Benjamin Hou
Text Embedded Swin-UMamba for DeepLesion Segmentation
Aug 08, 2025Abstract:Segmentation of lesions on CT enables automatic measurement for clinical assessment of chronic diseases (e.g., lymphoma). Integrating large language models (LLMs) into the lesion segmentation workflow offers the potential to combine imaging features with descriptions of lesion characteristics from the radiology reports. In this study, we investigate the feasibility of integrating text into the Swin-UMamba architecture for the task of lesion segmentation. The publicly available ULS23 DeepLesion dataset was used along with short-form descriptions of the findings from the reports. On the test dataset, a high Dice Score of 82% and low Hausdorff distance of 6.58 (pixels) was obtained for lesion segmentation. The proposed Text-Swin-UMamba model outperformed prior approaches: 37% improvement over the LLM-driven LanGuideMedSeg model (p < 0.001),and surpassed the purely image-based xLSTM-UNet and nnUNet models by 1.74% and 0.22%, respectively. The dataset and code can be accessed at https://github.com/ruida/LLM-Swin-UMamba
Longitudinal Assessment of Lung Lesion Burden in CT
Apr 09, 2025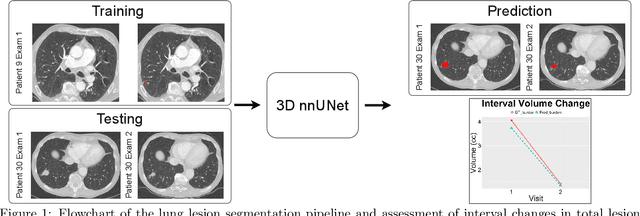
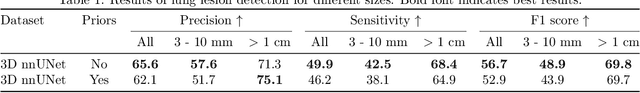

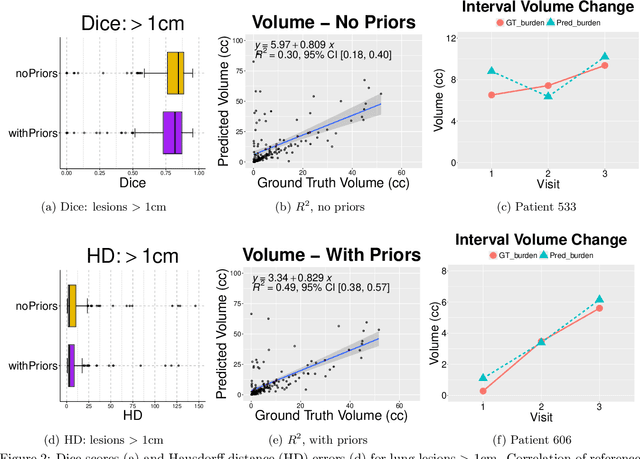
Abstract:In the U.S., lung cancer is the second major cause of death. Early detection of suspicious lung nodules is crucial for patient treatment planning, management, and improving outcomes. Many approaches for lung nodule segmentation and volumetric analysis have been proposed, but few have looked at longitudinal changes in total lung tumor burden. In this work, we trained two 3D models (nnUNet) with and without anatomical priors to automatically segment lung lesions and quantified total lesion burden for each patient. The 3D model without priors significantly outperformed ($p < .001$) the model trained with anatomy priors. For detecting clinically significant lesions $>$ 1cm, a precision of 71.3\%, sensitivity of 68.4\%, and F1-score of 69.8\% was achieved. For segmentation, a Dice score of 77.1 $\pm$ 20.3 and Hausdorff distance error of 11.7 $\pm$ 24.1 mm was obtained. The median lesion burden was 6.4 cc (IQR: 2.1, 18.1) and the median volume difference between manual and automated measurements was 0.02 cc (IQR: -2.8, 1.2). Agreements were also evaluated with linear regression and Bland-Altman plots. The proposed approach can produce a personalized evaluation of the total tumor burden for a patient and facilitate interval change tracking over time.
Efficient 2D CT Foundation Model for Contrast Phase Classification
Jan 23, 2025Abstract:Purpose: The purpose of this study is to harness the efficiency of a 2D foundation model to develop a robust phase classifier that is resilient to domain shifts. Materials and Methods: This retrospective study utilized three public datasets from separate institutions. A 2D foundation model was trained on the DeepLesion dataset (mean age: 51.2, s.d.: 17.6; 2398 males) to generate embeddings from 2D CT slices for downstream contrast phase classification. The classifier was trained on the VinDr Multiphase dataset and externally validated on the WAW-TACE dataset. The 2D model was also compared to three 3D supervised models. Results: On the VinDr dataset (146 male, 63 female, 56 unidentified), the model achieved near-perfect AUROC scores and F1 scores of 99.2%, 94.2%, and 93.1% for non-contrast, arterial, and venous phases, respectively. The `Other' category scored lower (F1: 73.4%) due to combining multiple contrast phases into one class. On the WAW-TACE dataset (mean age: 66.1, s.d.: 10.0; 185 males), the model showed strong performance with AUROCs of 91.0% and 85.6%, and F1 scores of 87.3% and 74.1% for non-contrast and arterial phases. Venous phase performance was lower, with AUROC and F1 scores of 81.7% and 70.2% respectively, due to label mismatches. Compared to 3D supervised models, the approach trained faster, performed as well or better, and showed greater robustness to domain shifts. Conclusion: The robustness of the 2D Foundation model may be potentially useful for automation of hanging protocols and data orchestration for clinical deployment of AI algorithms.
Demystifying Large Language Models for Medicine: A Primer
Oct 24, 2024



Abstract:Large language models (LLMs) represent a transformative class of AI tools capable of revolutionizing various aspects of healthcare by generating human-like responses across diverse contexts and adapting to novel tasks following human instructions. Their potential application spans a broad range of medical tasks, such as clinical documentation, matching patients to clinical trials, and answering medical questions. In this primer paper, we propose an actionable guideline to help healthcare professionals more efficiently utilize LLMs in their work, along with a set of best practices. This approach consists of several main phases, including formulating the task, choosing LLMs, prompt engineering, fine-tuning, and deployment. We start with the discussion of critical considerations in identifying healthcare tasks that align with the core capabilities of LLMs and selecting models based on the selected task and data, performance requirements, and model interface. We then review the strategies, such as prompt engineering and fine-tuning, to adapt standard LLMs to specialized medical tasks. Deployment considerations, including regulatory compliance, ethical guidelines, and continuous monitoring for fairness and bias, are also discussed. By providing a structured step-by-step methodology, this tutorial aims to equip healthcare professionals with the tools necessary to effectively integrate LLMs into clinical practice, ensuring that these powerful technologies are applied in a safe, reliable, and impactful manner.
Beyond Multiple-Choice Accuracy: Real-World Challenges of Implementing Large Language Models in Healthcare
Oct 24, 2024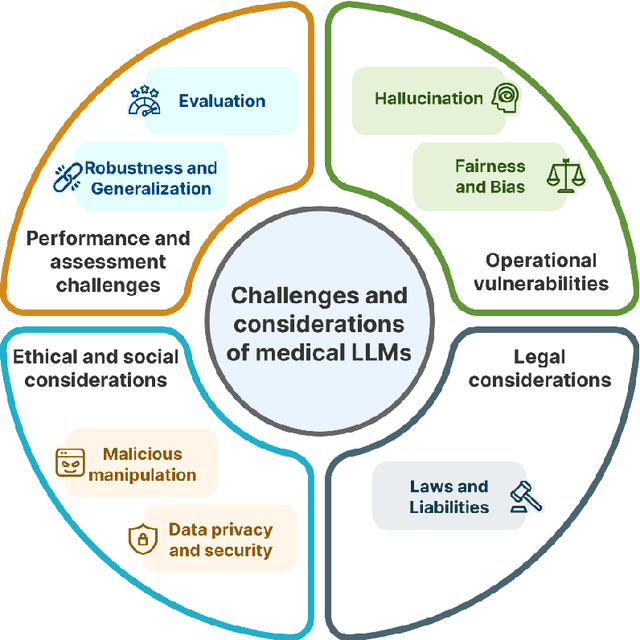
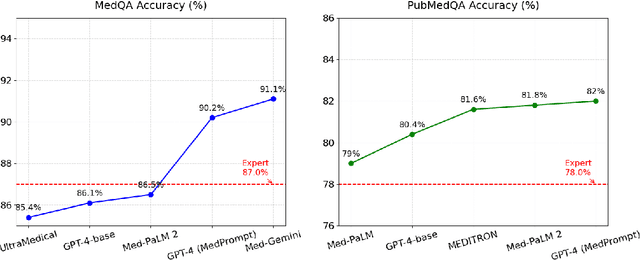
Abstract:Large Language Models (LLMs) have gained significant attention in the medical domain for their human-level capabilities, leading to increased efforts to explore their potential in various healthcare applications. However, despite such a promising future, there are multiple challenges and obstacles that remain for their real-world uses in practical settings. This work discusses key challenges for LLMs in medical applications from four unique aspects: operational vulnerabilities, ethical and social considerations, performance and assessment difficulties, and legal and regulatory compliance. Addressing these challenges is crucial for leveraging LLMs to their full potential and ensuring their responsible integration into healthcare.
Deep Learning Segmentation of Ascites on Abdominal CT Scans for Automatic Volume Quantification
Jun 23, 2024



Abstract:Purpose: To evaluate the performance of an automated deep learning method in detecting ascites and subsequently quantifying its volume in patients with liver cirrhosis and ovarian cancer. Materials and Methods: This retrospective study included contrast-enhanced and non-contrast abdominal-pelvic CT scans of patients with cirrhotic ascites and patients with ovarian cancer from two institutions, National Institutes of Health (NIH) and University of Wisconsin (UofW). The model, trained on The Cancer Genome Atlas Ovarian Cancer dataset (mean age, 60 years +/- 11 [s.d.]; 143 female), was tested on two internal (NIH-LC and NIH-OV) and one external dataset (UofW-LC). Its performance was measured by the Dice coefficient, standard deviations, and 95% confidence intervals, focusing on ascites volume in the peritoneal cavity. Results: On NIH-LC (25 patients; mean age, 59 years +/- 14 [s.d.]; 14 male) and NIH-OV (166 patients; mean age, 65 years +/- 9 [s.d.]; all female), the model achieved Dice scores of 0.855 +/- 0.061 (CI: 0.831-0.878) and 0.826 +/- 0.153 (CI: 0.764-0.887), with median volume estimation errors of 19.6% (IQR: 13.2-29.0) and 5.3% (IQR: 2.4-9.7) respectively. On UofW-LC (124 patients; mean age, 46 years +/- 12 [s.d.]; 73 female), the model had a Dice score of 0.830 +/- 0.107 (CI: 0.798-0.863) and median volume estimation error of 9.7% (IQR: 4.5-15.1). The model showed strong agreement with expert assessments, with r^2 values of 0.79, 0.98, and 0.97 across the test sets. Conclusion: The proposed deep learning method performed well in segmenting and quantifying the volume of ascites in concordance with expert radiologist assessments.
Shadow and Light: Digitally Reconstructed Radiographs for Disease Classification
Jun 06, 2024



Abstract:In this paper, we introduce DRR-RATE, a large-scale synthetic chest X-ray dataset derived from the recently released CT-RATE dataset. DRR-RATE comprises of 50,188 frontal Digitally Reconstructed Radiographs (DRRs) from 21,304 unique patients. Each image is paired with a corresponding radiology text report and binary labels for 18 pathology classes. Given the controllable nature of DRR generation, it facilitates the inclusion of lateral view images and images from any desired viewing position. This opens up avenues for research into new and novel multimodal applications involving paired CT, X-ray images from various views, text, and binary labels. We demonstrate the applicability of DRR-RATE alongside existing large-scale chest X-ray resources, notably the CheXpert dataset and CheXnet model. Experiments demonstrate that CheXnet, when trained and tested on the DRR-RATE dataset, achieves sufficient to high AUC scores for the six common pathologies cited in common literature: Atelectasis, Cardiomegaly, Consolidation, Lung Lesion, Lung Opacity, and Pleural Effusion. Additionally, CheXnet trained on the CheXpert dataset can accurately identify several pathologies, even when operating out of distribution. This confirms that the generated DRR images effectively capture the essential pathology features from CT images. The dataset and labels are publicly accessible at https://huggingface.co/datasets/farrell236/DRR-RATE.
MRISegmentator-Abdomen: A Fully Automated Multi-Organ and Structure Segmentation Tool for T1-weighted Abdominal MRI
May 09, 2024



Abstract:Background: Segmentation of organs and structures in abdominal MRI is useful for many clinical applications, such as disease diagnosis and radiotherapy. Current approaches have focused on delineating a limited set of abdominal structures (13 types). To date, there is no publicly available abdominal MRI dataset with voxel-level annotations of multiple organs and structures. Consequently, a segmentation tool for multi-structure segmentation is also unavailable. Methods: We curated a T1-weighted abdominal MRI dataset consisting of 195 patients who underwent imaging at National Institutes of Health (NIH) Clinical Center. The dataset comprises of axial pre-contrast T1, arterial, venous, and delayed phases for each patient, thereby amounting to a total of 780 series (69,248 2D slices). Each series contains voxel-level annotations of 62 abdominal organs and structures. A 3D nnUNet model, dubbed as MRISegmentator-Abdomen (MRISegmentator in short), was trained on this dataset, and evaluation was conducted on an internal test set and two large external datasets: AMOS22 and Duke Liver. The predicted segmentations were compared against the ground-truth using the Dice Similarity Coefficient (DSC) and Normalized Surface Distance (NSD). Findings: MRISegmentator achieved an average DSC of 0.861$\pm$0.170 and a NSD of 0.924$\pm$0.163 in the internal test set. On the AMOS22 dataset, MRISegmentator attained an average DSC of 0.829$\pm$0.133 and a NSD of 0.908$\pm$0.067. For the Duke Liver dataset, an average DSC of 0.933$\pm$0.015 and a NSD of 0.929$\pm$0.021 was obtained. Interpretation: The proposed MRISegmentator provides automatic, accurate, and robust segmentations of 62 organs and structures in T1-weighted abdominal MRI sequences. The tool has the potential to accelerate research on various clinical topics, such as abnormality detection, radiotherapy, disease classification among others.
Decomposing Vision-based LLM Predictions for Auto-Evaluation with GPT-4
Mar 08, 2024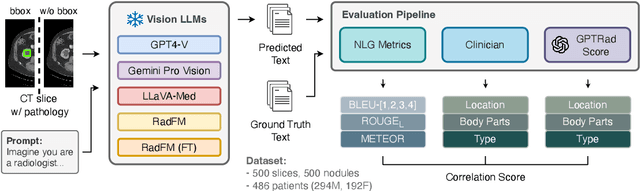

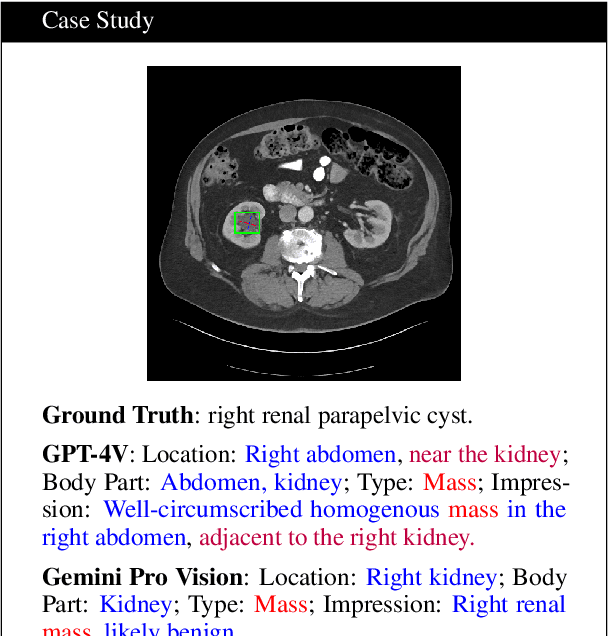

Abstract:The volume of CT exams being done in the world has been rising every year, which has led to radiologist burn-out. Large Language Models (LLMs) have the potential to reduce their burden, but their adoption in the clinic depends on radiologist trust, and easy evaluation of generated content. Presently, many automated methods are available to evaluate the reports generated for chest radiographs, but such an approach is not available for CT presently. In this paper, we propose a novel evaluation framework to judge the capabilities of vision-language LLMs in generating accurate summaries of CT-based abnormalities. CT slices containing an abnormality (e.g., lesion) were input to a vision-based LLM (GPT-4V, LLaVA-Med, and RadFM), and it generated a free-text summary of the predicted characteristics of the abnormality. Next, a GPT-4 model decomposed the summary into specific aspects (body part, location, type, and attributes), automatically evaluated the characteristics against the ground-truth, and generated a score for each aspect based on its clinical relevance and factual accuracy. These scores were then contrasted against those obtained from a clinician, and a high correlation ( 85%, p < .001) was observed. Although GPT-4V outperformed other models in our evaluation, it still requires overall improvement. Our evaluation method offers valuable insights into the specific areas that need the most enhancement, guiding future development in this field.
Leveraging Professional Radiologists' Expertise to Enhance LLMs' Evaluation for Radiology Reports
Feb 02, 2024Abstract:In radiology, Artificial Intelligence (AI) has significantly advanced report generation, but automatic evaluation of these AI-produced reports remains challenging. Current metrics, such as Conventional Natural Language Generation (NLG) and Clinical Efficacy (CE), often fall short in capturing the semantic intricacies of clinical contexts or overemphasize clinical details, undermining report clarity. To overcome these issues, our proposed method synergizes the expertise of professional radiologists with Large Language Models (LLMs), like GPT-3.5 and GPT-4 1. Utilizing In-Context Instruction Learning (ICIL) and Chain of Thought (CoT) reasoning, our approach aligns LLM evaluations with radiologist standards, enabling detailed comparisons between human and AI generated reports. This is further enhanced by a Regression model that aggregates sentence evaluation scores. Experimental results show that our "Detailed GPT-4 (5-shot)" model achieves a 0.48 score, outperforming the METEOR metric by 0.19, while our "Regressed GPT-4" model shows even greater alignment with expert evaluations, exceeding the best existing metric by a 0.35 margin. Moreover, the robustness of our explanations has been validated through a thorough iterative strategy. We plan to publicly release annotations from radiology experts, setting a new standard for accuracy in future assessments. This underscores the potential of our approach in enhancing the quality assessment of AI-driven medical reports.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge