Jianfei Liu
Utility of Pancreas Surface Lobularity as a CT Biomarker for Opportunistic Screening of Type 2 Diabetes
Nov 13, 2025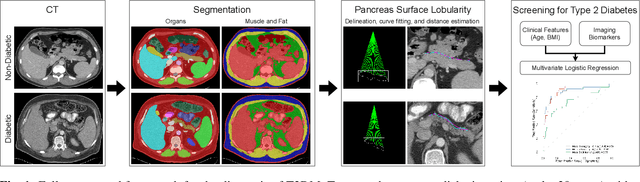
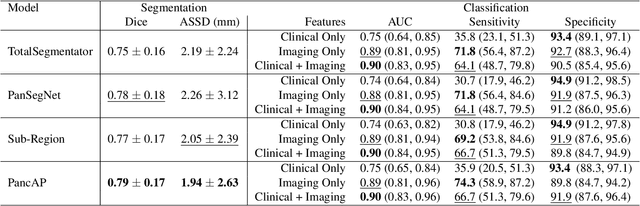
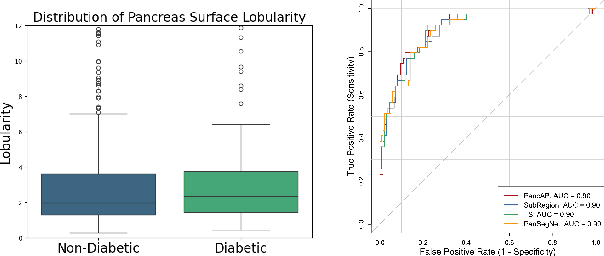
Abstract:Type 2 Diabetes Mellitus (T2DM) is a chronic metabolic disease that affects millions of people worldwide. Early detection is crucial as it can alter pancreas function through morphological changes and increased deposition of ectopic fat, eventually leading to organ damage. While studies have shown an association between T2DM and pancreas volume and fat content, the role of increased pancreatic surface lobularity (PSL) in patients with T2DM has not been fully investigated. In this pilot work, we propose a fully automated approach to delineate the pancreas and other abdominal structures, derive CT imaging biomarkers, and opportunistically screen for T2DM. Four deep learning-based models were used to segment the pancreas in an internal dataset of 584 patients (297 males, 437 non-diabetic, age: 45$\pm$15 years). PSL was automatically detected and it was higher for diabetic patients (p=0.01) at 4.26 $\pm$ 8.32 compared to 3.19 $\pm$ 3.62 for non-diabetic patients. The PancAP model achieved the highest Dice score of 0.79 $\pm$ 0.17 and lowest ASSD error of 1.94 $\pm$ 2.63 mm (p$<$0.05). For predicting T2DM, a multivariate model trained with CT biomarkers attained 0.90 AUC, 66.7\% sensitivity, and 91.9\% specificity. Our results suggest that PSL is useful for T2DM screening and could potentially help predict the early onset of T2DM.
Benchmarking Multi-Organ Segmentation Tools for Multi-Parametric T1-weighted Abdominal MRI
Apr 10, 2025Abstract:The segmentation of multiple organs in multi-parametric MRI studies is critical for many applications in radiology, such as correlating imaging biomarkers with disease status (e.g., cirrhosis, diabetes). Recently, three publicly available tools, such as MRSegmentator (MRSeg), TotalSegmentator MRI (TS), and TotalVibeSegmentator (VIBE), have been proposed for multi-organ segmentation in MRI. However, the performance of these tools on specific MRI sequence types has not yet been quantified. In this work, a subset of 40 volumes from the public Duke Liver Dataset was curated. The curated dataset contained 10 volumes each from the pre-contrast fat saturated T1, arterial T1w, venous T1w, and delayed T1w phases, respectively. Ten abdominal structures were manually annotated in these volumes. Next, the performance of the three public tools was benchmarked on this curated dataset. The results indicated that MRSeg obtained a Dice score of 80.7 $\pm$ 18.6 and Hausdorff Distance (HD) error of 8.9 $\pm$ 10.4 mm. It fared the best ($p < .05$) across the different sequence types in contrast to TS and VIBE.
Leveraging Anatomical Priors for Automated Pancreas Segmentation on Abdominal CT
Apr 09, 2025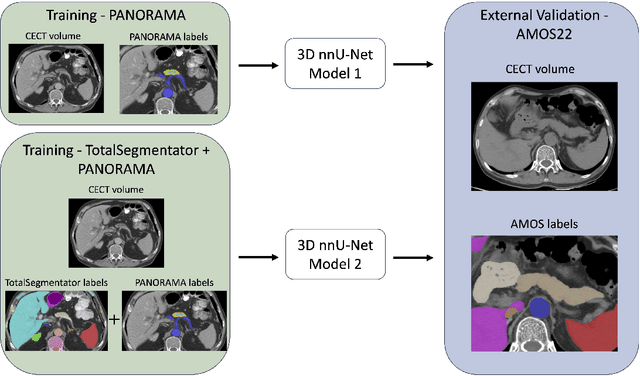


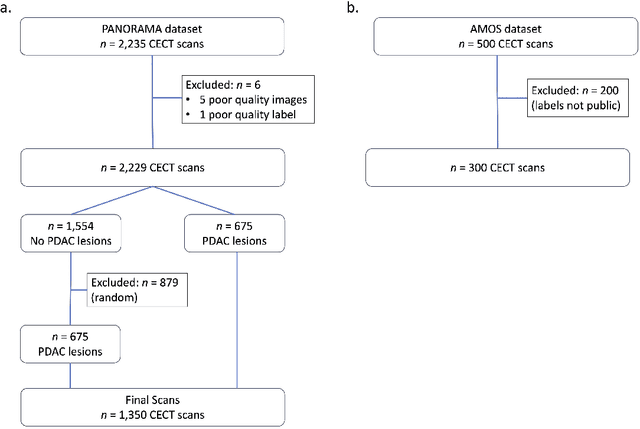
Abstract:An accurate segmentation of the pancreas on CT is crucial to identify pancreatic pathologies and extract imaging-based biomarkers. However, prior research on pancreas segmentation has primarily focused on modifying the segmentation model architecture or utilizing pre- and post-processing techniques. In this article, we investigate the utility of anatomical priors to enhance the segmentation performance of the pancreas. Two 3D full-resolution nnU-Net models were trained, one with 8 refined labels from the public PANORAMA dataset, and another that combined them with labels derived from the public TotalSegmentator (TS) tool. The addition of anatomical priors resulted in a 6\% increase in Dice score ($p < .001$) and a 36.5 mm decrease in Hausdorff distance for pancreas segmentation ($p < .001$). Moreover, the pancreas was always detected when anatomy priors were used, whereas there were 8 instances of failed detections without their use. The use of anatomy priors shows promise for pancreas segmentation and subsequent derivation of imaging biomarkers.
3D Universal Lesion Detection and Tagging in CT with Self-Training
Apr 07, 2025Abstract:Radiologists routinely perform the tedious task of lesion localization, classification, and size measurement in computed tomography (CT) studies. Universal lesion detection and tagging (ULDT) can simultaneously help alleviate the cumbersome nature of lesion measurement and enable tumor burden assessment. Previous ULDT approaches utilize the publicly available DeepLesion dataset, however it does not provide the full volumetric (3D) extent of lesions and also displays a severe class imbalance. In this work, we propose a self-training pipeline to detect 3D lesions and tag them according to the body part they occur in. We used a significantly limited 30\% subset of DeepLesion to train a VFNet model for 2D lesion detection and tagging. Next, the 2D lesion context was expanded into 3D, and the mined 3D lesion proposals were integrated back into the baseline training data in order to retrain the model over multiple rounds. Through the self-training procedure, our VFNet model learned from its own predictions, detected lesions in 3D, and tagged them. Our results indicated that our VFNet model achieved an average sensitivity of 46.9\% at [0.125:8] false positives (FP) with a limited 30\% data subset in comparison to the 46.8\% of an existing approach that used the entire DeepLesion dataset. To our knowledge, we are the first to jointly detect lesions in 3D and tag them according to the body part label.
Correcting Class Imbalances with Self-Training for Improved Universal Lesion Detection and Tagging
Apr 07, 2025Abstract:Universal lesion detection and tagging (ULDT) in CT studies is critical for tumor burden assessment and tracking the progression of lesion status (growth/shrinkage) over time. However, a lack of fully annotated data hinders the development of effective ULDT approaches. Prior work used the DeepLesion dataset (4,427 patients, 10,594 studies, 32,120 CT slices, 32,735 lesions, 8 body part labels) for algorithmic development, but this dataset is not completely annotated and contains class imbalances. To address these issues, in this work, we developed a self-training pipeline for ULDT. A VFNet model was trained on a limited 11.5\% subset of DeepLesion (bounding boxes + tags) to detect and classify lesions in CT studies. Then, it identified and incorporated novel lesion candidates from a larger unseen data subset into its training set, and self-trained itself over multiple rounds. Multiple self-training experiments were conducted with different threshold policies to select predicted lesions with higher quality and cover the class imbalances. We discovered that direct self-training improved the sensitivities of over-represented lesion classes at the expense of under-represented classes. However, upsampling the lesions mined during self-training along with a variable threshold policy yielded a 6.5\% increase in sensitivity at 4 FP in contrast to self-training without class balancing (72\% vs 78.5\%) and a 11.7\% increase compared to the same self-training policy without upsampling (66.8\% vs 78.5\%). Furthermore, we show that our results either improved or maintained the sensitivity at 4FP for all 8 lesion classes.
APDDv2: Aesthetics of Paintings and Drawings Dataset with Artist Labeled Scores and Comments
Nov 13, 2024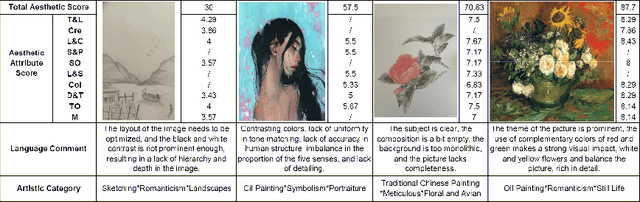
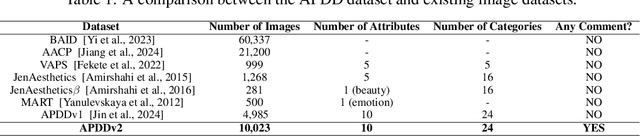
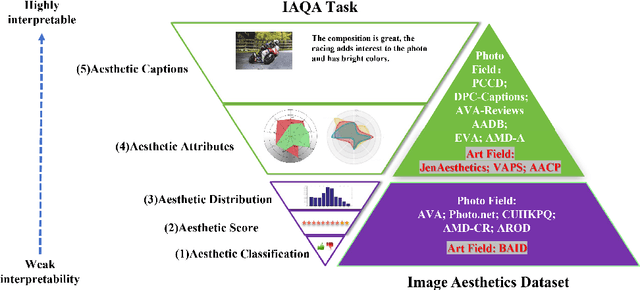
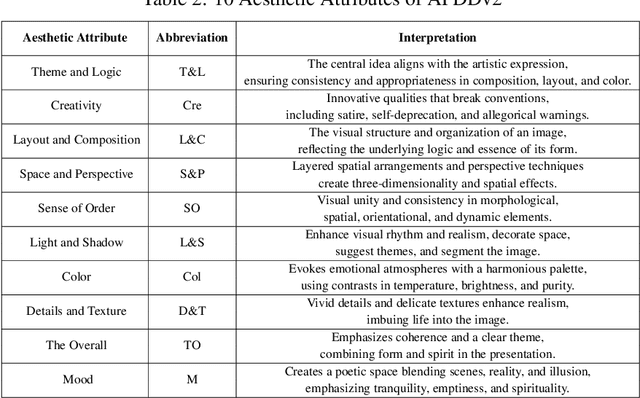
Abstract:Datasets play a pivotal role in training visual models, facilitating the development of abstract understandings of visual features through diverse image samples and multidimensional attributes. However, in the realm of aesthetic evaluation of artistic images, datasets remain relatively scarce. Existing painting datasets are often characterized by limited scoring dimensions and insufficient annotations, thereby constraining the advancement and application of automatic aesthetic evaluation methods in the domain of painting. To bridge this gap, we introduce the Aesthetics Paintings and Drawings Dataset (APDD), the first comprehensive collection of paintings encompassing 24 distinct artistic categories and 10 aesthetic attributes. Building upon the initial release of APDDv1, our ongoing research has identified opportunities for enhancement in data scale and annotation precision. Consequently, APDDv2 boasts an expanded image corpus and improved annotation quality, featuring detailed language comments to better cater to the needs of both researchers and practitioners seeking high-quality painting datasets. Furthermore, we present an updated version of the Art Assessment Network for Specific Painting Styles, denoted as ArtCLIP. Experimental validation demonstrates the superior performance of this revised model in the realm of aesthetic evaluation, surpassing its predecessor in accuracy and efficacy. The dataset and model are available at https://github.com/BestiVictory/APDDv2.git.
Self and Mixed Supervision to Improve Training Labels for Multi-Class Medical Image Segmentation
Mar 06, 2024Abstract:Accurate training labels are a key component for multi-class medical image segmentation. Their annotation is costly and time-consuming because it requires domain expertise. This work aims to develop a dual-branch network and automatically improve training labels for multi-class image segmentation. Transfer learning is used to train the network and improve inaccurate weak labels sequentially. The dual-branch network is first trained by weak labels alone to initialize model parameters. After the network is stabilized, the shared encoder is frozen, and strong and weak decoders are fine-tuned by strong and weak labels together. The accuracy of weak labels is iteratively improved in the fine-tuning process. The proposed method was applied to a three-class segmentation of muscle, subcutaneous and visceral adipose tissue on abdominal CT scans. Validation results on 11 patients showed that the accuracy of training labels was statistically significantly improved, with the Dice similarity coefficient of muscle, subcutaneous and visceral adipose tissue increased from 74.2% to 91.5%, 91.2% to 95.6%, and 77.6% to 88.5%, respectively (p<0.05). In comparison with our earlier method, the label accuracy was also significantly improved (p<0.05). These experimental results suggested that the combination of the dual-branch network and transfer learning is an efficient means to improve training labels for multi-class segmentation.
Automated Plaque Detection and Agatston Score Estimation on Non-Contrast CT Scans: A Multicenter Study
Feb 14, 2024Abstract:Coronary artery calcification (CAC) is a strong and independent predictor of cardiovascular disease (CVD). However, manual assessment of CAC often requires radiological expertise, time, and invasive imaging techniques. The purpose of this multicenter study is to validate an automated cardiac plaque detection model using a 3D multiclass nnU-Net for gated and non-gated non-contrast chest CT volumes. CT scans were performed at three tertiary care hospitals and collected as three datasets, respectively. Heart, aorta, and lung segmentations were determined using TotalSegmentator, while plaques in the coronary arteries and heart valves were manually labeled for 801 volumes. In this work we demonstrate how the nnU-Net semantic segmentation pipeline may be adapted to detect plaques in the coronary arteries and valves. With a linear correction, nnU-Net deep learning methods may also accurately estimate Agatston scores on chest non-contrast CT scans. Compared to manual Agatson scoring, automated Agatston scoring indicated a slope of the linear regression of 0.841 with an intercept of +16 HU (R2 = 0.97). These results are an improvement over previous work assessing automated Agatston score computation in non-gated CT scans.
Weakly Supervised Detection of Pheochromocytomas and Paragangliomas in CT
Feb 12, 2024Abstract:Pheochromocytomas and Paragangliomas (PPGLs) are rare adrenal and extra-adrenal tumors which have the potential to metastasize. For the management of patients with PPGLs, CT is the preferred modality of choice for precise localization and estimation of their progression. However, due to the myriad variations in size, morphology, and appearance of the tumors in different anatomical regions, radiologists are posed with the challenge of accurate detection of PPGLs. Since clinicians also need to routinely measure their size and track their changes over time across patient visits, manual demarcation of PPGLs is quite a time-consuming and cumbersome process. To ameliorate the manual effort spent for this task, we propose an automated method to detect PPGLs in CT studies via a proxy segmentation task. As only weak annotations for PPGLs in the form of prospectively marked 2D bounding boxes on an axial slice were available, we extended these 2D boxes into weak 3D annotations and trained a 3D full-resolution nnUNet model to directly segment PPGLs. We evaluated our approach on a dataset consisting of chest-abdomen-pelvis CTs of 255 patients with confirmed PPGLs. We obtained a precision of 70% and sensitivity of 64.1% with our proposed approach when tested on 53 CT studies. Our findings highlight the promising nature of detecting PPGLs via segmentation, and furthers the state-of-the-art in this exciting yet challenging area of rare cancer management.
Automated Classification of Body MRI Sequence Type Using Convolutional Neural Networks
Feb 12, 2024Abstract:Multi-parametric MRI of the body is routinely acquired for the identification of abnormalities and diagnosis of diseases. However, a standard naming convention for the MRI protocols and associated sequences does not exist due to wide variations in imaging practice at institutions and myriad MRI scanners from various manufacturers being used for imaging. The intensity distributions of MRI sequences differ widely as a result, and there also exists information conflicts related to the sequence type in the DICOM headers. At present, clinician oversight is necessary to ensure that the correct sequence is being read and used for diagnosis. This poses a challenge when specific series need to be considered for building a cohort for a large clinical study or for developing AI algorithms. In order to reduce clinician oversight and ensure the validity of the DICOM headers, we propose an automated method to classify the 3D MRI sequence acquired at the levels of the chest, abdomen, and pelvis. In our pilot work, our 3D DenseNet-121 model achieved an F1 score of 99.5% at differentiating 5 common MRI sequences obtained by three Siemens scanners (Aera, Verio, Biograph mMR). To the best of our knowledge, we are the first to develop an automated method for the 3D classification of MRI sequences in the chest, abdomen, and pelvis, and our work has outperformed the previous state-of-the-art MRI series classifiers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge