Nick Byrne
Right Regions, Wrong Labels: Semantic Label Flips in Segmentation under Correlation Shift
Apr 14, 2026Abstract:The robustness of machine learning models can be compromised by spurious correlations between non-causal features in the input data and target labels. A common way to test for such correlations is to train on data where the label is strongly tied to some non-causal cue, then evaluate on examples where that tie no longer holds. This idea is well established for classification tasks, but for semantic segmentation the specific failure modes are not well understood. We show that a model may achieve reasonable overlap while assigning the wrong semantic label, swapping one plausible foreground class for another, even when object boundaries are largely correct. We focus on this semantic label-flip behaviour and quantify it with a simple diagnostic (Flip) that counts how often ground truth foreground pixels are assigned the wrong foreground identity while remaining predicted as foreground. In a setting where category and scene are correlated during training, increasing the correlation consistently widens the gap between common and rare test conditions and increases these within-object label swaps on counterfactual groups. Overall, our results motivate assessing segmentation robustness under distribution shift beyond overlap by decomposing foreground errors into correct pixels, flipped-identity pixels, and missed-to-background pixels. We also propose an entropy-based, ground truth label-free `flip-risk' score, which is computed from foreground identity uncertainty, and show that it can flag flip-prone cases at inference time. Code is available at https://github.com/acharaakshit/label-flips.
A persistent homology-based topological loss for CNN-based multi-class segmentation of CMR
Jul 27, 2021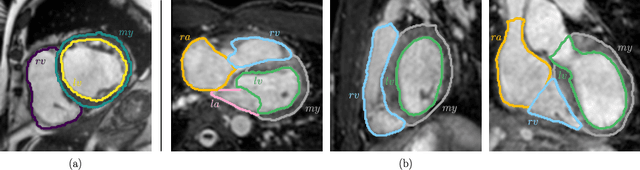
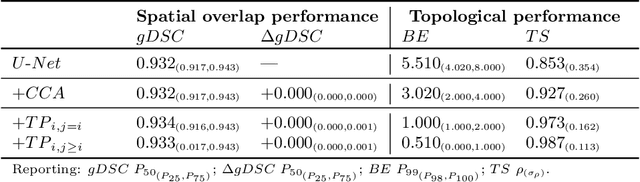
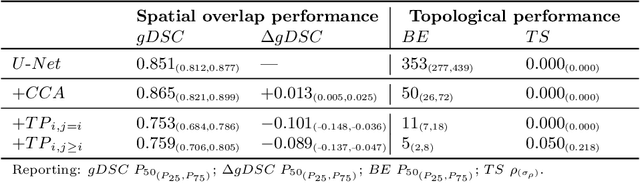

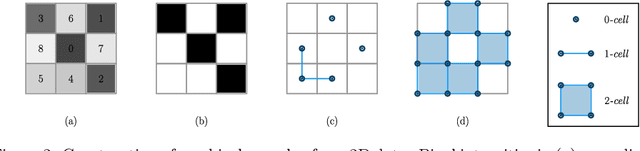

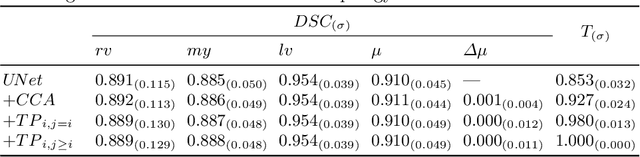
Abstract:Multi-class segmentation of cardiac magnetic resonance (CMR) images seeks a separation of data into anatomical components with known structure and configuration. The most popular CNN-based methods are optimised using pixel wise loss functions, ignorant of the spatially extended features that characterise anatomy. Therefore, whilst sharing a high spatial overlap with the ground truth, inferred CNN-based segmentations can lack coherence, including spurious connected components, holes and voids. Such results are implausible, violating anticipated anatomical topology. In response, (single-class) persistent homology-based loss functions have been proposed to capture global anatomical features. Our work extends these approaches to the task of multi-class segmentation. Building an enriched topological description of all class labels and class label pairs, our loss functions make predictable and statistically significant improvements in segmentation topology using a CNN-based post-processing framework. We also present (and make available) a highly efficient implementation based on cubical complexes and parallel execution, enabling practical application within high resolution 3D data for the first time. We demonstrate our approach on 2D short axis and 3D whole heart CMR segmentation, advancing a detailed and faithful analysis of performance on two publicly available datasets.
A persistent homology-based topological loss function for multi-class CNN segmentation of cardiac MRI
Aug 21, 2020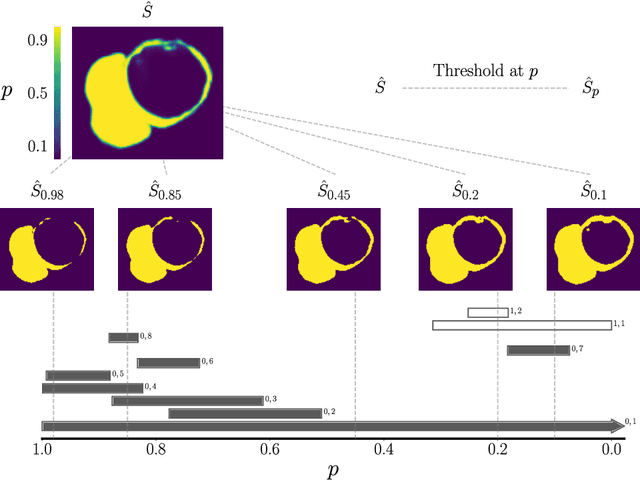
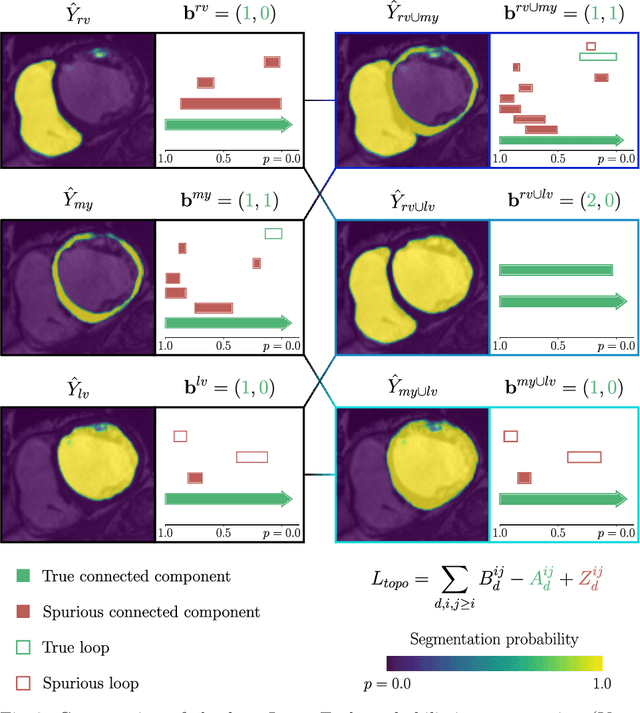
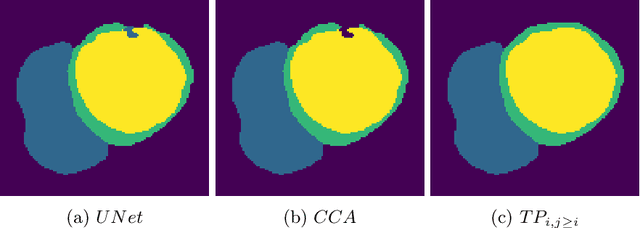
Abstract:With respect to spatial overlap, CNN-based segmentation of short axis cardiovascular magnetic resonance (CMR) images has achieved a level of performance consistent with inter observer variation. However, conventional training procedures frequently depend on pixel-wise loss functions, limiting optimisation with respect to extended or global features. As a result, inferred segmentations can lack spatial coherence, including spurious connected components or holes. Such results are implausible, violating the anticipated topology of image segments, which is frequently known a priori. Addressing this challenge, published work has employed persistent homology, constructing topological loss functions for the evaluation of image segments against an explicit prior. Building a richer description of segmentation topology by considering all possible labels and label pairs, we extend these losses to the task of multi-class segmentation. These topological priors allow us to resolve all topological errors in a subset of 150 examples from the ACDC short axis CMR training data set, without sacrificing overlap performance.
Topology-preserving augmentation for CNN-based segmentation of congenital heart defects from 3D paediatric CMR
Aug 23, 2019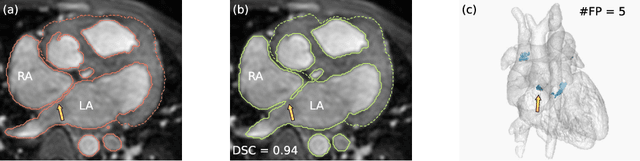
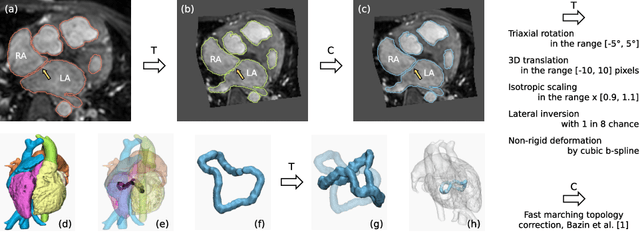
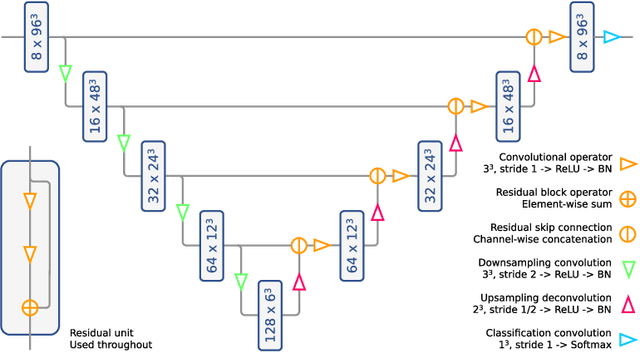
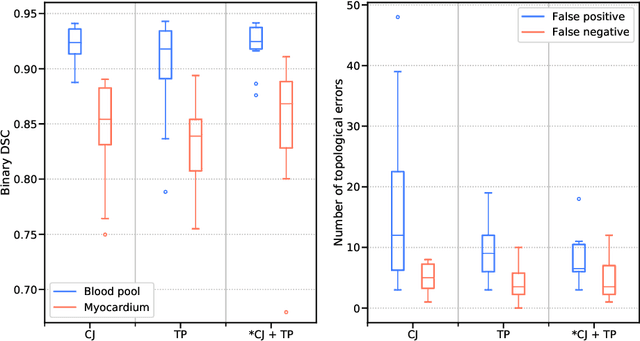
Abstract:Patient-specific 3D printing of congenital heart anatomy demands an accurate segmentation of the thin tissue interfaces which characterise these diagnoses. Even when a label set has a high spatial overlap with the ground truth, inaccurate delineation of these interfaces can result in topological errors. These compromise the clinical utility of such models due to the anomalous appearance of defects. CNNs have achieved state-of-the-art performance in segmentation tasks. Whilst data augmentation has often played an important role, we show that conventional image resampling schemes used therein can introduce topological changes in the ground truth labelling of augmented samples. We present a novel pipeline to correct for these changes, using a fast-marching algorithm to enforce the topology of the ground truth labels within their augmented representations. In so doing, we invoke the idea of cardiac contiguous topology to describe an arbitrary combination of congenital heart defects and develop an associated, clinically meaningful metric to measure the topological correctness of segmentations. In a series of five-fold cross-validations, we demonstrate the performance gain produced by this pipeline and the relevance of topological considerations to the segmentation of congenital heart defects. We speculate as to the applicability of this approach to any segmentation task involving morphologically complex targets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge