Nicolas Toussaint
Comparative validation of surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation in endoscopy: Results of the PhaKIR 2024 challenge
Jul 22, 2025Abstract:Reliable recognition and localization of surgical instruments in endoscopic video recordings are foundational for a wide range of applications in computer- and robot-assisted minimally invasive surgery (RAMIS), including surgical training, skill assessment, and autonomous assistance. However, robust performance under real-world conditions remains a significant challenge. Incorporating surgical context - such as the current procedural phase - has emerged as a promising strategy to improve robustness and interpretability. To address these challenges, we organized the Surgical Procedure Phase, Keypoint, and Instrument Recognition (PhaKIR) sub-challenge as part of the Endoscopic Vision (EndoVis) challenge at MICCAI 2024. We introduced a novel, multi-center dataset comprising thirteen full-length laparoscopic cholecystectomy videos collected from three distinct medical institutions, with unified annotations for three interrelated tasks: surgical phase recognition, instrument keypoint estimation, and instrument instance segmentation. Unlike existing datasets, ours enables joint investigation of instrument localization and procedural context within the same data while supporting the integration of temporal information across entire procedures. We report results and findings in accordance with the BIAS guidelines for biomedical image analysis challenges. The PhaKIR sub-challenge advances the field by providing a unique benchmark for developing temporally aware, context-driven methods in RAMIS and offers a high-quality resource to support future research in surgical scene understanding.
Screen Tracking for Clinical Translation of Live Ultrasound Image Analysis Methods
Jul 13, 2020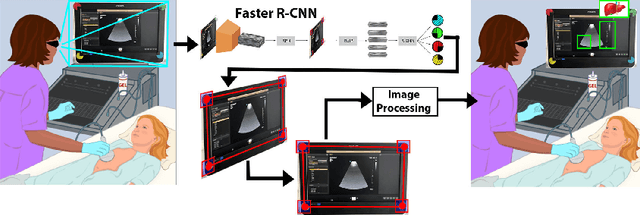
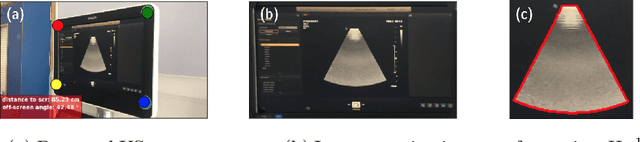
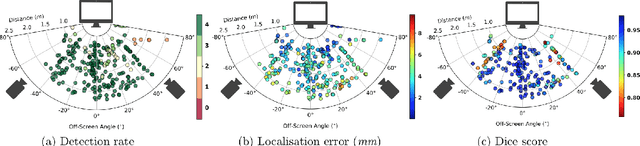
Abstract:Ultrasound (US) imaging is one of the most commonly used non-invasive imaging techniques. However, US image acquisition requires simultaneous guidance of the transducer and interpretation of images, which is a highly challenging task that requires years of training. Despite many recent developments in intra-examination US image analysis, the results are not easy to translate to a clinical setting. We propose a generic framework to extract the US images and superimpose the results of an analysis task, without any need for physical connection or alteration to the US system. The proposed method captures the US image by tracking the screen with a camera fixed at the sonographer's view point and reformats the captured image to the right aspect ratio, in 87.66 +- 3.73ms on average. It is hypothesized that this would enable to input such retrieved image into an image processing pipeline to extract information that can help improve the examination. This information could eventually be projected back to the sonographer's field of view in real time using, for example, an augmented reality (AR) headset.
Confident Head Circumference Measurement from Ultrasound with Real-time Feedback for Sonographers
Aug 07, 2019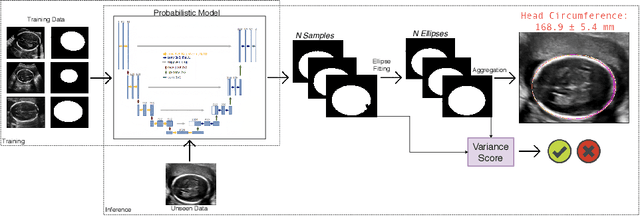
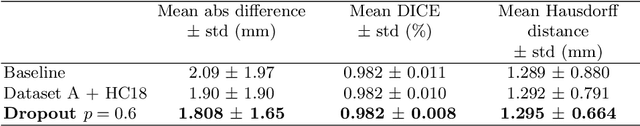
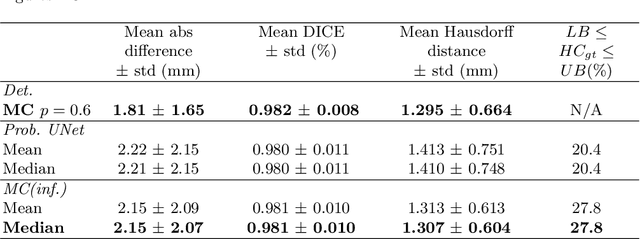
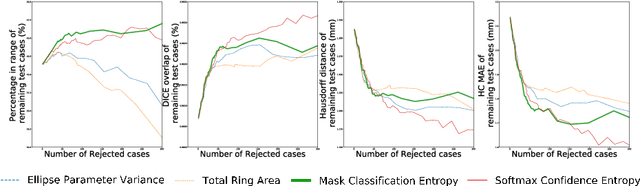
Abstract:Manual estimation of fetal Head Circumference (HC) from Ultrasound (US) is a key biometric for monitoring the healthy development of fetuses. Unfortunately, such measurements are subject to large inter-observer variability, resulting in low early-detection rates of fetal abnormalities. To address this issue, we propose a novel probabilistic Deep Learning approach for real-time automated estimation of fetal HC. This system feeds back statistics on measurement robustness to inform users how confident a deep neural network is in evaluating suitable views acquired during free-hand ultrasound examination. In real-time scenarios, this approach may be exploited to guide operators to scan planes that are as close as possible to the underlying distribution of training images, for the purpose of improving inter-operator consistency. We train on free-hand ultrasound data from over 2000 subjects (2848 training/540 test) and show that our method is able to predict HC measurements within 1.81$\pm$1.65mm deviation from the ground truth, with 50% of the test images fully contained within the predicted confidence margins, and an average of 1.82$\pm$1.78mm deviation from the margin for the remaining cases that are not fully contained.
Mechanically Powered Motion Imaging Phantoms: Proof of Concept
May 17, 2019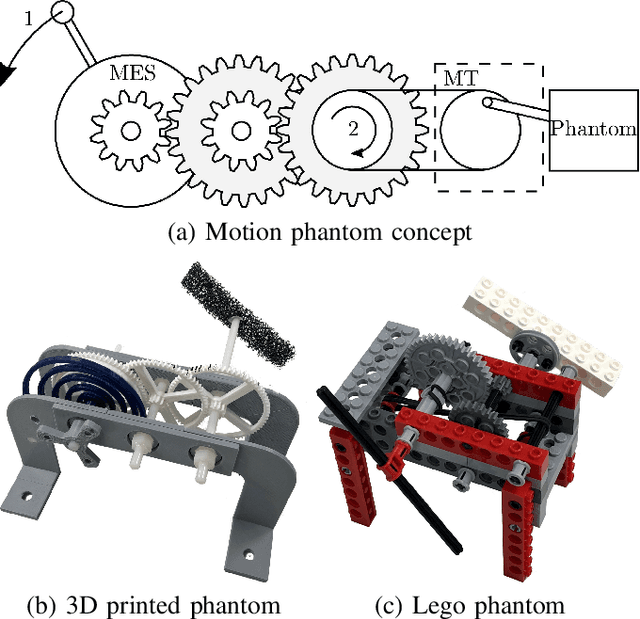
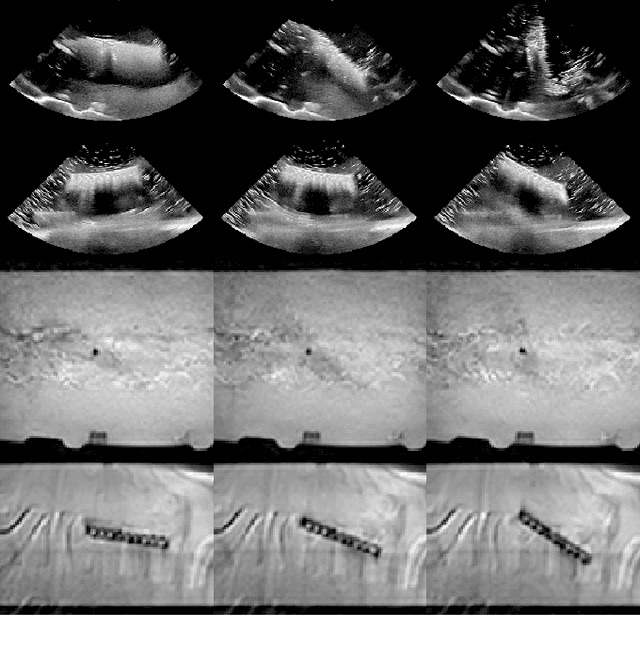
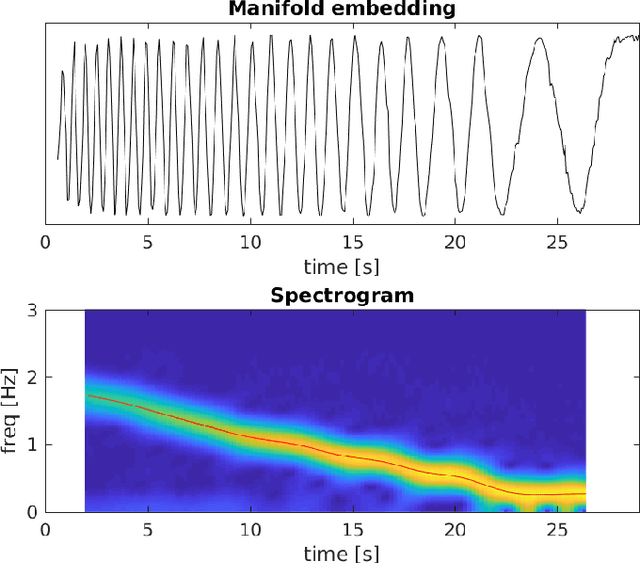
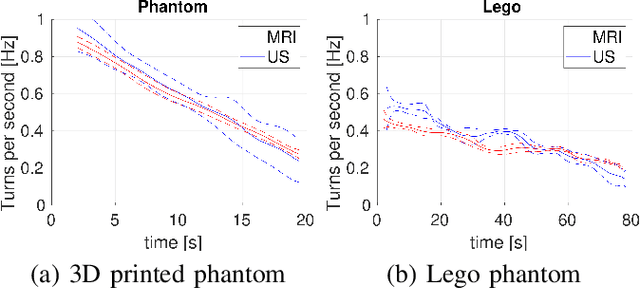
Abstract:Motion imaging phantoms are expensive, bulky and difficult to transport and set-up. The purpose of this paper is to demonstrate a simple approach to the design of multi-modality motion imaging phantoms that use mechanically stored energy to produce motion. We propose two phantom designs that use mainsprings and elastic bands to store energy. A rectangular piece was attached to an axle at the end of the transmission chain of each phantom, and underwent a rotary motion upon release of the mechanical motor. The phantoms were imaged with MRI and US, and the image sequences were embedded in a 1D non linear manifold (Laplacian Eigenmap) and the spectrogram of the embedding was used to derive the angular velocity over time. The derived velocities were consistent and reproducible within a small error. The proposed motion phantom concept showed great potential for the construction of simple and affordable motion phantoms
Robotic-assisted Ultrasound for Fetal Imaging: Evolution from Single-arm to Dual-arm System
Apr 10, 2019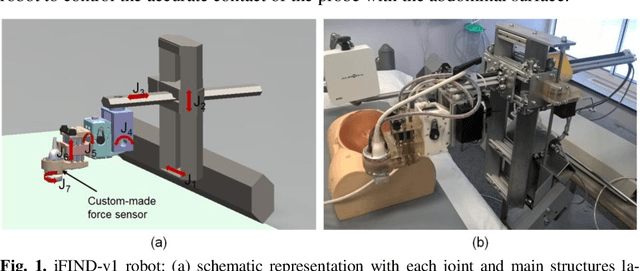
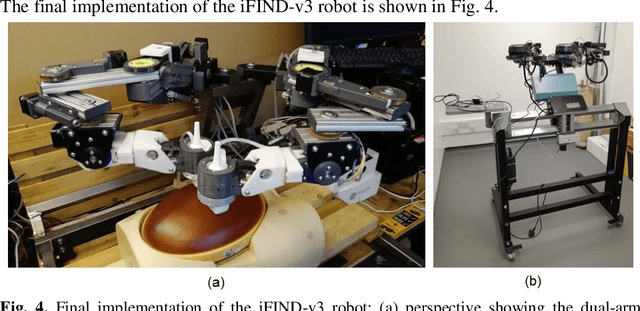
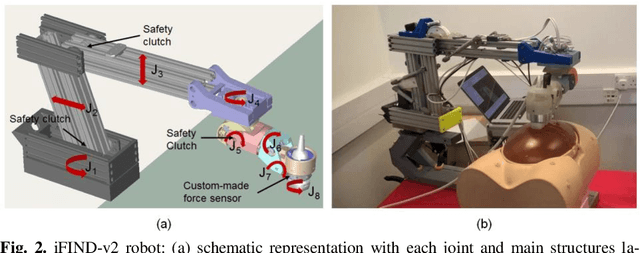
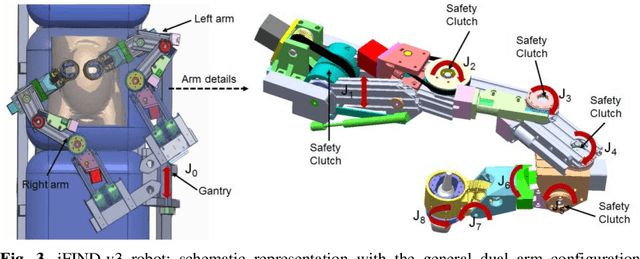

Abstract:The development of robotic-assisted extracorporeal ultrasound systems has a long history and a number of projects have been proposed since the 1990s focusing on different technical aspects. These aim to resolve the deficiencies of on-site manual manipulation of hand-held ultrasound probes. This paper presents the recent ongoing developments of a series of bespoke robotic systems, including both single-arm and dual-arm versions, for a project known as intelligent Fetal Imaging and Diagnosis (iFIND). After a brief review of the development history of the extracorporeal ultrasound robotic system used for fetal and abdominal examinations, the specific aim of the iFIND robots, the design evolution, the implementation details of each version, and the initial clinical feedback of the iFIND robot series are presented. Based on the preliminary testing of these newly-proposed robots on 42 volunteers, the successful and re-liable working of the mechatronic systems were validated. Analysis of a participant questionnaire indicates a comfortable scanning experience for the volunteers and a good acceptance rate to being scanned by the robots.
Weakly Supervised Estimation of Shadow Confidence Maps in Ultrasound Imaging
Nov 21, 2018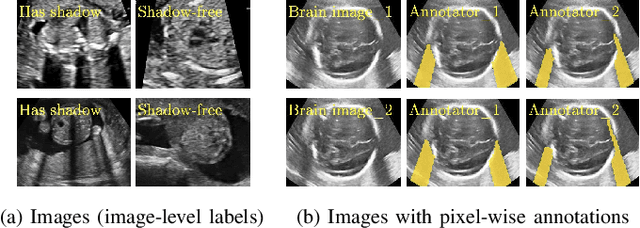
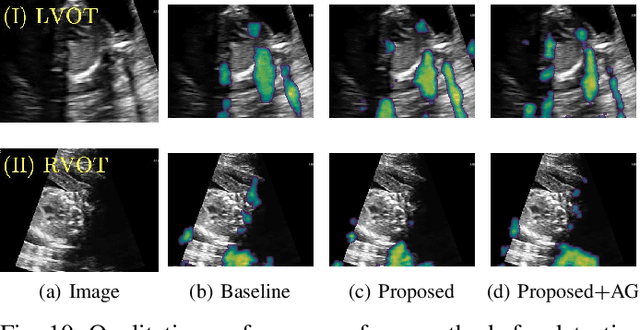

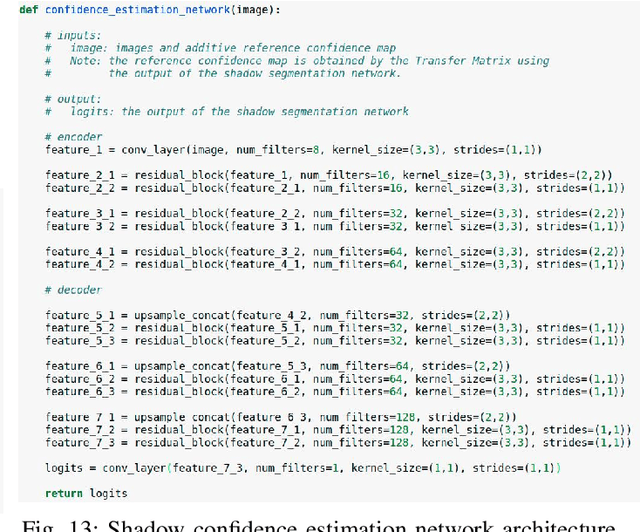
Abstract:Detecting acoustic shadows in ultrasound images is important in many clinical and engineering applications. Real-time feedback of acoustic shadows can guide sonographers to a standardized diagnostic viewing plane with minimal artifacts and can provide additional information for other automatic image analysis algorithms. However, automatically detecting shadow regions is challenging because pixel-wise annotation of acoustic shadows is subjective and time consuming. In this paper we propose a weakly supervised method for automatic confidence estimation of acoustic shadow regions, which is able to generate a dense shadow-focused confidence map. During training, a multi-task module for shadow segmentation is built to learn general shadow features according based image-level annotations as well as a small number of coarse pixel-wise shadow annotations. A transfer function is then established to extend the binary shadow segmentation to a reference confidence map. In addition, a confidence estimation network is proposed to learn the mapping between input images and the reference confidence maps. This confidence estimation network is able to predict shadow confidence maps directly from input images during inference. We evaluate DICE, soft DICE, recall, precision, mean squared error and inter-class correlation to verify the effectiveness of our method. Our method outperforms the state-of-the-art qualitatively and quantitatively. We further demonstrate the applicability of our method by integrating shadow confidence maps into tasks such as ultrasound image classification, multi-view image fusion and automated biometric measurements.
Weakly Supervised Localisation for Fetal Ultrasound Images
Aug 02, 2018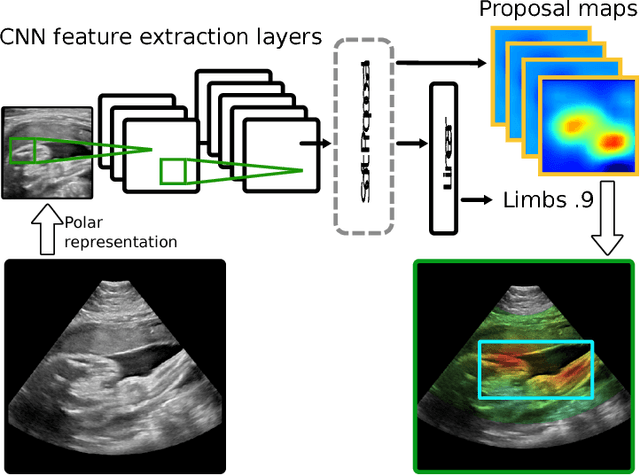
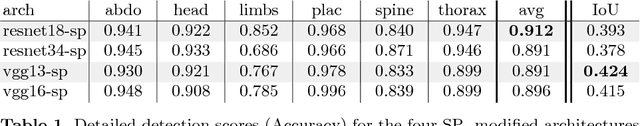
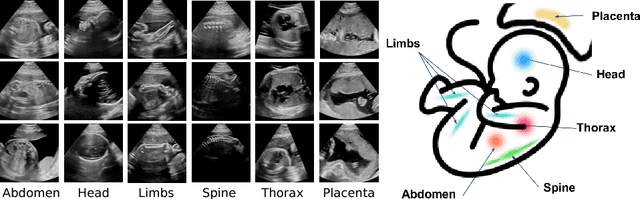
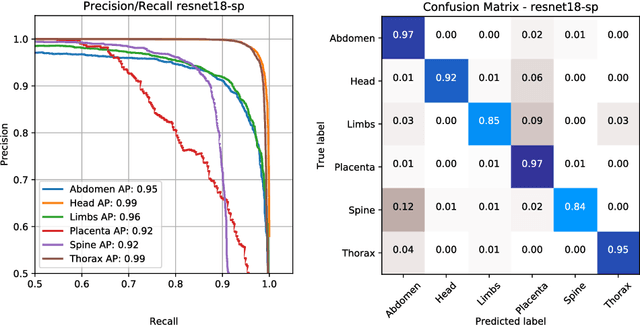
Abstract:This paper addresses the task of detecting and localising fetal anatomical regions in 2D ultrasound images, where only image-level labels are present at training, i.e. without any localisation or segmentation information. We examine the use of convolutional neural network architectures coupled with soft proposal layers. The resulting network simultaneously performs anatomical region detection (classification) and localisation tasks. We generate a proposal map describing the attention of the network for a particular class. The network is trained on 85,500 2D fetal Ultrasound images and their associated labels. Labels correspond to six anatomical regions: head, spine, thorax, abdomen, limbs, and placenta. Detection achieves an average accuracy of 90\% on individual regions, and show that the proposal maps correlate well with relevant anatomical structures. This work presents itself as a powerful and essential step towards subsequent tasks such as fetal position and pose estimation, organ-specific segmentation, or image-guided navigation. Code and additional material is available at https://ntoussaint.github.io/fetalnav
EchoFusion: Tracking and Reconstruction of Objects in 4D Freehand Ultrasound Imaging without External Trackers
Jul 19, 2018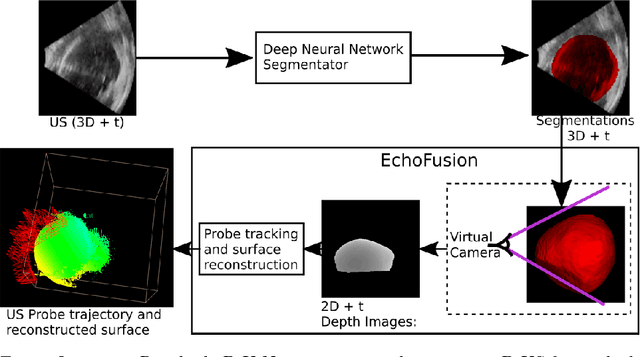
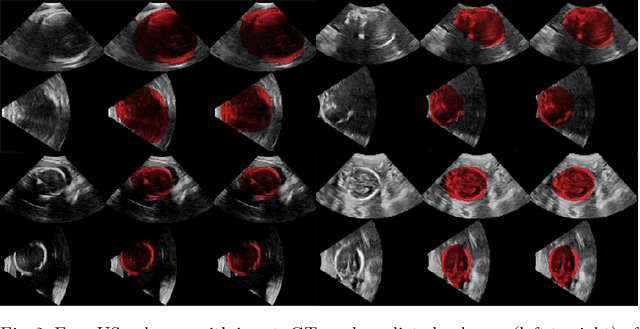


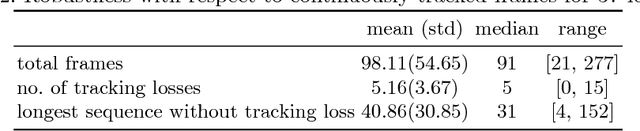
Abstract:Ultrasound (US) is the most widely used fetal imaging technique. However, US images have limited capture range, and suffer from view dependent artefacts such as acoustic shadows. Compounding of overlapping 3D US acquisitions into a high-resolution volume can extend the field of view and remove image artefacts, which is useful for retrospective analysis including population based studies. However, such volume reconstructions require information about relative transformations between probe positions from which the individual volumes were acquired. In prenatal US scans, the fetus can move independently from the mother, making external trackers such as electromagnetic or optical tracking unable to track the motion between probe position and the moving fetus. We provide a novel methodology for image-based tracking and volume reconstruction by combining recent advances in deep learning and simultaneous localisation and mapping (SLAM). Tracking semantics are established through the use of a Residual 3D U-Net and the output is fed to the SLAM algorithm. As a proof of concept, experiments are conducted on US volumes taken from a whole body fetal phantom, and from the heads of real fetuses. For the fetal head segmentation, we also introduce a novel weak annotation approach to minimise the required manual effort for ground truth annotation. We evaluate our method qualitatively, and quantitatively with respect to tissue discrimination accuracy and tracking robustness.
Oversegmenting Graphs
Jun 01, 2018



Abstract:We propose a novel method to adapt a graph to image data. The method drives the nodes of the graph towards image features. The adaptation process naturally lends itself to a measure of feature saliency which can then be used to retain meaningful nodes and edges in the graph. From the adapted graph, we propose the computation of a dual graph, which inherits the saliency measure from the adapted graph, and whose edges run along image features hence producing an oversegmenting graph. This dual graph captures the structure of the underlying image, and therefore constitutes a sparse representation of the image features and their topology. The proposed method is computationally efficient and fully parallelisable. We propose two distance measures find image saliency along graph edges, and evaluate its performance on synthetic images and on natural images from publicly available databases. In both cases, the salient-most nodes of the graph achieve average boundary recall over 90%. We also provide a qualitative comparison with two related techniques: superpixel clustering, and variational image meshing, showing potential for a large number of applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge