Zhizheng Wang
Rethinking Contrastive Learning in Session-based Recommendation
Jun 05, 2025



Abstract:Session-based recommendation aims to predict intents of anonymous users based on limited behaviors. With the ability in alleviating data sparsity, contrastive learning is prevailing in the task. However, we spot that existing contrastive learning based methods still suffer from three obstacles: (1) they overlook item-level sparsity and primarily focus on session-level sparsity; (2) they typically augment sessions using item IDs like crop, mask and reorder, failing to ensure the semantic consistency of augmented views; (3) they treat all positive-negative signals equally, without considering their varying utility. To this end, we propose a novel multi-modal adaptive contrastive learning framework called MACL for session-based recommendation. In MACL, a multi-modal augmentation is devised to generate semantically consistent views at both item and session levels by leveraging item multi-modal features. Besides, we present an adaptive contrastive loss that distinguishes varying contributions of positive-negative signals to improve self-supervised learning. Extensive experiments on three real-world datasets demonstrate the superiority of MACL over state-of-the-art methods.
Knowledge-guided Contextual Gene Set Analysis Using Large Language Models
Jun 04, 2025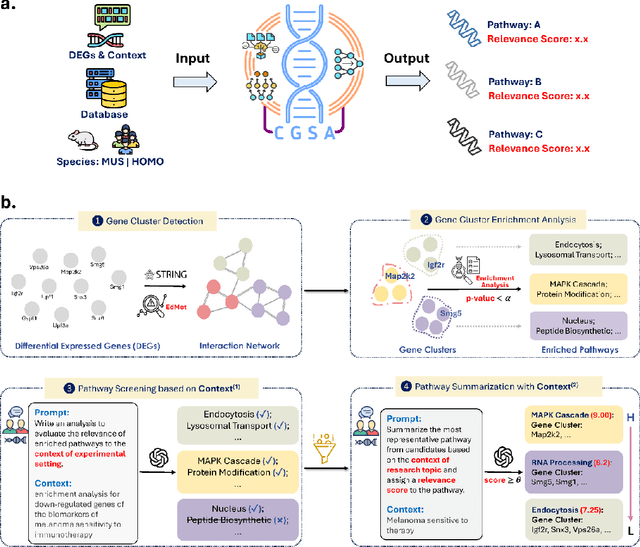

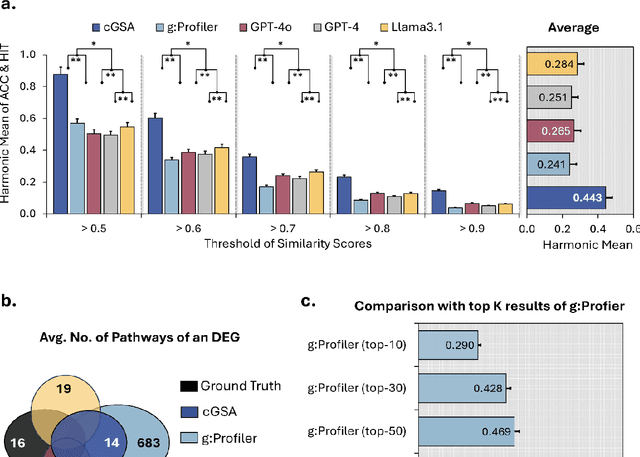

Abstract:Gene set analysis (GSA) is a foundational approach for interpreting genomic data of diseases by linking genes to biological processes. However, conventional GSA methods overlook clinical context of the analyses, often generating long lists of enriched pathways with redundant, nonspecific, or irrelevant results. Interpreting these requires extensive, ad-hoc manual effort, reducing both reliability and reproducibility. To address this limitation, we introduce cGSA, a novel AI-driven framework that enhances GSA by incorporating context-aware pathway prioritization. cGSA integrates gene cluster detection, enrichment analysis, and large language models to identify pathways that are not only statistically significant but also biologically meaningful. Benchmarking on 102 manually curated gene sets across 19 diseases and ten disease-related biological mechanisms shows that cGSA outperforms baseline methods by over 30%, with expert validation confirming its increased precision and interpretability. Two independent case studies in melanoma and breast cancer further demonstrate its potential to uncover context-specific insights and support targeted hypothesis generation.
Beyond Multiple-Choice Accuracy: Real-World Challenges of Implementing Large Language Models in Healthcare
Oct 24, 2024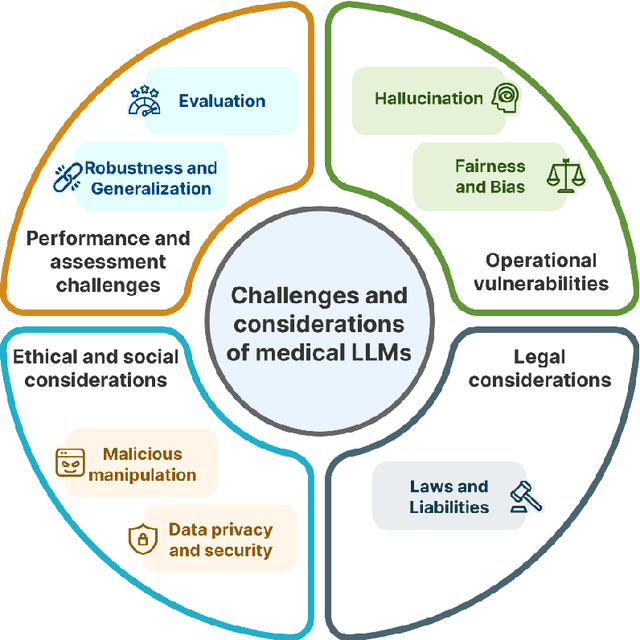
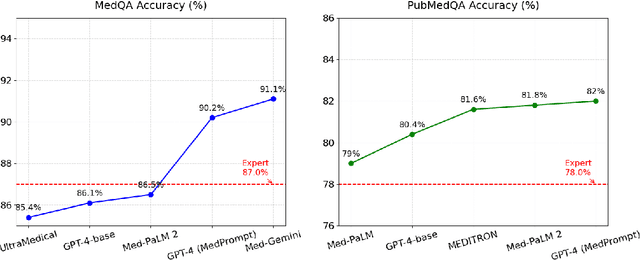
Abstract:Large Language Models (LLMs) have gained significant attention in the medical domain for their human-level capabilities, leading to increased efforts to explore their potential in various healthcare applications. However, despite such a promising future, there are multiple challenges and obstacles that remain for their real-world uses in practical settings. This work discusses key challenges for LLMs in medical applications from four unique aspects: operational vulnerabilities, ethical and social considerations, performance and assessment difficulties, and legal and regulatory compliance. Addressing these challenges is crucial for leveraging LLMs to their full potential and ensuring their responsible integration into healthcare.
Demystifying Large Language Models for Medicine: A Primer
Oct 24, 2024



Abstract:Large language models (LLMs) represent a transformative class of AI tools capable of revolutionizing various aspects of healthcare by generating human-like responses across diverse contexts and adapting to novel tasks following human instructions. Their potential application spans a broad range of medical tasks, such as clinical documentation, matching patients to clinical trials, and answering medical questions. In this primer paper, we propose an actionable guideline to help healthcare professionals more efficiently utilize LLMs in their work, along with a set of best practices. This approach consists of several main phases, including formulating the task, choosing LLMs, prompt engineering, fine-tuning, and deployment. We start with the discussion of critical considerations in identifying healthcare tasks that align with the core capabilities of LLMs and selecting models based on the selected task and data, performance requirements, and model interface. We then review the strategies, such as prompt engineering and fine-tuning, to adapt standard LLMs to specialized medical tasks. Deployment considerations, including regulatory compliance, ethical guidelines, and continuous monitoring for fairness and bias, are also discussed. By providing a structured step-by-step methodology, this tutorial aims to equip healthcare professionals with the tools necessary to effectively integrate LLMs into clinical practice, ensuring that these powerful technologies are applied in a safe, reliable, and impactful manner.
GeneAgent: Self-verification Language Agent for Gene Set Knowledge Discovery using Domain Databases
May 25, 2024



Abstract:Gene set knowledge discovery is essential for advancing human functional genomics. Recent studies have shown promising performance by harnessing the power of Large Language Models (LLMs) on this task. Nonetheless, their results are subject to several limitations common in LLMs such as hallucinations. In response, we present GeneAgent, a first-of-its-kind language agent featuring self-verification capability. It autonomously interacts with various biological databases and leverages relevant domain knowledge to improve accuracy and reduce hallucination occurrences. Benchmarking on 1,106 gene sets from different sources, GeneAgent consistently outperforms standard GPT-4 by a significant margin. Moreover, a detailed manual review confirms the effectiveness of the self-verification module in minimizing hallucinations and generating more reliable analytical narratives. To demonstrate its practical utility, we apply GeneAgent to seven novel gene sets derived from mouse B2905 melanoma cell lines, with expert evaluations showing that GeneAgent offers novel insights into gene functions and subsequently expedites knowledge discovery.
Decomposing Vision-based LLM Predictions for Auto-Evaluation with GPT-4
Mar 08, 2024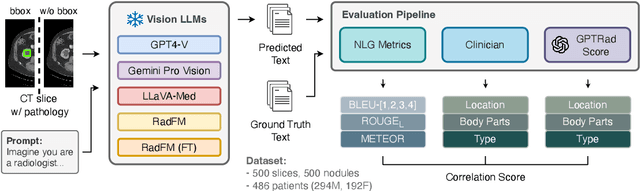

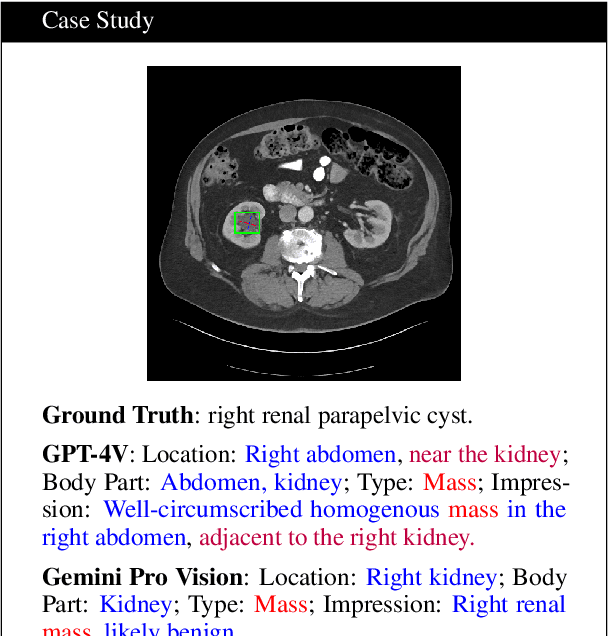

Abstract:The volume of CT exams being done in the world has been rising every year, which has led to radiologist burn-out. Large Language Models (LLMs) have the potential to reduce their burden, but their adoption in the clinic depends on radiologist trust, and easy evaluation of generated content. Presently, many automated methods are available to evaluate the reports generated for chest radiographs, but such an approach is not available for CT presently. In this paper, we propose a novel evaluation framework to judge the capabilities of vision-language LLMs in generating accurate summaries of CT-based abnormalities. CT slices containing an abnormality (e.g., lesion) were input to a vision-based LLM (GPT-4V, LLaVA-Med, and RadFM), and it generated a free-text summary of the predicted characteristics of the abnormality. Next, a GPT-4 model decomposed the summary into specific aspects (body part, location, type, and attributes), automatically evaluated the characteristics against the ground-truth, and generated a score for each aspect based on its clinical relevance and factual accuracy. These scores were then contrasted against those obtained from a clinician, and a high correlation ( 85%, p < .001) was observed. Although GPT-4V outperformed other models in our evaluation, it still requires overall improvement. Our evaluation method offers valuable insights into the specific areas that need the most enhancement, guiding future development in this field.
AgentMD: Empowering Language Agents for Risk Prediction with Large-Scale Clinical Tool Learning
Feb 20, 2024Abstract:Clinical calculators play a vital role in healthcare by offering accurate evidence-based predictions for various purposes such as prognosis. Nevertheless, their widespread utilization is frequently hindered by usability challenges, poor dissemination, and restricted functionality. Augmenting large language models with extensive collections of clinical calculators presents an opportunity to overcome these obstacles and improve workflow efficiency, but the scalability of the manual curation process poses a significant challenge. In response, we introduce AgentMD, a novel language agent capable of curating and applying clinical calculators across various clinical contexts. Using the published literature, AgentMD has automatically curated a collection of 2,164 diverse clinical calculators with executable functions and structured documentation, collectively named RiskCalcs. Manual evaluations show that RiskCalcs tools achieve an accuracy of over 80% on three quality metrics. At inference time, AgentMD can automatically select and apply the relevant RiskCalcs tools given any patient description. On the newly established RiskQA benchmark, AgentMD significantly outperforms chain-of-thought prompting with GPT-4 (87.7% vs. 40.9% in accuracy). Additionally, we also applied AgentMD to real-world clinical notes for analyzing both population-level and risk-level patient characteristics. In summary, our study illustrates the utility of language agents augmented with clinical calculators for healthcare analytics and patient care.
PubTator 3.0: an AI-powered Literature Resource for Unlocking Biomedical Knowledge
Jan 19, 2024

Abstract:PubTator 3.0 (https://www.ncbi.nlm.nih.gov/research/pubtator3/) is a biomedical literature resource using state-of-the-art AI techniques to offer semantic and relation searches for key concepts like proteins, genetic variants, diseases, and chemicals. It currently provides over one billion entity and relation annotations across approximately 36 million PubMed abstracts and 6 million full-text articles from the PMC open access subset, updated weekly. PubTator 3.0's online interface and API utilize these precomputed entity relations and synonyms to provide advanced search capabilities and enable large-scale analyses, streamlining many complex information needs. We showcase the retrieval quality of PubTator 3.0 using a series of entity pair queries, demonstrating that PubTator 3.0 retrieves a greater number of articles than either PubMed or Google Scholar, with higher precision in the top 20 results. We further show that integrating ChatGPT (GPT-4) with PubTator APIs dramatically improves the factuality and verifiability of its responses. In summary, PubTator 3.0 offers a comprehensive set of features and tools that allow researchers to navigate the ever-expanding wealth of biomedical literature, expediting research and unlocking valuable insights for scientific discovery.
A Note on Graph-Based Nearest Neighbor Search
Dec 21, 2020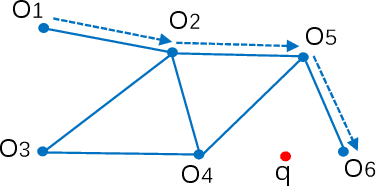
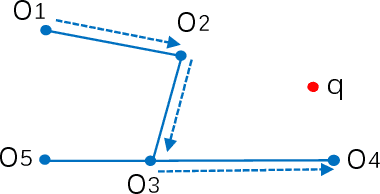
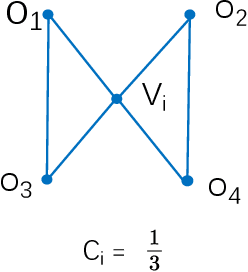
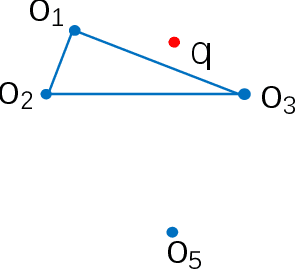
Abstract:Nearest neighbor search has found numerous applications in machine learning, data mining and massive data processing systems. The past few years have witnessed the popularity of the graph-based nearest neighbor search paradigm because of its superiority over the space-partitioning algorithms. While a lot of empirical studies demonstrate the efficiency of graph-based algorithms, not much attention has been paid to a more fundamental question: why graph-based algorithms work so well in practice? And which data property affects the efficiency and how? In this paper, we try to answer these questions. Our insight is that "the probability that the neighbors of a point o tends to be neighbors in the KNN graph" is a crucial data property for query efficiency. For a given dataset, such a property can be qualitatively measured by clustering coefficient of the KNN graph. To show how clustering coefficient affects the performance, we identify that, instead of the global connectivity, the local connectivity around some given query q has more direct impact on recall. Specifically, we observed that high clustering coefficient makes most of the k nearest neighbors of q sit in a maximum strongly connected component (SCC) in the graph. From the algorithmic point of view, we show that the search procedure is actually composed of two phases - the one outside the maximum SCC and the other one in it, which is different from the widely accepted single or multiple paths search models. We proved that the commonly used graph-based search algorithm is guaranteed to traverse the maximum SCC once visiting any point in it. Our analysis reveals that high clustering coefficient leads to large size of the maximum SCC, and thus provides good answer quality with the help of the two-phase search procedure. Extensive empirical results over a comprehensive collection of datasets validate our findings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge