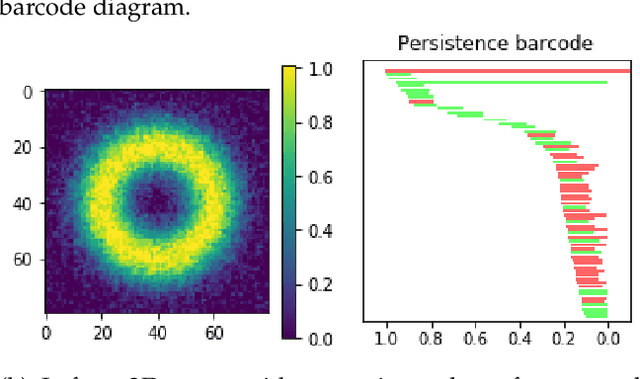
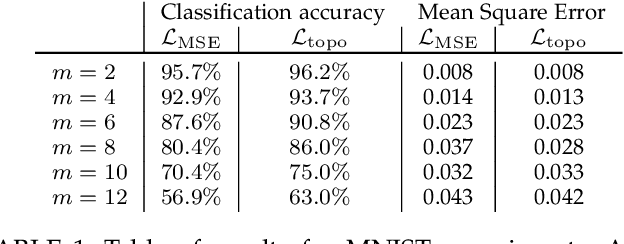
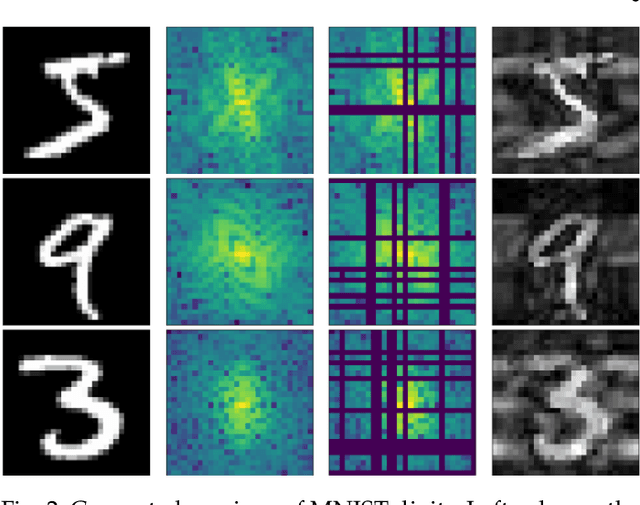
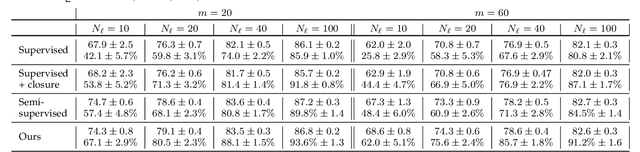
Abstract:We introduce a method for training neural networks to perform image or volume segmentation in which prior knowledge about the topology of the segmented object can be explicitly provided and then incorporated into the training process. By using the differentiable properties of persistent homology, a concept used in topological data analysis, we can specify the desired topology of segmented objects in terms of their Betti numbers and then drive the proposed segmentations to contain the specified topological features. Importantly this process does not require any ground-truth labels, just prior knowledge of the topology of the structure being segmented. We demonstrate our approach in three experiments. Firstly we create a synthetic task in which handwritten MNIST digits are de-noised, and show that using this kind of topological prior knowledge in the training of the network significantly improves the quality of the de-noised digits. Secondly we perform an experiment in which the task is segmenting the myocardium of the left ventricle from cardiac magnetic resonance images. We show that the incorporation of the prior knowledge of the topology of this anatomy improves the resulting segmentations in terms of both the topological accuracy and the Dice coefficient. Thirdly, we extend the method to 3D volumes and demonstrate its performance on the task of segmenting the placenta from ultrasound data, again showing that incorporating topological priors improves performance on this challenging task. We find that embedding explicit prior knowledge in neural network segmentation tasks is most beneficial when the segmentation task is especially challenging and that it can be used in either a semi-supervised or post-processing context to extract a useful training gradient from images without pixelwise labels.
dAUTOMAP: decomposing AUTOMAP to achieve scalability and enhance performance
Sep 25, 2019
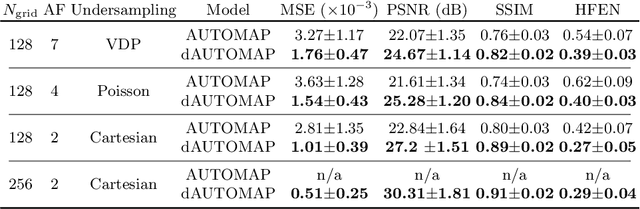
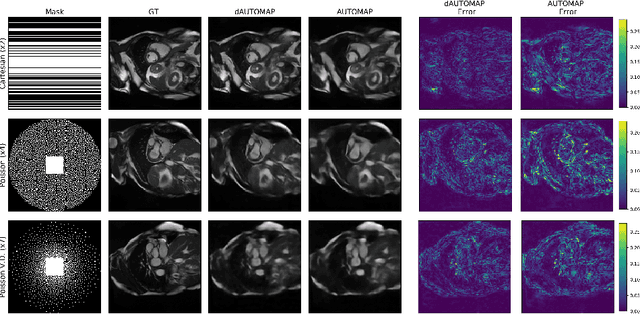

Abstract:AUTOMAP is a promising generalized reconstruction approach, however, it is not scalable and hence the practicality is limited. We present dAUTOMAP, a novel way for decomposing the domain transformation of AUTOMAP, making the model scale linearly. We show dAUTOMAP outperforms AUTOMAP with significantly fewer parameters.
Self-supervised Recurrent Neural Network for 4D Abdominal and In-utero MR Imaging
Aug 28, 2019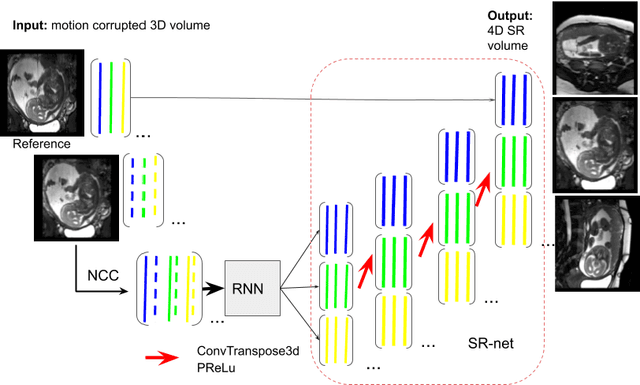



Abstract:Accurately estimating and correcting the motion artifacts are crucial for 3D image reconstruction of the abdominal and in-utero magnetic resonance imaging (MRI). The state-of-art methods are based on slice-to-volume registration (SVR) where multiple 2D image stacks are acquired in three orthogonal orientations. In this work, we present a novel reconstruction pipeline that only needs one orientation of 2D MRI scans and can reconstruct the full high-resolution image without masking or registration steps. The framework consists of two main stages: the respiratory motion estimation using a self-supervised recurrent neural network, which learns the respiratory signals that are naturally embedded in the asymmetry relationship of the neighborhood slices and cluster them according to a respiratory state. Then, we train a 3D deconvolutional network for super-resolution (SR) reconstruction of the sparsely selected 2D images using integrated reconstruction and total variation loss. We evaluate the classification accuracy on 5 simulated images and compare our results with the SVR method in adult abdominal and in-utero MRI scans. The results show that the proposed pipeline can accurately estimate the respiratory state and reconstruct 4D SR volumes with better or similar performance to the 3D SVR pipeline with less than 20\% sparsely selected slices. The method has great potential to transform the 4D abdominal and in-utero MRI in clinical practice.
Topology-preserving augmentation for CNN-based segmentation of congenital heart defects from 3D paediatric CMR
Aug 23, 2019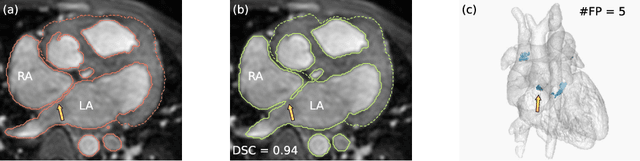
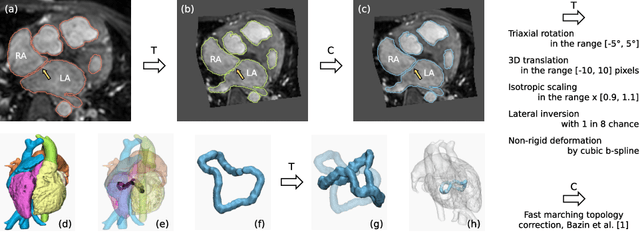
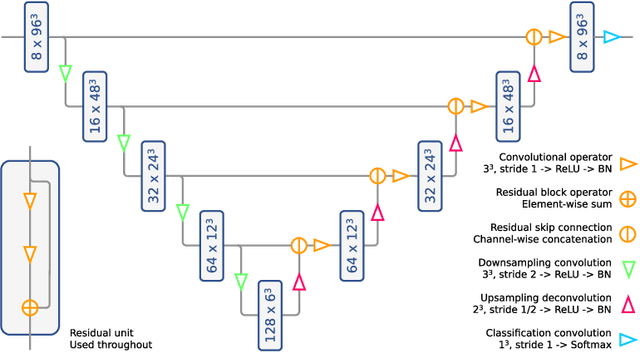
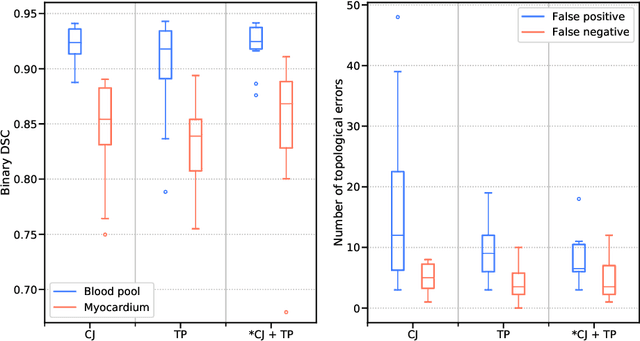
Abstract:Patient-specific 3D printing of congenital heart anatomy demands an accurate segmentation of the thin tissue interfaces which characterise these diagnoses. Even when a label set has a high spatial overlap with the ground truth, inaccurate delineation of these interfaces can result in topological errors. These compromise the clinical utility of such models due to the anomalous appearance of defects. CNNs have achieved state-of-the-art performance in segmentation tasks. Whilst data augmentation has often played an important role, we show that conventional image resampling schemes used therein can introduce topological changes in the ground truth labelling of augmented samples. We present a novel pipeline to correct for these changes, using a fast-marching algorithm to enforce the topology of the ground truth labels within their augmented representations. In so doing, we invoke the idea of cardiac contiguous topology to describe an arbitrary combination of congenital heart defects and develop an associated, clinically meaningful metric to measure the topological correctness of segmentations. In a series of five-fold cross-validations, we demonstrate the performance gain produced by this pipeline and the relevance of topological considerations to the segmentation of congenital heart defects. We speculate as to the applicability of this approach to any segmentation task involving morphologically complex targets.
Assessing the Impact of Blood Pressure on Cardiac Function Using Interpretable Biomarkers and Variational Autoencoders
Aug 13, 2019

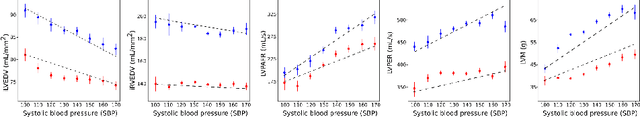

Abstract:Maintaining good cardiac function for as long as possible is a major concern for healthcare systems worldwide and there is much interest in learning more about the impact of different risk factors on cardiac health. The aim of this study is to analyze the impact of systolic blood pressure (SBP) on cardiac function while preserving the interpretability of the model using known clinical biomarkers in a large cohort of the UK Biobank population. We propose a novel framework that combines deep learning based estimation of interpretable clinical biomarkers from cardiac cine MR data with a variational autoencoder (VAE). The VAE architecture integrates a regression loss in the latent space, which enables the progression of cardiac health with SBP to be learnt. Results on 3,600 subjects from the UK Biobank show that the proposed model allows us to gain important insight into the deterioration of cardiac function with increasing SBP, identify key interpretable factors involved in this process, and lastly exploit the model to understand patterns of positive and adverse adaptation of cardiac function.
Global and Local Interpretability for Cardiac MRI Classification
Jun 14, 2019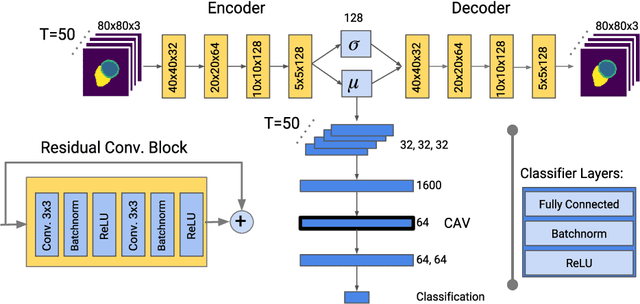



Abstract:Deep learning methods for classifying medical images have demonstrated impressive accuracy in a wide range of tasks but often these models are hard to interpret, limiting their applicability in clinical practice. In this work we introduce a convolutional neural network model for identifying disease in temporal sequences of cardiac MR segmentations which is interpretable in terms of clinically familiar measurements. The model is based around a variational autoencoder, reducing the input into a low-dimensional latent space in which classification occurs. We then use the recently developed `concept activation vector' technique to associate concepts which are diagnostically meaningful (eg. clinical biomarkers such as `low left-ventricular ejection fraction') to certain vectors in the latent space. These concepts are then qualitatively inspected by observing the change in the image domain resulting from interpolations in the latent space in the direction of these vectors. As a result, when the model classifies images it is also capable of providing naturally interpretable concepts relevant to that classification and demonstrating the meaning of those concepts in the image domain. Our approach is demonstrated on the UK Biobank cardiac MRI dataset where we detect the presence of coronary artery disease.
Mechanically Powered Motion Imaging Phantoms: Proof of Concept
May 17, 2019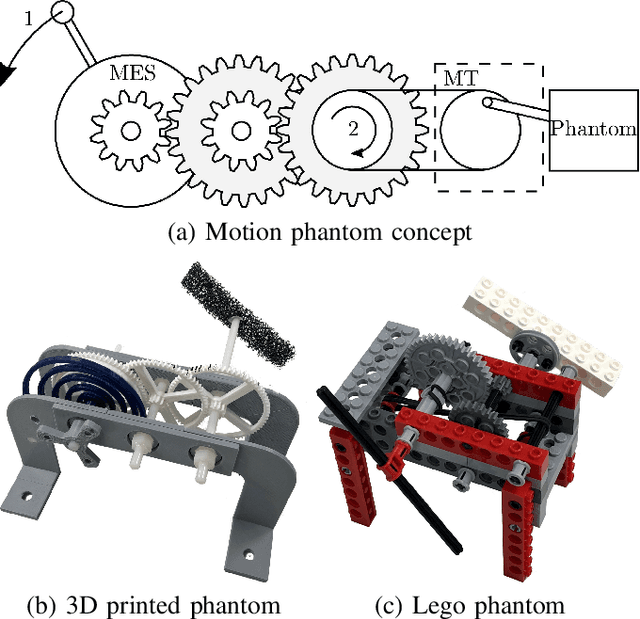
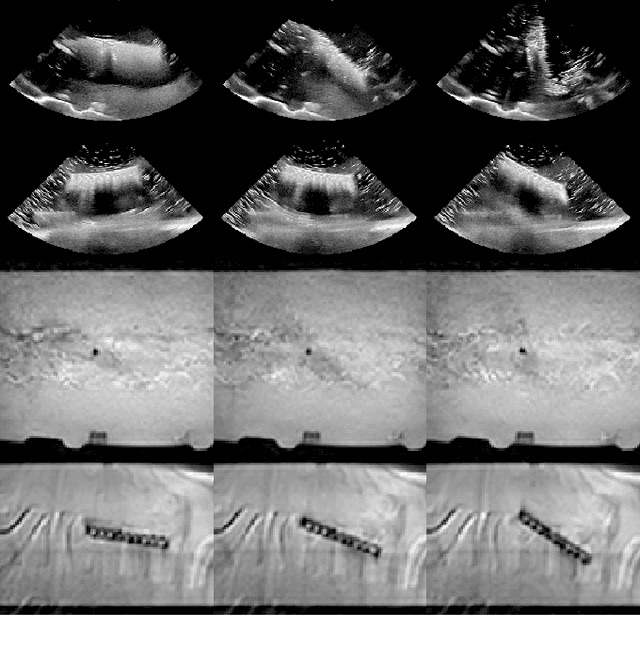
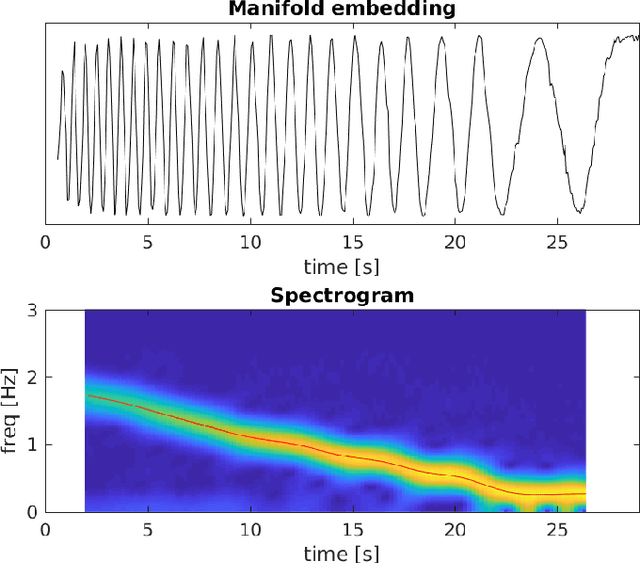
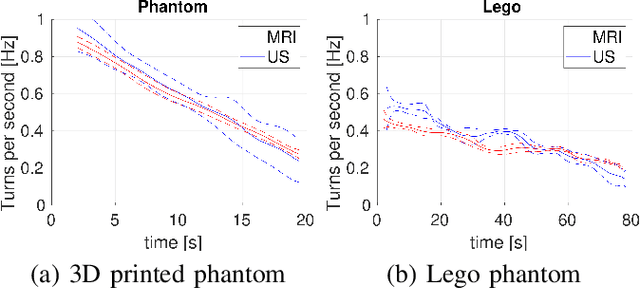
Abstract:Motion imaging phantoms are expensive, bulky and difficult to transport and set-up. The purpose of this paper is to demonstrate a simple approach to the design of multi-modality motion imaging phantoms that use mechanically stored energy to produce motion. We propose two phantom designs that use mainsprings and elastic bands to store energy. A rectangular piece was attached to an axle at the end of the transmission chain of each phantom, and underwent a rotary motion upon release of the mechanical motor. The phantoms were imaged with MRI and US, and the image sequences were embedded in a 1D non linear manifold (Laplacian Eigenmap) and the spectrogram of the embedding was used to derive the angular velocity over time. The derived velocities were consistent and reproducible within a small error. The proposed motion phantom concept showed great potential for the construction of simple and affordable motion phantoms
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge